We carried out our coronary heart failure prediction pipeline in Python and included publicly obtainable language fashions from Hugging Face ( Our pipeline required GPU assets for in depth computations with language fashions, which have been carried out on the Excessive-Efficiency Cluster of the RWTH Aachen College with entry to GPU computing assets powered by the NVIDIA H100 Tensor-Core-GPU.
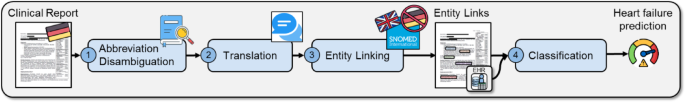
Our coronary heart failure prediction pipeline consists of 4 steps: (1) medical abbreviation disambiguation, (2) translation of German scientific notes to English, (3) medical entity linking of scientific notes to SNOMED-CT ideas, and (4) classification of sufferers based mostly on their EHR and entity hyperlink knowledge to diagnose coronary heart failure.
Disambiguating medical abbreviations
Abbreviation disambiguation approaches use abbreviation dictionaries containing medical abbreviations and their expansions. Our strategy disambiguates abbreviations in English and German scientific notes with two abbreviation dictionaries, since medical abbreviations are language-dependent. For English scientific notes, we compiled a complete abbreviation-expansion dictionary consisting of abbreviations from three totally different sources.
-
1.
An overtly obtainable, manually curated English medical abbreviation-expansion dictionary from present analysis, overlaying 5 information sources with a complete of 3758 distinctive abbreviations and 5794 distinctive abbreviation–growth pairs13. This dataset was created utilizing web-scale reverse substitution (WSRS), which substitutes medical abbreviations with their expansions in web-derived sentences, facilitating scientific abbreviation detection and growth. For simplicity, this dataset is recognized as WSRS Scientific Abbreviation dataset.
-
2.
The Scientific Abbreviation Sense Stock (CASI) dataset17, which dietary supplements the WSRS dataset with further abbreviation-expansion pairs.
-
3.
Abbreviation-expansion pairs from the UMLS, which have been extracted mechanically by analyzing UMLS phrases containing hyphens. For example, within the abbreviation “AP” with the growth “abdominal pain” within the UMLS time period “AP – abdominal pain” was recognized for the reason that letters of the abbreviation matched the primary letters of the time period “abdominal pain” after the hyphen.
For the abbreviation disambiguation in German scientific notes, we supplemented 110 abbreviations supplied by the Division of Inside Drugs I on the College Hospital Aachen with 1757 abbreviations from the medical affiliation of Mecklenburg Vorpommern, which have been aggregated for the competence coaching of physicians58.
To allow context-dependent disambiguation of abbreviations in our pipeline, we collected 30 context samples for every abbreviation, together with surrounding context from English scientific notes within the established MIMIC-III dataset23. The variety of 30 samples was chosen based mostly on findings in earlier analysis on efficient context-based disambiguation14. For the disambiguation of German abbreviations, we aggregated 30 context samples from the overtly obtainable GGPONC dataset24, which represents one of many largest German datasets with medical paperwork based mostly on scientific observe tips for oncology.
The abbreviation disambiguation in our strategy entails two steps: first, figuring out an abbreviation via widespread sample matching based mostly on recognized abbreviations in our abbreviation-expansion dictionary, and second, increasing the abbreviation based mostly on its surrounding context (Fig. 4). To disambiguate the abbreviation, the encompassing context of the abbreviation within the sentence is in contrast by a language mannequin pre-trained on semantic similarity (SS-LM) with a context assortment for doable expansions of the abbreviation, following prior analysis59.
Overview of the abbreviation disambiguation strategy. Our zero-shot abbreviation disambiguation strategy detects abbreviations in scientific notes via sample matching in step one and expands abbreviations within the second textual content via context comparability. On this instance, step one of our strategy identifies the abbreviation “HT” via sample matching within the enter phrase and finds the growth candidates “hypertension” and “head trauma” via the supplied abbreviation dictionary. Within the second step, context samples for each growth candidates are in contrast relating to semantic similarity via a language mannequin with the context supplied via the enter phrase to pick the growth “hypertension” with essentially the most comparable context.
We use the pre-trained SS-LM BioLORD60 for English textual content and paraphrase-multilingual-MiniLM-L12-v261 for German textual content. BioLORD was particularly skilled to generate representations of scientific sentences and biomedical ideas from UMLS and SNOMED-CT, outperforming different state-of-the-art fashions in duties corresponding to semantic textual similarity as computed via cosine similarity, biomedical idea illustration and biomedical idea title normalization60.
The SS-LM computes the similarity between every context pattern for a given growth candidate and the present textual content snippet, as an illustration, a sentence. The growth candidate with the very best similarity is chosen. We apply a threshold filter of 0.3 based mostly on earlier analysis to solely broaden an abbreviation, if the context has a cosine similarity of 0.3 or above, to keep away from expansions when no comparable context justifies them14.
We measured the abbreviation disambiguation efficiency via recall, precision, growth accuracy and complete accuracy. Recall displays the proportion of appropriately retrieved floor reality annotations, for instance, abbreviations or idea identifiers with corresponding spans, and is outlined as
$$:Recall:=frac{True:positives}{True:positives+False:negatives}$$
(1)
.
The precision signifies how lots of the retrieved gadgets are right and is outlined as
$$:Precision:=frac{True:positives}{True:positives+False:positives}$$
(2)
.
The growth accuracy is outlined as
$$:Growth:accuracy=left(frac{textual content{C}textual content{o}textual content{r}textual content{r}textual content{e}textual content{c}textual content{t}:textual content{e}textual content{x}textual content{p}textual content{a}textual content{n}textual content{s}textual content{i}textual content{o}textual content{n}}{textual content{C}textual content{o}textual content{r}textual content{r}textual content{e}textual content{c}textual content{t}:textual content{e}textual content{x}textual content{p}textual content{a}textual content{n}textual content{s}textual content{i}textual content{o}textual content{n}+Incorrect:Growth}proper)$$
(3)
and the overall accuracy outlined as
$$:Whole:accuracy=left(Recallcdot:Growth:accuracyright),$$
(4)
to make sure comparability with earlier analysis13. Furthermore, the harmonic imply of recall and precision varieties the F1 rating and is outlined as:
$$:F1_Score=2left(frac{Precision:cdot::Recall}{Precision+Recall}proper).$$
(5)
The efficiency of our abbreviation disambiguation in English was evaluated on the CASI dataset17 and the WSRS Scientific Abbreviation dataset13 (Fig. 5). We shaped a subset of the CASI dataset for the analysis to align our experiment with present analysis, which resulted in 21,564 textual content snippets containing 63 distinctive abbreviations and 119 distinctive abbreviation expansions13.
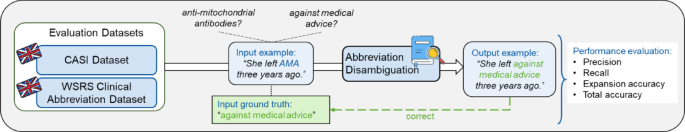
Overview of the abbreviation disambiguation analysis. The abbreviation disambiguation step was evaluated on the exterior CASI and WSRS Scientific Abbreviation datasets with the efficiency metrics precision, recall, growth accuracy, and complete accuracy. Particular person knowledge factors of each datasets consisting of phrases with abbreviations are disambiguated by the pipeline. The output phrase with an expanded abbreviation is checked in opposition to the bottom reality from the datasets to compute the efficiency metrics.
Translation of scientific notes
For the interpretation of our German scientific notes to English, we use the open-source language mannequin fb/wmt19-en-de on account of its open supply availability and excessive efficiency with a BiLingual Analysis Understudy (BLEU) rating of 40.862. Scientific notes have been divided into smaller chunks for the interpretation to handle enter restrictions relating to the utmost variety of processable tokens of the language mannequin. After translation, the concatenated chunks type the translated scientific word. The interpretation step isn’t evaluated individually, as we didn’t modify the interpretation mannequin and seek advice from its printed efficiency62.
Medical entity linking
Preparation of terminology knowledge
To arrange our entity linking strategy with the English scientific notes, we parsed the terminologies SNOMED-CT and UMLS. For every idea within the terminologies, we gathered the idea descriptions, such because the synonyms in addition to the identifier and class in an idea database. To facilitate quick lookup for comparable idea descriptions for a given textual content portion, we embedded and saved all idea descriptions as vectors utilizing respective SS-LMs. Totally different ideas, e.g. with totally different identifiers, hierarchy positions and relations, could be related via the identical description, e.g. synonym, leading to round 40% of ideas within the MedMentions dataset requiring disambiguation14. We remedy this problem analogous to the abbreviation disambiguation by leveraging aggregated context samples for every ambiguous idea description. To make sure comparability of our outcomes with printed MedCAT outcomes, we use the UMLS 2018AB launch. For the entity linking in our German scientific notes to SNOMED-CT, we use the SNOMED-CT Worldwide Might 2023 launch.
SNOMED-CT idea candidate identification
The SNOMED-CT idea candidate identification within the entity linking course of comprises 4 steps (Fig. 6).
First, our pipeline identifies recognized abbreviations from the abbreviation-expansion dictionary and shops positions and expansions of the abbreviations in an abbreviation overview desk. The longest abbreviations are matched first to stop the untimely growth of nested abbreviations.
Second, the scientific word is tokenized by splitting up strings surrounded by non-alphanumeric characters to type n-grams, phrases of sequentially occurring n adjoining tokens, such because the 2-gram “the patient”.
Third, it removes phrases containing prohibited infixes, which could be outlined per use case. In our use case, full stops or hyphens adopted by a whitespace as infixes are prohibited since they’ll point out a time period spanning two sentences or bullet factors, main presumably to inaccurate entity hyperlinks. Abbreviations within the remaining n-grams are expanded based mostly on the desk created in step one. The growth at this stage ensures that the beginning and finish positions of phrases could be saved earlier than modifications, which retains the saved positions aligned with the positions from analysis datasets.
Fourth, we retrieve the top-k most comparable SNOMED-CT idea descriptions for every n-gram based mostly on the cosine similarity of the respective SS-LM embedding. Retrieving a number of candidates improves recall potential, as the right candidate, in accordance with floor reality annotations, might not all the time be the top-ranked match.
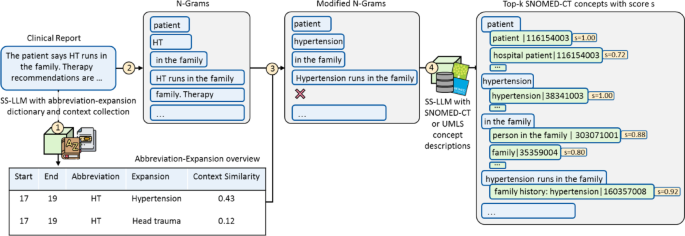
Overview of candidate identification steps. The candidate identification within the entity linking course of comprises 4 steps. In step one on this instance, the abbreviation “HT” within the scientific word is recognized via sample matching and disambiguated based mostly on its surrounding context. Within the second step, n-grams corresponding to “HT runs in the family” are shaped. Within the third step, abbreviations are expanded in every n-gram based mostly on the abbreviation-expansion overview, leading to modified n-grams corresponding to “Hypertension runs in the family”. N-grams corresponding to “family. Therapy” are dropped since a forbidden infix, a full cease, is contained. Within the fourth step, for every n-gram the top-k most comparable SNOMED-CT descriptions with respective idea identifiers are decided via the language mannequin pre-trained for semantic similarity (SS-LM).
SNOMED-CT idea candidate choice
The SNOMED-CT idea candidate choice consists of 4 steps to cut back redundancy and choose essentially the most correct ideas for the entity hyperlinks from the beforehand generated candidates. This process allows the omission of redundant and out of date ideas whereas adapting to knowledge akin to particular use circumstances by way of iterative on-line studying. The method makes use of the entity hyperlink candidates from the scientific word of the earlier step as enter (Fig. 7).
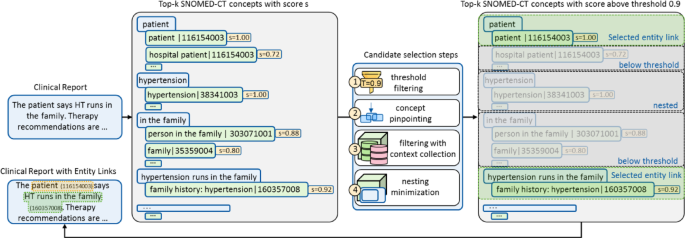
Overview of candidate choice steps. The candidate choice course of consists of 4 sequential steps to refine entity hyperlinks by eradicating out of date and redundant candidates, guaranteeing solely essentially the most correct hyperlinks are retained. (1) threshold filtering: Ideas with similarity scores under a predefined threshold, e.g., 0.9, are discarded. For instance, the n-gram “patient” initially hyperlinks to each “hospital patient” and “patient,” however solely “patient” is retained as its similarity rating of 1.0 exceeds the brink. (2) idea pinpointing: if a number of n-grams hyperlink to the identical idea, solely essentially the most consultant one is saved. For example, each “The patient” and “patient” might hyperlink to “patient | 116154003”, however solely “patient” is retained. (3) filtering with context-collection: ideas are additional refined based mostly on surrounding contextual data. (4) nesting minimization: entity hyperlinks containing semantically nested ideas are eliminated. For instance, “hypertension | 38341003” is nested inside “family history: hypertension | 160357008” and is due to this fact eradicated.
Step one removes any entity hyperlinks with a cosine similarity under an outlined similarity threshold, guaranteeing that solely extremely comparable ideas are linked to the spans within the textual content. The second step pinpoints the identical ideas, that are linked in a number of overlapping n-grams with various similarity to the n-gram with the very best similarity whereas the opposite hyperlinks are eliminated. The third step removes out of date ideas via a group of constructive and unfavorable context samples for ideas alongside the SS-LM, to resolve redundancy in entity hyperlinks of ambiguous idea descriptions and allow on-line studying for improved efficiency. We use the aggregated context from scientific notes in MIMIC-III for ambiguous idea descriptions, which hyperlink to multiple idea within the goal terminology. These context samples function constructive context and are supplemented through the entity linking. Damaging context is supplied via cases the place our pipeline incorrectly linked a free-text portion to an idea that was not linked within the floor reality dataset. If an idea has solely unfavorable context samples, the pipeline removes entity hyperlinks containing that idea to keep away from repeated incorrect hyperlinks. Any ideas linked within the floor reality dataset are added to the constructive context assortment, alongside their surrounding context. If an idea in an entity hyperlink has each constructive and unfavorable context samples, it’s dismissed if the very best unfavorable context similarity exceeds the very best constructive context similarity within the present context. The danger of information leakage is addressed by guaranteeing processing of the coaching dataset prior earlier than processing the check dataset and solely including context samples after the respective entity linking course of. The analysis of entity hyperlinks computed after processing a scientific word can enhance efficiency in an online-learning style and tailor our pipeline mechanically to particular datasets or use-cases.
The fourth step minimizes nesting wherever doable. Our pipeline analyzes whether or not semantic nesting is current or whether or not nesting is justified on account of various kinds of data being linked. The pipeline evaluates how omitting the nested n-gram modifications the similarity of the outer entity hyperlink to its goal idea. If an outer n-gram with no nested n-gram yields a decrease similarity to the goal idea, it signifies that the nested n-gram is semantically nested data. Subsequently, the nested entity hyperlink is dismissed. An elevated similarity as an alternative would point out no nesting and the presence of recent data within the entity hyperlink, justifying the nested entity hyperlinks to stop data loss.
Validation
We evaluated the entity linking efficiency in English and German scientific notes utilizing three totally different datasets (Fig. 8). For English, we validated our technique on the established MedMentions and ShARe/CLEF 2014 datasets with the goal terminologies UMLS and SNOMED-CT and efficiency metrics recall, precision, and F1-score (Eqs. (1), (2), (5)). For German, we assessed the entity linking efficiency on the DARIO dataset with the goal terminology SNOMED-CT via an skilled survey involving three cardiologists.
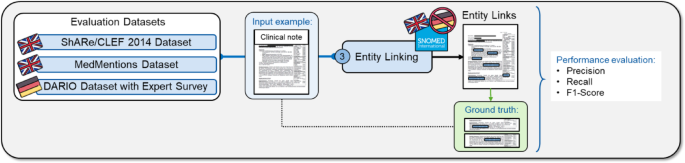
Overview of the analysis of the entity linking step. The entity linking step was evaluated on the exterior datasets ShARe/CLEF 2014, MedMentions, and our inner DARIO dataset with the efficiency metrics recall, precision, and F1-score. On this instance, a scientific word is an enter for the entity linking step, which then hyperlinks medical entities within the free textual content to SNOMED-CT ideas. The entity linking efficiency is evaluated via the efficiency metrics by evaluating the output with the linked entities within the floor reality.
MedMentions
MedMentions consists of over 4,000 abstracts of PubMed publications with greater than 350,000 manually linked entities19. Mentions of medical entities within the abstracts have been linked by skilled annotators to ideas of the UMLS 2017AA model, which comprises round 3.2 million distinctive ideas19. For our analysis, we used the pre-defined “disorders only” subset as outlined within the dataset description to concentrate on health-related ideas and omit different biomedical ideas.
ShARe/CLEF 2014
ShARe/CLEF 2014 contains 433 de-identified scientific notes from over 30,000 sufferers admitted to the intensive-care unit (ICU), which have been annotated by 15 scientific professionals and informatics specialists63. We use the event set containing 300 paperwork of 4 differing kinds: discharge abstract, radiology report, electrocardiogram and echocardiogram. The annotations include ideas of the UMLS model 2011AA belonging to the “Disorder” semantic group and which are linked to SNOMED-CT18. To guage our entity linking strategy, we solely retained annotations linking textual content spans to UMLS identifiers, excluding “CUI-less” annotations, that are annotations with out distinctive idea identifier (CUI), that make up round 30% of th dataset64,65.
We evaluated the entity linking efficiency of our pipeline by measuring the recall after candidate identification and the recall, precision and F1 rating after the candidate choice.
An entity hyperlink is taken into account right provided that it matches the precise textual content portion and idea identifier as the bottom reality annotations. The entity linking was evaluated throughout excessive similarity thresholds starting from 0.9 to 0.99 to keep away from degrading the efficiency via linking textual content portion to semantically much less dissimilar ideas and preserve resource-intensive computations low. To guage the efficiency acquire in our pipeline with the abbreviation disambiguation step, we additionally evaluated the efficiency distinction in our strategy by disambiguating abbreviations within the scientific notes previous to the entity linking step.
DARIO dataset
To guage the entity linking efficiency on German scientific notes, we used affected person knowledge from our inner coronary heart failure cohort dataset (DARIO dataset). The DARIO dataset contains retrospective well being information from 2012 to 2016 for 846 sufferers from the Division of Inside Drugs I on the College Hospital Aachen, with 351 sufferers with no coronary heart failure (41.5%), 193 sufferers with HFrEF (22.8%), 175 sufferers with HFmrEF (20.7) and 127 sufferers with HFpEF (15%). The distribution of the DARIO cohort displays the real-world prevalence of coronary heart failure subtypes in specialised scientific settings. Our retrospective research was authorised by the Ethics Committee of the Medical College at RWTH Aachen College (EK 416 − 21). The affected person knowledge consists of structured EHR knowledge containing demographic data, diagnoses, circumstances, and laboratory values as obtained from blood exams or electrocardiography in a tabular format and unstructured knowledge as scientific free-text notes within the type of discharge summaries. The scientific notes cowl totally different medical features such because the analysis, cardiac historical past, household historical past, social historical past, affected person development, process and beneficial remedy of the affected person. Moreover, the scientific word might include a report sheet with examination findings and different feedback. Sufferers could be recognized with coronary heart failure (HF) and differentiated into three subtypes based mostly on the ejection fraction (EF): lowered (HFrEF), mildly lowered (HFmrEF) or preserved (HFpEF).
For the reason that DARIO dataset lacks idea hyperlinks to SNOMED-CT or UMLS, we can not mechanically consider linked ideas in opposition to floor reality annotations. As an alternative, we evaluated the goodness of the linked entities by our entity linking strategy from the German scientific notes of the DARIO dataset via an skilled survey performed with three cardiologists from the Division of Inside Drugs I of the College Hospital Aachen. The survey questions consisted of 120 randomly sampled entity hyperlinks of the SNOMED-CT classes “Body structure”, “Procedures” and “Clinical finding”. Every query contained a textual content snippet, during which a portion of the textual content akin to a linked SNOMED-CT idea was highlighted and the SNOMED-CT idea was listed with its corresponding identifier. The physicians have been requested to fee randomly-sampled linked entities on a 3-Level LIKERT scale from “no match”, “partial match” and “exact match”. Solely SNOMED-CT ideas of the three classes “Procedures”, “Body Structures” and “Clinical Findings” have been evaluated to make sure feasibility of the skilled analysis. Linked entities are sampled in a stratified method based mostly on coronary heart failure classification and the corresponding SNOMED-CT hierarchy. In complete, 120 entity hyperlinks are sampled, with 30 entity hyperlinks per coronary heart failure class, together with 10 entity hyperlinks for every of the three SNOMED-CT hierarchies. The survey consists of three questionnaires, every containing 40 entity hyperlinks of the identical SNOMED-CT idea class throughout all 4 coronary heart failure affected person lessons. The inter-annotator settlement (IAA) of the physicians is evaluated utilizing the Cohen’s Kappa rating48, with values starting from − 1 to 1 to point probability settlement with low values as much as sturdy settlement with excessive values.
Coronary heart failure analysis
For coronary heart failure prediction our novel pipeline makes use of a Assist Vector Machine (SVM) classifier, which was skilled with affected person knowledge of the DARIO dataset. We targeted on diagnostic help of coronary heart failure of hospital sufferers, to cut back frequent misdiagnoses of sufferers and consider our novel complete pipeline to serve in future use circumstances with affected person knowledge of early illness phases, corresponding to scientific notes of common practitioners.
The classifier is skilled with affected person knowledge in type of EHR and obtained SNOMED-CT ideas from the beforehand computed entity hyperlinks to compute coronary heart failure predictions for every affected person (Fig. 9).
Within the classification step of our pipeline, knowledge of sufferers from the DARIO undertaking cohort in type of scientific notes and EHR knowledge is processed individually. Info in scientific notes is linked to SNOMED-CT ideas, for which all related ancestor ideas are retrieved and assigned a weight based mostly on the time period frequency-inverted doc frequency (TF-IDF) weighting scheme. We use the scikit-learn TF-IDF transformer match on the coaching dataset to assign weights to every SNOMED-CT idea based mostly on its frequency in a scientific word in relation to the frequency throughout the scientific notes of all sufferers, much like prior analysis assigning TF-IDF weights to key phrases of radiological reviews66. The EHR knowledge underwent knowledge cleansing to handle lacking values and inconsistencies, adopted by normalization to assemble every affected person’s EHR vector. The ensuing SNOMED-CT vector and EHR vector are mixed to symbolize every affected person with a complete vector, which serves as enter for the SVM classifier for coronary heart failure prediction. For the reason that TF-IDF transformer is match on the coaching knowledge solely to stop knowledge leakage, the SNOMED-CT vector is sparse when encoding SNOMED-CT ideas per affected person within the check dataset. We selected an SVM classifier on account of excessive efficiency when going through the sparsity and excessive dimensionality of the SNOMED-CT vector. The SVM classifier is then skilled with affected person vectors to foretell a coronary heart failure threat for every of the 4 coronary heart failure teams of sufferers having no coronary heart failure (None), coronary heart failure (HF) with lowered ejection fraction (HFrEF), mildly lowered ejection fraction (HFmrEF) or preserved ejection fraction (HFpEF). For the SVM implementation we used the scikit-learn SGDClassifier and modified the default parameters with the ‘hinge’ loss to coach a SVM with a hard and fast random_state of ‘2026’ and the class_weight set as ‘balanced’ to account for the dataset imbalance.
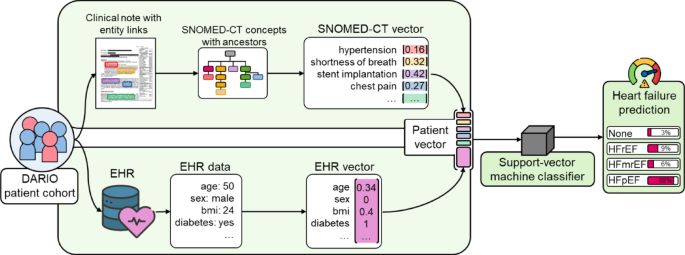
Overview of the classification strategy. The classification step leverages affected person knowledge within the type of scientific notes with entity hyperlinks and digital well being information (EHR). Ideas from entity hyperlinks, corresponding to hypertension or chest ache, alongside corresponding SNOMED-CT ancestor ideas make up a weighted SNOMED-CT vector for every affected person. Every affected person’s SNOMED-CT vector is mixed and standardized with a vector containing their EHR knowledge, corresponding to their age or BMI, to type a affected person vector. Affected person vectors are used for coaching and testing of the Assist-vector machine classifier, which predicts a coronary heart failure analysis for every affected person – on this case with an 82% likelihood of the affected person affected by HFpEF.
To evaluate the extent to which coronary heart failure classification could be improved by utilizing knowledge extracted from scientific notes, we in contrast the efficiency of three classifiers: one skilled solely on entity hyperlink knowledge, one other on tabular knowledge within the type of EHR knowledge, and a classifier skilled on each forms of knowledge.
All classifications came about with stratified 5-fold cross-validation throughout all 4 coronary heart failure lessons and with a hard and fast random seed ‘2026’ for reproducibility.
Furthermore, to validate the efficiency acquire of our carried out pipeline, we in contrast the efficiency to a fine-tuned medBERT.de mannequin for the classification job, which instantly produces a coronary heart failure prediction based mostly on scientific notes with out requiring entity linking to SNOMED-CT ideas. medBERT.de is a novel complete German BERT mannequin skilled on 4.7 million various medical paperwork and achieved state-of-the-art efficiency on a number of medical benchmarks. medBERT.de poses a neural baseline with two classification approaches: first, the classification is carried out solely on scientific notes and second, via late-fusion scientific notes and EHR are mixed for a complete analysis. For the late-fusion strategy, we carried out a multimodal neural structure extending the medBERT.de classification strategy. The textual content knowledge is processed by way of the medBERT.de transformer to extract a semantic embedding classify token (CLS token), whereas the structured EHR knowledge is processed via a parallel Multi-Layer Perceptron (MLP) department consisting of two linear layers with Rectified Linear Unit (ReLU) activation and dropout. These two representations are then concatenated to type a joint function vector, which serves because the enter to a ultimate classification head. This structure permits to concurrently optimize the textual content embeddings and the tabular function weights through the coaching course of to determine cross-modal correlations. The medBERT.de text-based and multimodal strategy with the MLP have been configured with a studying fee of 2e-5, analysis technique set to ‘epoch’ with a most of 10 epochs, early stopping persistence set to three and early stopping threshold of 0.001 to permit as much as three consecutive coaching iterations with out efficiency enchancment earlier than stopping the coaching course of. The optimization came about in direction of the ‘f1’ rating with a hard and fast random seed set to ‘2026’.