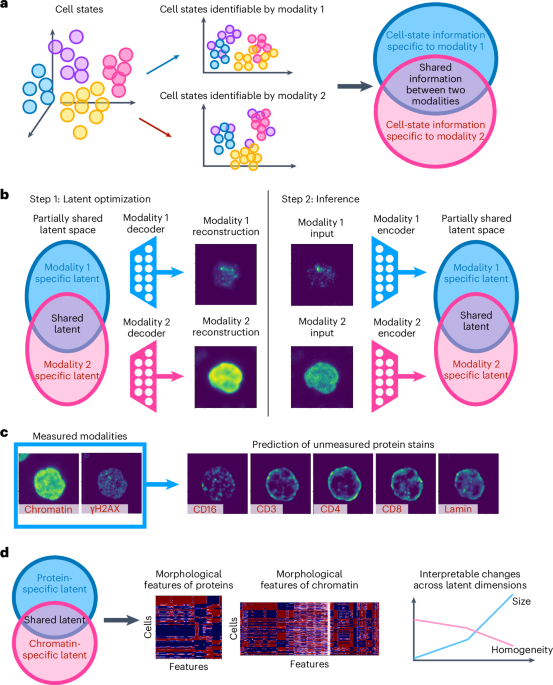
Latent optimization
In apply, much like variational sampling in autoencoders, to enhance generalization to unseen samples, we add Gaussian noise to every element of the latent area and add a regularization time period (ℓ2-norm of the latent options) along with the reconstruction losses. Thus, the complete goal perform is:
$$start{array}{l}{{bf{E}}}_{({x}^{(1)},{x}^{(2)}) sim P({X}^{(1)},{X}^{(2)})}[L({x}^{(1)},{D}_{1}({z}_{S}+{epsilon }_{S},{z}_{{S}_{1}}+{epsilon }_{{S}_{1}})) +L({x}^{(2)},{D}_{2}({z}_{S}+{epsilon }_{S},{z}_{{S}_{2}}+{epsilon }_{{S}_{2}}))+L({x}^{(1)},{D}_{1}^{{prime} }({z}_{S}+{epsilon }_{S})) +L({x}^{(2)},{D}_{2}^{{prime} }({z}_{S}+{epsilon }_{S}))]+lambda (parallel {z}_{S}{parallel }_{2}+parallel {z}_{{S}_{1}}{parallel }_{2}+parallel {z}_{{S}_{2}}{parallel }_{2}),finish{array}$$
(1)
the place ϵS, ({epsilon }_{{S}_{1}}) and ({epsilon }_{{S}_{2}}) denote Gaussian noise in every element of the latent area and λ is a hyperparameter. We reduce equation (1) to acquire the decoders D1, D2, ({D}_{1}^{{prime} }) and ({D}_{2}^{{prime} }) in addition to the shared latent options zS and the modality-specific latent options ({z}_{{S}_{1}}) and ({z}_{{S}_{2}}).
Mannequin structure and coaching of paired scRNA-seq and scATAC-seq
For each coaching steps, we cut up the information into coaching and validation units, as is normal in neural community coaching. We use the identical randomly chosen 85% of cells from the SHARE-seq dataset5 to coach our mannequin and the remaining 15% to validate and take a look at the generalization efficiency of our mannequin.
Step 1 latent optimization
On this step, we practice the latent areas and the corresponding decoders to reconstruct the scRNA-seq and scATAC-seq knowledge (Prolonged Information Fig. 5b and Supplementary Fig. 1a). The coaching was carried out on one 24 GB GPU for 35 hours with 10.8 seconds per epoch.
Latent areas of paired scRNA-seq and scATAC-seq
The shared latent area has 50 dimensions and every of the 2 modality-specific latent areas have 20 dimensions. The shared latent area is chosen to be a lot bigger than the modality-specific latent areas to make sure that the shared area has sufficient capability to include all shared info. Latent areas are initialized utilizing ‘torch.nn.Embedding’ with the default parameters. Unbiased Gaussian noise with zero imply and unit variance is added to the latent areas at every coaching epoch. The hyperparameter λ in equation (1) for the ℓ2 regularization of the latent areas is ready to 0.001 and the training charge of the ADAM optimizer is ready to 0.001, that are the identical hyperparameter values as in ref. 37.
Decoders of paired-sequencing-based modalities
4 decoders are educated in step 1: (1) a decoder that reconstructs scRNA-seq from the shared latent area; (2) a decoder that reconstructs scRNA-seq from the complete RNA-seq latent area (the shared latent area embedding concatenated with the RNA-specific latent area embedding); (3) a decoder that reconstructs scATAC-seq from the shared latent area; and (4) a decoder that reconstructs scATAC-seq from the complete ATAC-seq latent area (the shared latent area embedding concatenated with the ATAC-specific latent area embedding). The architectures of the decoders are proven in Prolonged Information Fig. 5b. The enter characteristic dimension is 70 for the complete latent area decoder and 50 for the shared latent area decoder besides when completely different latent area sizes are examined (Fig. 2a). Every decoder has 3 hidden layers, every of which has dimension 1,024. All hidden layers are linear layers with a dropout charge of 0.01 and are adopted by LeakyReLU activation and a batch normalization layer. The output layer is a linear layer with a dropout charge of 0.01 and sigmoid activation. This decoder structure follows normal autoencoder set-ups23,58. For each modalities, we use binary cross-entropy loss to reduce the target perform outlined in equation (1), which is calculated utilizing ‘torch.nn.BCEWithLogitsLoss’ based mostly on the output earlier than the sigmoid activation with pos_weight set inversely proportional to the entire counts of genes or peaks. The educational charge of the decoders is 0.0001 and ADAM is used for optimization, which is identical as in ref. 37.
Step 2 inference
On this step, we practice the encoders to deduce the realized latent areas from the scRNA-seq and scATAC-seq knowledge with out updating the latent areas or the decoders (Prolonged Information Fig. 5c and Supplementary Fig. 1b). The coaching was carried out on one 24 GB GPU for 9 hours with 7.5 seconds per epoch to coach the mannequin previous convergence.
Encoders of paired scRNA-seq and scATAC-seq
We practice two separate encoders for scRNA-seq and scATAC-seq with an identical buildings (Prolonged Information Fig. 5c). Every encoder begins with two linear layers with a dropout charge of 0.01, every of which has 1,024 dimensions and is adopted by LeakyReLU activation and batch normalization, which is an ordinary set-up for autoencoders23,58. After the primary two hidden layers, two separate linear layers with LeakyReLU activation are used to acquire separate hidden layers for the shared and modality-specific latent areas. A linear layer is utilized to every hidden layer to acquire the shared or modality-specific latent area. The inferred latent areas from the autoencoder are in contrast with the latent areas realized in step 1 by way of MSE loss. The MSE loss is minimized utilizing the ADAM optimizer with a studying charge of 0.0001, the identical studying charge because the decoder.
Utility to paired scRNA-seq and protein abundance
The identical set-up is utilized to the paired scRNA-seq and protein abundance knowledge measured by CITE-seq25, with dimension 50 for the shared latent area and dimension 30 for the modality-specific latent area. On condition that cross-modality prediction isn’t the purpose on this utility and to exhibit robustness of our method, the mannequin is educated with out the 2 decoders ({D}_{1}^{{prime} }) and ({D}_{2}^{{prime} }) that map from the shared latent area to every of the modalities (Prolonged Information Fig. 6a).
Pre-processing of scRNA-seq and scATAC-seq knowledge
The paired scRNA-seq and scATAC-seq datasets from ref. 5 are individually filtered for genes or peaks which are non-zero in at the very least 300 cells after which filtered for cells with at the very least 300 non-zero genes or peaks. The frequent cells between the 2 modalities are then chosen for one more spherical of filtering with the identical standards. These two rounds of filtering lead to 28,098 cells with 9,153 genes within the scRNA-seq knowledge, and 58,170 peaks within the scATAC-seq knowledge. The information are then log-transformed and min–max scaled, such that the minimal depend in every cell is 0 and the utmost depend in every cell is 1. Our filtering and normalization steps observe normal procedures of preprocessing scRNA-seq and scATAC-seq knowledge (see, for instance, refs. 5,22,23,42).
Preprocessing of scRNA-seq and mobile floor protein abundance knowledge
Following the method taken by a earlier examine that analyzes this dataset25, we use the highest 4,005 extremely variable genes within the scRNA-seq knowledge and all 110 proteins. Identical as within the preprocessing of scRNA-seq and scATAC-seq described above, the information are log-transformed and min–max scaled, such that the minimal depend in every cell is 0 and the utmost depend in every cell is 1.
Different fashions for scRNA-seq and mobile floor protein abundance knowledge
We use the default parameter setting for the Seurat WNN technique17. For testing the usual multi-modal autoencoder technique, we use the identical encoder and decoder architectures as the complete APOLLO mannequin with 80 latent dimensions, which is the same as the complete latent area dimension of the APOLLO mannequin. As an alternative of performing a two-step coaching, this mannequin trains the encoder and decoder collectively in a single step, with out immediately updating the latent areas as parameters for optimization. MSE loss is added between the latent areas obtained from the 2 encoders of the 2 enter modalities. Much like the APOLLO mannequin that provides Gaussian noise within the latent area, this mannequin performs a variational sampling step as in an ordinary variational autoencoder and the ensuing latent areas are handed to the decoder for reconstruction. In every epoch, we alternate between coaching the scRNA-seq autoencoder and coaching the protein autoencoder.
Cell-type prediction based mostly on the inferred latent areas of scRNA-seq and scATAC-seq
We practice 4 separate neural community classifiers with the identical structure to foretell cell varieties based mostly on one of many following inputs: (1) shared latent area inferred from scRNA-seq knowledge utilizing the RNA encoder; (2) shared latent area inferred from scATAC-seq utilizing the ATAC encoder; (3) each shared and modality-specific latent areas inferred from scRNA-seq knowledge utilizing the RNA encoder; and (4) each shared and modality-specific latent areas inferred from scATAC-seq knowledge utilizing the ATAC encoder. We use an ordinary feedforward neural community structure in our classifiers; see, for instance, ref. 19. Every classifier consists of 4 layers and outputs the chance of a given cell being assigned to every of the 23 cell varieties. The primary three layers are adopted by LeakyReLU activation and batch normalization. All 4 layers have a dropout charge of 0.1 and a hidden dimension of 128. We use the ADAM optimizer with a studying charge of 0.00001 to reduce the cross-entropy loss between our prediction and the cell-type labels assigned by ref. 5. We use the identical practice–validation cut up to coach the classifiers as within the coaching of the APOLLO mannequin, which suggests the identical 85% of cells are used to coach the APOLLO mannequin and the cell-type classifiers.
Mannequin structure and coaching of paired chromatin and protein imaging
For each coaching steps, we hold-out all photographs from one affected person in every of the 4 phenotypes for testing the mannequin (Supplementary Fig. 4).
Step 1 latent optimization
On this step, we practice the latent areas, protein IDs, and the corresponding decoders to reconstruct the chromatin and protein photographs (Prolonged Information Fig. 8a and Supplementary Fig. 5a). The coaching was carried out on one 24 GB GPU with 40 seconds per epoch.
Latent areas and protein IDs of paired chromatin and protein imaging
For every protein, we randomly initialize a trainable 64-dimensional vector of protein ID that’s shared throughout all photographs of that protein by utilizing ‘torch.nn.Embedding’. The protein IDs are utilized in each the encoding and decoding steps by concatenating them to the latent area embeddings, much like a conditional autoencoder mannequin (Prolonged Information Fig. 8). The shared latent area has 1,024 dimensions and every of the 2modality-specific latent areas has 200 dimensions besides when completely different latent area sizes are examined (Fig. 3b). Much like the appliance to paired scRNA-seq and scATAC-seq, the shared latent area is chosen to be a lot bigger than the modality-specific latent areas to make sure that the shared area has sufficient capability to include all of the shared info. Latent areas are initialized utilizing ‘torch.nn.Embedding’ with the default parameters. Unbiased Gaussian noise with zero imply and unit variance is added to the latent areas at every coaching epoch. The hyperparameter λ in equation (1) for the ℓ2 regularization of the latent areas is ready to 0.001 and the training charge of the ADAM optimizer is ready to 0.001, that are the identical hyperparameter values as in ref. 37.
Decoders of paired chromatin and protein imaging
4 decoders are educated in step 1: (1) a decoder that reconstructs chromatin photographs from the shared latent area; (2) a decoder that reconstructs chromatin photographs from the complete chromatin latent area (the shared latent area embedding concatenated with the chromatin-specific latent area embedding); (3) a decoder that reconstructs protein photographs from the shared latent area; and (4) a decoder that reconstructs protein photographs from the complete protein latent area (the shared latent area embedding concatenated with the protein-specific latent area embedding). The architectures of the decoders are proven in Prolonged Information Fig. 8a. The latent area embedding, after including noise and being concatenated with the protein ID, is handed by way of a linear layer with ReLU activation and reshaped to 4 × 4 × 96 dimensions for subsequent convolutions. That is adopted by 5 convolutional layers with a kernel measurement of 4 and stride of two. The variety of channels in every hidden layer is listed in Prolonged Information Fig. 8a. The primary 4 convolutional layers are adopted by batch normalization and LeakyReLU activation. The final convolutional layer is adopted by sigmoid activation to scale the output picture from 0 to 1. For each imaging modalities, we use binary cross-entropy loss to reduce the target perform outlined in equation (1), which is calculated utilizing ‘torch.nn.BCEWithLogitsLoss’ based mostly on the output earlier than the sigmoid activation. The educational charge of the decoders is 0.0001 and ADAM is used for optimization. The decoder architectures and coaching process observe normal set-ups for autoencoders (see, for instance, refs. 23,37,45).
Step 2 inference
On this step, we practice the encoders to deduce the realized latent areas from the chromatin and protein photographs with out updating the latent areas, protein IDs or the decoders (Prolonged Information Fig. 8b and Supplementary Fig. 5b). The coaching was carried out on one 24 GB GPU with 20 seconds per epoch.
Encoders of paired chromatin and protein imaging
We practice two separate encoders for chromatin and protein photographs with an identical buildings (Prolonged Information Fig. 8b). Every encoder begins with 5 convolutional layers with LeakyReLU activation, an ordinary set-up for autoencoders23,37,45. The scale of the hidden layers are listed in Prolonged Information Fig. 8b. The output of the final convolutional layer is split into two units of channels which are used to derive the shared and modality-specific latent areas respectively. Eighty out of the 96 channels of the final hidden layer are flattened, concatenated with protein ID, and handed by way of a linear layer to acquire the shared latent area. Equally, the modality-specific latent area is obtained from the remaining 16 channels. The inferred latent areas from the autoencoder are in contrast with the latent areas realized in step 1 by way of MSE loss. The MSE loss is minimized utilizing the ADAM optimizer with a studying charge of 0.001.
Cross-modality predictions
To foretell unmeasured proteins, the shared latent area is first inferred from the chromatin picture of a cell utilizing the chromatin encoder. Then every protein picture will be predicted by decoding the inferred shared latent area by way of the protein decoder utilizing the protein ID of the goal protein (Prolonged Information Fig. 8a).
Utility to HPA knowledge
The identical set-up is utilized to the HPA knowledge35, with a shared latent area dimension of 1,024 and a modality-specific latent area dimension of 200. The 2 decoders ({D}_{1}^{{prime} }) and ({D}_{2}^{{prime} }) that map from the shared latent area to every of the modalities are usually not used, because the omission doesn’t impression cross-modality-prediction efficiency (Fig. 3b) and ends in right disentanglement of the shared and modality-specific info (Fig. 2f).
Different fashions
The fashions used for benchmarking our full APOLLO mannequin are described under. Along with utilizing binary cross entropy (BCE) loss for the decoder outputs, we additionally examined the usage of another loss perform, specifically, the MSE loss. All mannequin coaching and testing use the identical practice–take a look at cut up. For benchmarking cross-modality prediction, the prediction outcomes are first thresholded utilizing mode depth after which in contrast utilizing ℓ1 loss, which isn’t used for coaching any of the fashions.
Commonplace autoencoder coaching utilizing a single step
This mannequin has the identical encoder and decoder architectures as the complete APOLLO mannequin, and the identical dimensions of the shared, chromatin-specific and protein-specific latent areas are used. Protein IDs are concatenated to the hidden layers throughout encoding and decoding as described within the full mannequin. Nonetheless, as a substitute of performing a two-step coaching, this mannequin trains the encoder and decoder collectively in a single step, with out immediately updating the latent areas as parameters for optimization (Prolonged Information Fig. 10a). MSE loss is added between the shared latent areas obtained from the 2 encoders of the 2 enter modalities. Much like the APOLLO mannequin that provides Gaussian noise within the latent area, this mannequin performs a variational sampling step as in an ordinary variational autoencoder and the ensuing latent areas are handed to the decoder for reconstruction. In every epoch, we alternate between coaching the chromatin autoencoder and coaching the protein autoencoder.
Our mannequin with out modality-specific latent area
This mannequin has the identical two-step coaching process as the complete APOLLO mannequin, however doesn’t separate the shared and the modality-specific latent areas (Prolonged Information Fig. 10b). There are solely two decoders that decode the 2 modalities from the shared latent area and they’re educated in step 1. The encoders solely output the shared latent area, which has the identical dimension because the mixed dimension of the shared and the modality-specific latent areas within the full APOLLO mannequin.
Inpainting mannequin
For the inpainting mannequin developed in ref. 45, we tailored the unique code from TensorFlow to PyTorch. We used the identical hyperparameter values as within the unique publication, together with the usage of MSE loss, besides the slight modification to adapt to the single-channel enter picture and not using a microtubule channel.
Information simulation and evaluation of APOLLO’s disentanglement efficiency
To systematically consider APOLLO’s means to disentangle shared and modality-specific representations, we generate simulated multi-modal datasets with ground-truth latent construction. Given the current theoretical outcomes on identifiability of the shared and modality-specific variables36, we solely allowed directed edges from the shared latent variables to the modality-specific latent variables however not vice versa. For all simulations, we assign the latent variables Z1 and Z2 to the shared latent area, Z3 and Z4 to the modality 1 particular latent area, and Z5 to the modality 2 particular latent area. The latent causal graph and the corresponding chance distribution are laid out in panel a of Prolonged Information Figs. 1–4, from which 2,000 samples of latent options are independently drawn for every simulation and visualized in panel b of Prolonged Information Figs. 1–4. Given the causal graphs, we count on an correct disentanglement to establish that the modality 1 particular latent area additionally captures Z2 in simulations 3–5 however not in simulations 1 and a couple of. Every noticed characteristic is a linear mixture of a number of latent options with the coefficients independently drawn from an ordinary regular distribution. In simulations 1 and three, the place every noticed characteristic is determined by simply 1 latent characteristic, 10 noticed options per modality are generated from every latent characteristic, leading to a characteristic dimension of 40 in modality 1 and 30 in modality 2. In simulations 2, 4 and 5, the place we embrace noticed options with a number of dad and mom, 10 noticed options in modality 1 are generated from: (1) every considered one of Z2, Z3 and Z4 (30 noticed options whole); (2) every pair of Z2, Z3, and Z4 (30 noticed options whole); (3) Z2, Z3, and Z4; and (4) all 4 modality 1 latent options. Equally, a complete of 40 noticed options in modality 2 are generated from: (1) every considered one of Z2 and Z5 (20 noticed options whole); (2) Z2 and Z5 (10 noticed options); and (3) all three modality 2 latent options. This ends in a characteristic dimension of 40 in modality 1 and 30 in modality 2. In simulation 5, we examined the consequences of upper weights for the noticed options that rely solely on one latent characteristic by utilizing the identical set-up as simulation 4 whereas drawing coefficients of the noticed options from ({mathcal{N}}(0,16)). For APOLLO coaching, we use absolutely related decoders with the same structure as within the paired-sequencing functions and an MSE loss. Every of the shared and modality-specific latent areas has two dimensions. Accuracy in disentanglement is assessed by the separation of the binary variables Z1 and Z3 within the three latent areas, which we quantify utilizing silhouette scores59 and UMAP visualizations (Prolonged Information Figs. 1–4).
Pre-processing of paired chromatin and protein imaging
The imaging knowledge are obtained from ref. 43, the place the sufferers are annotated with one of many following 4 phenotypic courses: wholesome, meningioma, glioma, or head and neck tumor (Supplementary Fig. 4). Every single-cell picture of every protein stain and chromatin stain is min–max scaled such that the minimal pixel worth is 0 and the utmost is 1, an ordinary picture normalization step for neural networks43,45. For every cell nucleus, a picture patch centered on the centroid of the nucleus of dimension 128 × 128 pixels is cropped from the entire picture. A complete of 29,174 cell photographs have been utilized in coaching.
Pre-processing of the HPA photographs
The identical process as in a earlier examine32 is used to pre-process the pictures. We use all photographs of U2OS cells (the cell line with essentially the most knowledge within the HPA) which are additionally utilized in ref. 32 for finding out protein localization variabiltiy (2,973 proteins used) and exclude proteins which are stained in lower than 150 cells, leading to a complete of 141 proteins used for coaching the mannequin. The subset of proteins has been proven in ref. 32 to have various subcellular localizations. Nuclear segmentation obtained utilizing StarDist60 is used for computing the intranuclear proportion of the protein in every picture. The usual deviation of the intranuclear proportion of every protein is computed throughout all photographs stained for the protein. The 25 proteins with the most important normal deviations are used for deciphering the disentanglement outcomes (Fig. 5).
Phenotype classification utilizing actual or reconstructed protein and chromatin photographs
We use the ResNet-18 mannequin61, a preferred neural community mannequin for picture classification, in PyTorch for the phenotype classifiers to foretell phenotypes from actual protein/chromatin photographs, reconstructed protein/chromatin photographs, or protein photographs predicted from chromatin (Fig. 3d). The primary layer of the ResNet-18 mannequin is adjusted to take single-channel enter photographs. We use the cross-entropy loss and set the burden of every phenotype class to be inversely proportional to the fraction of cells in that individual phenotype class. The classifiers are educated utilizing the ADAM optimizer with a studying charge of 0.001. The coaching of every classifier is repeated 36 occasions with 6 completely different teams of held-out sufferers, the place every held-out group accommodates one affected person from every phenotype class. One-sided two-sample t-test and Wilcoxon signed-rank take a look at are used for the null speculation that the phenotype prediction accuracy utilizing the reconstruction from the complete latent area is smaller than or equal to the accuracy utilizing the reconstruction from the shared latent area.
Interpretation of the partially shared latent areas
Within the following, we describe a process to investigate the shared latent area in addition to the modality-specific latent area, which will be utilized to any knowledge modality. We compute the PCs of the latent areas and group the cells based mostly on their positions alongside the PCs. For each of our functions, we group cells into 11 bins with equal percentile ranges alongside every PC. Within the following description of our method, we denote the percentile bin of PCi that’s centered round 0 as ({mathrm{PC}}_{i}^{0}) and the bins ordered from detrimental to optimistic PC values are denoted as ({mathrm{PC}}_{i}^{-5},cdots {mathrm{PC}}_{i}^{-1},{mathrm{PC}}_{i}^{0},{mathrm{PC}}_{i}^{1},cdots {mathrm{PC}}_{i}^{5}). When evaluating cells alongside PCi we solely use the cells on the heart of all different PCs, that means the 15% of cells with the smallest Euclidean distance to the PC origin calculated utilizing all PCj for j ≠ i and j < 10. The variety of PCs thought of within the evaluation will be adjusted given the variance defined by every PC in a specific dataset. Utilizing cells on the heart of all different PCs, besides the PC being interpreted, ensures that we’re solely contemplating the variation of cells alongside one PC to tease aside the variations defined by the completely different PCs. Within the following, we clarify this process in additional element within the context of the paired-sequencing and paired-imaging datasets, the place we additionally carry out subsampling for a extra strong identification of the options defined by every PC.
Interpretation of the latent areas of paired scRNA-seq and scATAC-seq
We take a look at for vital gene expression and peak depend modifications alongside every PC. The cells are divided into 36 random practice–take a look at splits with 20% of cells used for testing in every cut up, to validate the numerous genes and peaks recognized alongside every PC. We take into account three teams of cells for every PCi which are all on the heart of different PCs: (1) cells in ({mathrm{PC}}_{i}^{-5}) and ({mathrm{PC}}_{i}^{-4}); (2) cells in ({mathrm{PC}}_{i}^{0}); and (3) cells in ({mathrm{PC}}_{i}^{4}) and ({mathrm{PC}}_{i}^{5}). Cells in every group are in contrast with all cells within the different two teams by way of t-tests utilizing Scanpy’s ‘rank_genes_groups’ perform42. The edge of P values after a number of testing correction is ready to 0.05 for each modalities. The variety of occasions in all practice–take a look at splits a gene or peak is examined to be vital in each the coaching and testing cells with the identical route of fold change are plotted for various thresholds of log fold change magnitude (Fig. 2c and Supplementary Figs. 2 and three). The ATAC-seq peaks within the shared and modality-specific latent areas are summarized in Fig. 2nd. For every ATAC-seq peak, we counted the variety of PCs within the shared and modality-specific latent areas respectively, by which the height is discovered to be vital. The genes are then grouped by their GO annotations and the entire counts of PCs within the two latent areas are normalized to 1.
Interpretation of the latent areas of paired chromatin and protein imaging
Imaging knowledge will be visually inspected by sampling cells at every percentile bin alongside a specific PC (Fig. 4a). As well as, for a extra complete evaluation, predefined handcrafted picture options43,62 can be utilized to check for vital characteristic modifications alongside every PC. For the next evaluation, we use a complete of 234 chromatin picture options and round 30 picture options per protein obtained from43. The sufferers are divided into 12 completely different practice–take a look at splits, that are summarized in Supplementary Fig. 4, to validate the numerous options recognized alongside every PC. As within the utility to paired scRNA-seq and scATAC-seq, we take a look at for vital characteristic modifications amongst three teams of cells for every PCi which are all on the heart of different PCs: (1) cells in ({mathrm{PC}}_{i}^{-5}) and ({mathrm{PC}}_{i}^{-4}); (2) cells in ({mathrm{PC}}_{i}^{0}); and (3) cells in ({mathrm{PC}}_{i}^{4}) and ({mathrm{PC}}_{i}^{5}). A characteristic is taken into account vital if the P worth after a number of testing correction is lower than 0.05 and absolutely the worth of log fold change is larger than log2(FCt) in each the coaching and testing sufferers, the place FCt is ready to be 1.2 for chromatin options and 1.4 for protein options. The fold-change thresholds are chosen so that the majority consultant morphological options are represented by at the very least one PC. As well as, the route of fold change is required to be the identical within the coaching and testing sufferers. Determine 4a and Supplementary 19 present the chromatin options which are vital in at the very least 8 out of the 12 practice–take a look at splits (Supplementary Fig. 4). We summarize the morphological options in all prime PCs within the shared and modality-specific latent areas respectively in Fig. 4b, and Supplementary Figs. 9c, 10c, 11c and 14c. For this, we counted the variety of PCs for which every characteristic is discovered to be vital within the shared and modality-specific latent areas and normalized the entire counts within the two latent areas to 1.
To acquire a extra concise illustration, we group the morphological options of chromatin and every protein by Pearson correlation throughout all cells and plot one consultant characteristic per group in Fig. 4a,b, and Supplementary Figs. 8–14. The characteristic groupings for every stain are obtained as follows: we construct a community with the options as nodes and an edge is added between every pair of options whose Pearson correlation is larger than 0.7. Options throughout the similar related element of the graph are thought of in the identical group. The consultant characteristic of a bunch is the node with the very best diploma. This ends in 18 characteristic teams for chromatin, 5 teams for γH2AX, 5 teams for lamin, 6 teams for CD8, 4 teams for CD4, 5 teams for CD3 and 4 teams for CD16.
Interpretation of the latent areas of the HPA photographs
For every subset of U2OS cells stained with the identical protein, we individually carry out ok-means (ok = 2) clustering within the shared and the 2 modality-specific latent areas of every mannequin. The clustering in every latent area is repeated 5 occasions with completely different random seeds. For every random seed, a t-test is carried out to check the proportion of intranuclear protein localization within the two clusters, and absolutely the worth of the distinction between the cluster means is computed. A clustering reveals vital distinction of intranuclear proportions if P < 0.00022 and absolutely the distinction between clusters >0.01 in all 5 random initializations. The P-value threshold is chosen by making use of a Bonferroni correction to the general chance of kind 1 error at 0.05 for a complete of 225 checks carried out for 25 proteins in every of the three latent areas of the three separate fashions.
γH2AX characteristic prediction utilizing chromatin photographs
We use the ResNet-18 mannequin61 in PyTorch to coach separate regression fashions that predict the predefined γH2AX options from chromatin photographs. Much like the earlier part, for every of the three γH2AX options (Fig. 4b), a regression mannequin is educated 12 occasions with completely different practice–take a look at splits of the sufferers (Supplementary Fig. 4). The outputs of every regression mannequin on the held-out sufferers are then in contrast with the bottom fact utilizing Pearson’s correlation (Supplementary Fig. 13c).
Statistics and reproducibility
Publicly out there datasets have been used on this examine. Commonplace filtering steps have been carried out, and in any other case, no knowledge have been excluded from our analyses. Randomization and blinding weren’t relevant on this examine.
Reporting abstract
Additional info on analysis design is obtainable within the Nature Portfolio Reporting Abstract linked to this text.