Used microscopy datasets
Here, we describe the ten microscopy datasets we have used in this work, eight of which have been generated de novo by the authors; two others were taken from a previous publication3. All datasets will be publicly available, enabling full reproducibility of all experiments and results we have presented, and allowing others to directly compare their own methodological improvements to the approach we have presented. The datasets cover a broad spectrum of noise levels (SNR), number of imaged fluorescent channels, overall intensity and relative intensity of these channels, imaging modality (for example confocal, spinning disk confocal, structured illumination, live, fixed, and expanded samples, 2D and 3D). Using these datasets, we have trained \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) models for a total of 36 tasks in total, covering two-, three- and four-channel semantic unmixing experiments (as shown throughout all figures and summarized in Table 1 and Supplementary Table 2). All datasets are openly available ( Next, we describe the datasets. In Table 2, we provide a brief overview of the different datasets used in this work.
The HT-P23A and HT-P23B datasets
This dataset was acquired by the Pigino group at Human Technopole, Milan, Italy. The data generation procedure was based on previous work23.
Cryo-ExM of MDCK-II cells
MDCK-II cells were seeded on 12-mm coverslips contained within six-well plates at a density of 300,000 cells per well, grown at 37 °C and 5% CO2. When cells reached approximately 50% confluence, a subset of coverslips was incubated with 150 nM MitoTracker Deep Red FM (Thermo Fisher Scientific, M22426) for 30 min and checked for dye incorporation using an EVOS M5000 Imaging System (Thermo Fisher Scientific). Cells were then immediately processed for cryo-ExM1. In brief, cells on coverslips were rapidly plunged frozen in −180 °C liquid ethane using a manual plunger. Coverslips were transferred to an Eppendorf tube containing frozen, desiccated acetone supplemented with 0.1% paraformaldehyde (PFA) and 0.02% glutaraldehyde (GA) for overnight freeze substitution in dry ice. The next morning, the coverslips underwent successive ethanol baths in progressively decreasing percentages, as follows: ethanol 100% (5 min), ethanol 100% (5 min), ethanol 95% (5 min), ethanol 95% (5 min), ethanol 70% (3 min), ethanol 50% (3 min) and PBS. Substituted cells then underwent anchoring in 2% acrylamide and 1.4% formaldehyde in PBS for 3 h. Coverslips were then exposed to activated monomer solution containing sodium acrylate (19%), acrylamide (10%), N,N’-methylenbisacrylamide (0.1%) and ddH2O, activated by APS and TEMED (0.05%). Expansion gels polymerized for 1 h at 37 °C, and were then denatured in denaturation solution containing SDS (200 mM), NaCl (200 mM) and Tris (50 mM) in ultrapure water at 95 °C for 1.5 h. Expanded gels were then washed in PBS 3×, 10 min for each wash. Final concentrations of solutions are reported.
Expansion immunolabeling
The following day, gels were stained with anti-α-tubulin (ABCD Antibodies, ABCD_AA345) and anti-β-tubulin antibodies (ABCD antibodies, AA_344), diluted 1:300 in PBS 2% BSA. Half of the MitoTracker labeled gels were given Mouse IgG2a raised antibody, and the rest of the gels were stained with guinea pig IgG raised antibody. Tubulin-labeled gels were incubated at 37 °C for 3 h while gently shaking. After primary labeling, gels were washed 3× in PBS 0.1% Tween, 10 min for each wash. Mouse IgG2a tubulin-labeled gels were then stained with goat anti-mouse IgG secondary antibody Alexa Fluor 488 (Thermo Fisher Scientific, A-11001), while guinea pig IgG tubulin-labeled antibodies were stained with goat anti-guinea pig IgG secondary antibody Alexa Fluor 647 (Thermo Fisher Scientific, A-21450). Secondary antibody labeled gels were then incubated at 37 °C for 3 h while gently shaking. The gels were washed 3× in PBS 0.1% Tween, 10 min for each wash. Gels were further washed 2×, 30 min in ddH2O, and then placed in ddH2O overnight for expansion.
In HT-P23A, the fluorescence intensity of labeled microtubules strongly overpowered the mitochondrial signal in the condition in which both microtubules and mitochondria were co-fluorescent in the 647 channel. To ensure a more equal signal, we imaged HT-P23B, where we reduced the concentration of tubulin guinea pig IgG labeling to 1:450, and the concentration of anti-guinea pig IgG secondary antibody to 1:450 to ensure a more equal signal.
Imaging expansion gels
Expanded gels were cut into approximately 1.5-cm2 pieces and placed on a 24-mm coverslip coated in poly-D-lysine (Sigma, P7886) and placed within a 35-mm imaging chamber. Imaging was performed on a Zeiss LSM980-NLO, in confocal mode. z-stacks with a step size of 0.15 μm were collected with a frame size of 2,024 × 2,024 pixels using a ×63 oil immersion objective (1.4 NA).
The Pavia-P23 dataset
This dataset was acquired at the Synthetic Physiology Laboratory at the University of Pavia, Italy.
HaCaT cell line culture and medium
The HaCaT cell line was kindly gifted by H.de Jonge (Department of Molecular Medicine, University of Pavia). Cells were maintained in DMEM/F12 no phenol red (Gibco, 21041-025) with 10% fetal bovine serum (FBS) (Gibco, 10270-106) and 1% penicillin–streptomycin (P–S) (HiMedia, A001). Cells were never allowed to go beyond 75–80% confluency during routine splitting.
Generation of the HaCaT FUCCIplex-prototype clonal cell line
The HaCaT cell line expressing the structural actin and tubulin fluorophores and the FUCCIplex sensor was generated as described previously24. In brief, cells were plated at the density of 6 × 104 cells per well on a 12-well plate and infected with the FUCCIplex lentivirus (10 μl of concentrated FUCCIplex lentivirus particles (>108 TU ml−1, VectorBuilder). Positive cells were then selected with 20 μl ml−1 of hygromycin B (50 mg ml−1 PBS) (Invitrogen, 10687010) and expanded. Positive cells expressing the FUCCIplex sensor were then infected with a genetically encoded Lifeact-ACTB-RFP probe (rLV Ubi-LifeAct-TagRFP, 1 × 107 TU ml−1, 10 μl) that specifically tags the (F)-actin filaments with a red fluorescent reporter. Last, the α-tubulin locus (TUBA1B, NM006082.3) was genome-edited via CRISPR/Cas9 using the Thermo Fisher Scientific True Tag system (A42992) to tag the tubulin N terminus of the protein with an EGFP fluorophore. Positive cells were clonally selected and expanded. The True Tag homology arm primer forward sequence is TCCTGTCGCCTTCGCCTCCTAATCCCTAGCCACTATGGGAGGTAAGCCCTTGCATTCG and the reverse is CCTGAAAGCAGCCGGGAGCCGCACGGCTTACTCACACCGCTTCCACTACCTGAACC. The TrueGuide synthetic gRNA (sgRNA) sequence is GCACGGCTTACTCACCATAG. For imaging experiments, HaCaT cells were plated in culture medium into porcine skin-coated (0.2%, w/v) ibidi μ-slide eight-well-chambered coverslips (Ibidi, 80807) at a density of 50,000 cells per well.
Imaging methods for Pavia-P24 dataset
Imaging was performed using a Nikon Ti2 Eclipse inverted microscope integrated with a Crest V3 X-Light spinning disk confocal unit and a Teledyne Photometrics Kinetix Scientific sCMOS camera. The dataset was acquired using a CFI SR HP Plan Apo Lambda S ×100C objective lens (silicon oil immersion, NA 1.35, WD 0.31−0.28 mm, MRD73950).
A Lumencore Celesta Light Engine (TSX5030FV, Lumencore) provided illumination for fluorescence imaging, delivering light at multiple wavelengths (405 nm, 446 nm, 477 nm, 520 nm, 546 nm, 638 nm and 740 nm).
Specifically, the dataset was acquired using 446-nm and 477-nm laser lines. In the 477-nm line optical configuration, a multiband dichroic and excitation filter set (MXR00543-CELESTA-DA/FI/TR/Cy5/Cy7-A, CELESTA) was used, along with a FITC emission filter (MXR00541 FITC, CELESTA). Instead, the 446 nm optical configuration was equipped with dual-band filters for dichroic, excitation and emission (MXR00544, CELESTA CFP/YFP, CELESTA).
In addition, the microscope has an advanced environment setup to support live imaging experiments. The Oko-Cage incubation system (Okolab) equipped with environment controllers can maintain optimal conditions for live-cell imaging, including a temperature setpoint of 37 °C, passive humidity control and CO2 levels of 5%.
Nikon NIS Elements software (v.5.42.06, Nikon) was used to manage the imaging system. Spinning disk confocal mode was selected for this dataset of two-channel z-stacks. In detail, channels 477 nm and 446 nm were imaged sequentially with identical exposure settings (200 ms). Splitting performance was tested by changing two factors: the SNR level (high, mid and low) and the relative laser power percentage of each channel, as shown in Supplementary Table 10. Multiple z-stacks of six planes with a step size of 1 μm were acquired to populate the dataset for each condition.
The HT-H24 dataset
The HT-H24 dataset was imaged by the Harschnitz group at Human Technopole and contains immunofluorescent staining of SOX2 (555) and MAP2 (488) in DIV25 dorsal forebrain organoids generated from WTC-11 (UCSFi001-A) induced pluripotent stem (iPS) cells (UCSFi001-A). Human iPS cell-derived forebrain organoids were generated following a triple-inhibition patterning protocol, without LIF supplementation, as published in a previous work25. At days in vitro (DIV) 25, forebrain organoids were fixed in 4% paraformaldehyde at 4 °C overnight, transferred in PBS with 30% sucrose until fully immersed, embedded in OCT compound (Scigen), frozen rapidly, and stored at −20 °C. The 15-μm-thick sections were sliced on a cryostat (Leica). For immunofluorescence staining, organoid slices were rinsed in PBS, incubated with Retrieval Solution (Dako) for 45 min at 70 °C, permeabilized with 0.5% Triton X-100 in PBS 1× for 20 min and blocked with 0.1% Triton X-100, 10% donkey serum in PBS 1× for 1 h at room temperature. Sections were incubated with either rat α SOX2 1:200 (Invitrogen, 14-9811-82) and chicken α MAP2 1:5,000 (Invitrogen, PA1-10005) or rat α CTIP2 1:500 (Abcam, ab18465) overnight, and counterstained with 1:1,000 donkey α chicken Alexa Fluor 488 (Jackson ImmunoResearch, 703-545-155) or 1:1,000 donkey α rat Alexa Fluor Plus 555 (Invitrogen, A48270). Nuclei were stained with DAPI (Thermo Fisher, 62248). Immunostained sections were scanned using Ti2 CREST spinning disk (Nikon), with an objective magnification of ×40.
The HT-T24 dataset
This dataset is a three-channel confocal microscopy imaging of fixed E37 ferret brain sections acquired by the Taverna group at Human Technopole. It is a three-channel 2D dataset. The first two channels contain the SOX2 (transcription factor used to label stem cell nuclei) and Golgi marker Grasp65, and the last channel contains the superimposed image containing the above-mentioned two structures. For more details, please look into the metadata of the dataset files. ‘Organism’ is ferret (Mustela furo) and the ‘Sample’ is cryosection of E37 ferret brain stained with SOX2 and GRASP65 antibodies.
Experimental animals
All experimental procedures were conducted in agreement with the German Animal Welfare Legislation after approval by the Landesdirektion Sachsen (license for ferret TVV2/2015). Animals used for this study were kept in standardized hygienic conditions at the Biomedical Services Facility (BMS) of the MPI-CBG with free access to food and water. All experiments were performed in the dorsolateral telencephalon of ferret embryos, at a medial position along the rostro–caudal axis at a stage corresponding to mid-neurogenesis.
Protocol for immunofluorescence staining
After incubation at 70 °C for 30 min in Dako Target Retrieval Solution, citrate, pH 6, cryosections were permeabilized with 0.3% Triton X-100 in PBS for 30 min at room temperature. Blocking was performed in a blocking solution (0.2% gelatin, 300 mM NaCl and 0.3% Triton X-100 in PBS) for 30 min. Primary antibodies were incubated in the blocking solution overnight at 4 °C. The following antibodies were used: SOX2 (goat polyclonal, AF2018, 1:200 dilution, R&D Systems) and GRASP65 (rabbit polyclonal, PA3-910, 1:200, Invitrogen). Subsequently, the sections were washed three times in PBS and incubated for 1 h at room temperature with the following secondary antibodies: donkey anti-goat IgG (H+L) highly cross-adsorbed secondary antibody, Alexa Fluor Plus 555 (A32816, 1:500 dilution, Invitrogen) and donkey anti-rabbit IgG (H+L) highly cross-adsorbed secondary antibody, Alexa Fluor Plus 647 (A32795, 1:500 dilution, Invitrogen). After three washes in PBS, stained sections were mounted with Mowiol.
Acquisitions of HT-T24 dataset
All images were acquired using a spinning disk confocal system, consisting of a CrestOptics V3 Light scanhead (configured with 50-μm pinholes) mounted on a Nikon Ti2-E inverted microscope equipped with a motorized stage and four Photometrics Prime 95B 25 mm cameras (pixel size 11 μm). The samples were acquired in confocal mode with a Plan Apochromat Lambda S ×100/1.35 silicon immersion objective using Celesta Lumencor solid-state lasers as the light source. Fluorescence was collected using the following elements: channel 1 (C1), excitation wavelength 638 nm laser lines at 20%, excitation filter and dichroic mirror MXR00543-CELESTA-DAPI/FITC/TRITC/Cy5/Cy7-Full Multiband Penta, Cy5 emission filter; Channel 2 (C2), excitation wavelength 547 nm laser lines at 20%, excitation filter and dichroic mirror MXR00543-CELESTA-DAPI/FITC/TRITC/Cy5/Cy7-Full Multiband Penta, TRITC emission filter; superimposed channel (Input), excitation wavelength 547 nm and 638 nm laser lines both at 20%, excitation filter and dichroic mirror MXR00543-CELESTA-DAPI/FITC/TRITC/Cy5/Cy7-Full Multiband Penta, no emission filter. All images were acquired with the same camera parameters: binning 1, 16-bit and 400 ms of exposure time. For every field of view, a z-stack with a 1-μm-step size was acquired. Once the parameters of acquisition had been defined, they were kept constant. The software used for all acquisitions was NIS Elements AR 5.42.02 (Nikon).
The HT-LIF24 dataset
This dataset was acquired at the Light Imaging Facility at Human Technopole, Milan, Italy.
Sample preparation of HT-LIF24 dataset
HeLa cells were maintained in DMEM (EuroClone) containing 10% FBS (Thermo Fisher Scientific), supplemented with 2 mM L-glutamine (EuroClone) and penicillin/streptomycin both 100 μg ml−1 (EuroClone), at 37 °C in a humidified atmosphere and 5% CO2. Cells were plated onto glass number 1.5 coverslips for immunofluorescence microscopy and then fixed with 4% PFA for 10 min. They were permeabilized with 0.1% Triton X-100 and 0.2% BSA in PBS for 10 min. Blocking was performed in a blocking solution (2% BSA in PBS) for 30 min. Primary antibodies were incubated in the blocking solution for 1 h. The following antibodies were used: anti-α-tubulin mouse IgG monoclonal (Sigma-Aldrich, T5168; 1:100 dilution); anti-laminin B1 rabbit IgG polyclonal (Abcam, ab16048; 1:200 dilution); and anti-centromere protein human IgG polyclonal (Antibodies Incorporated, 15-234; 1:400 dilution). Subsequently, after washing in PBS, cells were incubated for 40 min with the following secondary antibodies in blocking solution- Alexa Fluor 488 donkey anti-mouse IgG (Thermo Fisher Scientific, A-21202; 1:400 dilution); Cy3 donkey anti-rabbit IgG (Jackson Immunoresearch, 711-165-152; 1:400 dilution); and Cy5 goat anti-human IgG (Jackson Immunoresearch, 109-175-088; 1:50 dilution). Finally, cells were also counterstained with 4,6-diamidino-2-phenylindole (DAPI) (Sigma-Aldrich, D9542; 1:40,000 dilution) for 15 min before the mounting step. All the steps were performed at room temperature. The samples were then mounted in Mowiol-DABCO and acquired with a spinning disk system as described below. At least 200 nonoverlapping and randomly distributed fields of view were acquired and analyzed.
Acquisitions of HT-LIF24 dataset
All images were acquired using a spinning disk confocal system, consisting of a CrestOptics V3 Light scanhead (configured with 50-μm pinholes) mounted on a Nikon Ti2-E inverted microscope equipped with a motorized stage and a Photometrics Prime 95B 25-mm camera (pixel size 11 μm). The samples were acquired with a Plan Apochromat Lambda S ×40/1.25 silicon immersion objective using Celesta Lumencor solid-state lasers as the light source. Fluorescence was collected using 19 different lightpath configurations. In each configuration, a penta-band excitation filter (MXR00543-CELESTA-DAPI/FITC/TRITC/Cy5/Cy7-Full Multiband Penta), a penta-band dichroic filter (MXR00543-CELESTA-DAPI/FITC/TRITC/Cy5/Cy7-Full Multiband Penta) and a penta-band emission filter (Semrock FF01-441/511/593/684/817-25) were used. Ground truth images were acquired using a specific additional band-pass emission filter inserted before the penta-band emission filter (in particular the DAPI, FITC, TRITC and Cy5 emission filters were added for GT-A, GT-B, GT-C and GT-D configurations, respectively), whereas for all the other channels only the penta-band emission filter was used. The excitation wavelengths of the 19 channels were set as follows: GT-A, 405 nm (40%); GT-B, 477 nm (35%); GT-C, 547 nm (5%); GT-D, 638 nm (50%); A, 405 nm (40%); B, 477 nm (35%); C, 547 nm (5%); D, 638 nm (50%); AB, 405 nm (40%), 477 nm (35%); AC, 405 nm (40%), 547 nm (5%); AD, 405 nm (40%), 638 nm (50%); BC, 477 nm (35%), 547 nm (5%); BD, 477 nm (35%), 638 nm (50%); CD, 547 nm (5%), 638 nm (50%); ABC, 405 nm (40%), 477 nm (35%), 547 nm (5%); ABD, 405 nm (40%), 477 nm (35%), 638 nm (50%); ACD, 405 nm (40%), 547 nm (5%), 638 nm (50%); BCD, 477 nm (35%), 547 nm (5%), 638 nm (50%); and ABCD, 405 nm (40%), 477 nm (35%), 547 nm (5%), 638 nm (50%). All images were acquired at binning 1. For each field of view, a series of images was captured using the following exposure times: 2 ms, 3 ms, 5 ms, 20 ms and 500 ms. The software used for all acquisitions was NIS Element AR 5.42.02 (Nikon).
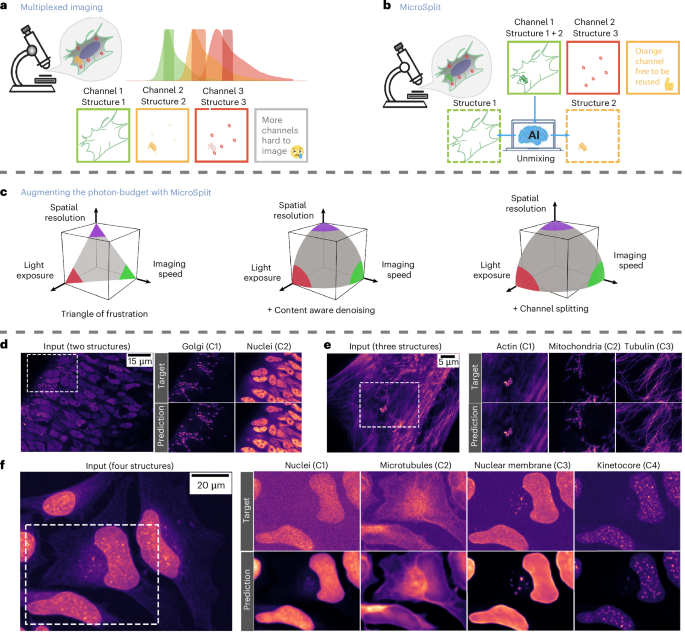
Simply put, the data contain four different structures: (1) the whole nuclei (DAPI staining); (2) microtubules, one component of the cytoskeleton, which is a filamentous system in the cytoplasm (α-tubulin staining); (3) nuclear envelope, the membrane that separates the nucleus from the cytoplasm (lamin B1 staining); and (4) the kinetochore/centromere-specific area along the chromosomes-DNA, which is used to connect the chromosomes themselves to the microtubules during mitosis (CREST staining).
The Chicago-Sch23 dataset
This dataset is four-color structured illumination super-resolution microscopy imaging of live human BJ fibroblast cells acquired at the Scherer Lab at the University of Chicago, Chicago, USA26,27,28,29. It has four channels of different structures: (1) actins (CellMask Orange); (2) mitochondria (MitoTracker Green); (3) microtubules (Tubulin Tracker Deep Red); and (4) nuclei (Hoechst).
Sample preparation
Human BJ fibroblast cells were cultured in high-glucose DMEM (Life Technologies, 10569) supplemented with 10% FBS (Life Technologies, 26140) and penicillin–streptomycin. Cells were maintained in a humidified incubator at 37 °C with 5% carbon dioxide. Before imaging, live cells were washed with PBS (Life Technologies, 15140) and trypsinized using 2.5 ml of 0.05% trypsin (Life Technologies, 25300) at 70–80% confluence. The cells were then transferred to 35-mm glass-bottom dishes (MatTek, P35G-1.5-14-C) for microscopy.
The staining solution was prepared by diluting Tubulin Tracker Deep Red (Thermo Fisher Scientific, T34077) to 1×, CellMask Orange Actin Tracking Stain (Thermo Fisher Scientific, A57247) to 2×, MitoTracker Green FM (Thermo Fisher Scientific, M7514) to 100 nM, and Hoechst 34580 (Thermo Fisher Scientific, H21486) to 5 μg ml−1 in growth medium. For imaging, cells were incubated with 1 ml of the staining solution for 30 min at 37 °C, rinsed five times with FluoroBrite DMEM (Thermo Fisher Scientific, A1896701) and subsequently imaged and analyzed in FluoroBrite DMEM.
Multicolor structured illumination microscopy super-resolution imaging
A custom-built structured illumination microscope (SIM) was used for multicolor imaging. Excitation wavelengths of 642 nm (Spectra-Physics, Excelsior One 642), 532 nm (Spectra-Physics, Millennia V), 488 nm (Spectra-Physics, Excelsior One 488) and 405 nm (Spectra-Physics, Excelsior One 405) were employed to excite Tubulin Tracker, CellMask Orange, MitoTracker and Hoechst, respectively. The four laser beams were combined using three dichroic mirrors (Semrock, Di03-R405-t1; Semrock, Di03-R488-t1; Thorlabs, DMSP550R) and subsequently expanded by 4×. The combined beams were first diffracted into monochromatic beams by a blazed grating (Thorlabs, GR13-0605) and then recombined at the plane of a digital micromirror device (DMD; Texas Instruments, DLP9000X VIS WQXGA). The DMD-generated patterns were projected onto the sample plane through an objective lens (Nikon, SR Plan Apo, ×60, 1.27 WI). A multiband dichroic mirror (Semrock, Di01-R405/488/532/635-25×36) and an emission filter (Semrock, FF01-446/510/581/703-25) were used to separate excitation and emission light. Additionally, a 2× beam expander was employed to further magnify the emission signal, resulting in a total magnification of ×120. Fluorescence images were captured using an sCMOS detector (Photometrics, Kinetix).
For SIM super-resolution imaging, striped binary patterns with a second-order spatial frequency of 2.86 μm−1 at the sample plane were displayed on the DMD. Patterns at three different angles with six phase shifts each were sequentially projected, with an exposure time of 100 ms per pattern. Super-resolution SIM reconstruction was performed using FairSIM 2, an ImageJ-based open-source software. Each raw image stack of 2,048 × 2,048 × 18 was reconstructed into 4,096 × 4,096 super-resolution images, where each pixel corresponds to 27 nm.
The HHMI-D25 dataset
Animal experiments
Heterozygous PhAMexcised female mice carrying the Mito-Dendra2 transgene were generated through in-house breeding. First, PhAMexcised heterozygous males and females (strain #018397, The Jackson Laboratory) were crossed to derive homozygous males, which were then bred with wild-type C57BL/6J females. Mice were housed in sound-attenuated, temperature- and humidity-controlled rooms under a 12-h light–dark cycle, with food and water provided ad libitum. All procedures were conducted in accordance with National Institutes of Health guidelines and were approved by the Institutional Animal Care and Use Committee at the Janelia Research Campus (protocol number 22-0229.04). Livers were collected via cardiac perfusion: first with 1× PBS to remove blood, followed by 30 ml of 4% paraformaldehyde (PFA) at a flow rate of 2.5 ml min−1 to minimize endothelial damage. Tissues were post-fixed in 4% PFA for 24 h, rinsed three times in 1× PBS and stored in 1% PFA until further processing.
Immunostaining and image acquisition
For imaging, samples were embedded in 4.6% low-melting-point agarose and sectioned into 120-μm slices using a Leica VT 1200S vibratome. Sections were blocked in 10% FBS with 0.5% Triton X-100 for 1 h and incubated with primary antibodies at 4 °C for 48 h. Mouse anti-PMP70 (MilliporeSigma, SAB4200181, 1:75 dilution) and rabbit anti-LAMP1 (Abcam, AB208943, 1:50 dilution) were used to label peroxisomes and lysosomes, respectively. Secondary antibodies Alexa Fluor 647 goat anti-mouse (Thermo Fisher, A21235, 1:500 dilution) and Alexa Fluor 750 goat anti-rabbit (Thermo Fisher, A21039, 1:500 dilution) were applied overnight at 4 °C. Additional markers included Alexa Fluor Plus 555 Phalloidin (Thermo Fisher, A30106, 1:100 dilution) and HCS LipidTox Red (Thermo Fisher, H34476, 1:100 dilution) to label actin and lipid droplets, respectively. Nuclei were stained with DAPI (Thermo Fisher, D3571, 1:1,000 dilution of 1 mg ml−1 stock) during a 25-min PBS wash the following day. Sections were cleared using EasyIndex (LifeCanvas Technologies, EI-500-1.52) by incubating samples first in 50% EasyIndex for 1 h, followed by 100% EasyIndex for 3–5 h, and mounted using Secure-Seal spacers (Thermo Fisher, 0523073). Imaging was performed on a Leica Stellaris 8 confocal microscope using a ×63/1.40 NA oil immersion objective. Images were acquired at 2,048 × 2,048 pixels using bidirectional scanning, ×2 optical zoom, and a pinhole size of 0.5 Airy units. A total depth of 10 μm was captured across 50 z-sections. Fluorophores were imaged using two acquisition lines: the first included mitochondria, actin and lysosomes, while the second included nuclei, lipid droplets and peroxisomes.
Evaluation metrics
In this work, we have predominantly used the metrics CARE-PSNR3 and \({\mathbb{M}}{\rm{icroMS}}{\mbox{-}}{\rm{SSIM}}\)8, which are slight alterations of PSNR (peak SNR) and SSIM, respectively1. We did so, as CARE-PSNR and \({\mathbb{M}}{\rm{icroMS}}{\mbox{-}}{\rm{SSIM}}\) are designed to better assess the quality of fluorescence microscopy images than their classical alternatives3,8. Please refer to Supplementary Note 3 for more details.
Model architecture and training
As shown in Fig. 1a, a superimposed image patch is fed to \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) as input. For a k channel semantic unmixing task (in this work, k ∈ [2, 3, 4]), \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) outputs k predicted images, each containing one of the structures that are superimposed in the given input patch.
In \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\), we have combined the benefits of μSplit6 and \({\rm{denoi}}{\mathbb{S}}{\rm{plit}}\)7, cast those ideas in a common learning framework, enabled direct training and prediction on volumetric data, and have extensively tested and evaluated its performance on a wide range of datasets and semantic unmixing tasks. We have also made openly available all training and prediction code and all data we used. We do this to foster rapid adoption of \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) by the scientific community and to allow others to improve our approach and compare our results with relative ease. In this section, we will discuss in detail the aspects of the above-mentioned works that were integrated into \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\). Subsequently, we also describe our loss function and important training hyperparameters.
From μSplit, we inherit the ability to efficiently incorporate spatial context using additional inputs, called LC inputs6. Here we feed, next to the primary input, for which the prediction will be made, additional LC inputs that help the network to better understand the image context from which the primary input is taken (Fig. 2a and Extended Data Fig. 1). These successive LC inputs are larger and larger patches centered on the primary input patch, but downscaled to the same pixel dimensions as the primary input itself. Hence, LC inputs capture the spatial context around the primary input patch but do so at lower resolutions to ensure efficient learning and predictions in a reasonably sized overall network.
The network architectures we proposed come in flavors that trade computational complexity and GPU consumption with the best-possible prediction quality. Its most GPU-efficient variant, Lean-LC, can train on a single GPU using less than 5 GB of GPU memory. If more resources are available, it is advisable to opt for setups such as Deep-LC, which show better predictive performance at increased computational cost.
In Extended Data Fig. 1, we have briefly described the three μSplit variants. The white regions correspond to features originating from the primary input patch. As in U-Net architectures, spatial resolution halves at each successive hierarchy level through pooling operations, causing these white areas to progressively diminish in size. In μSplit (and hence also in \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\)), the pooled embeddings undergo zero-padding before being concatenated with feature maps from lateral contextualization (LC) inputs. These LC features are processed through dedicated ‘Input branch’ sub-networks consisting of convolutional layers with nonlinear activations, dropout, and normalization components. ‘Input branch’ does not have pooling operations, and so their feature maps, at each hierarchy level, maintain spatial dimensions identical to the primary input (represented by gray regions). This preservation enables merged embeddings from both pathways to retain the original spatial dimensions throughout the network hierarchy (gray squares). This is the core idea of LC, which \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) has inherited from μSplit. In the caption of Extended Data Fig. 1, we provide more details regarding the differences in the three variants of μSplit. Ref. 6 provides further details.
From \({\rm{denoi}}{\mathbb{S}}{\rm{plit}}\), we inherit the ability to jointly perform unsupervised denoising, using suitable noise models. This also enables \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) to sample diverse predictions from a learned approximate posterior that captures a notion of the data uncertainty, as demonstrated by our trained networks being calibrated (see Section on ‘Error estimation, data uncertainty and calibration’). While also μSplit is a variational approach that is, in theory, capable of generating multiple predictions from its posterior, we found that \({\rm{denoi}}{\mathbb{S}}{\rm{plit}}\), arguably due to its different Kullback–Leibler (KL) loss formulation, produces a higher diversity that is better in line with the uncertainty in the data. In Extended Data Fig. 1, we present the architecture of \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\), which has LC inputs and noise models, all integrated into a single setup.
As mentioned above, we also enabled \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) to operate directly on volumetric image data, a possibility that was absent in both μSplit and \({\rm{denoi}}{\mathbb{S}}{\rm{plit}}\).
Loss function used to train \({\bf{Micro}}{{\mathbb{S}}}{{{\bf{plit}}}}\)
The loss function of \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) is the weighted average between the μSplit loss and \({\rm{denoi}}{\mathbb{S}}{\rm{plit}}\) loss:
$${{\rm{loss}}}_{{\rm{Micro}}{\mathbb{S}}{\rm{plit}}}=w\times {{\rm{loss}}}_{{\rm{denoi}}{\mathbb{S}}{\rm{plit}}}+(1-w)\times {{\rm{loss}}}_{\upmu {\rm{Split}}}$$
(1)
Unless explicitly specified, w = 0.9 is used in all experiments we conducted. This simple design also gives us the ability to switch to pure μSplit or \({\rm{denoi}}{\mathbb{S}}{\rm{plit}}\) mode by simply setting w to 0 or 1, respectively.
To incorporate LC inputs into the \({\rm{denoi}}{\mathbb{S}}{\rm{plit}}\) setup, we observed the need to modify the KL loss formulation used in \({{\rm{loss}}}_{{\rm{denoi}}{\mathbb{S}}{\rm{plit}}}\). In \({\rm{denoi}}{\mathbb{S}}{\rm{plit}}\), pixel-wise KL divergence is computed at every hierarchy level. Let KLi denote the pixel-wise KL divergence tensor at the ith hierarchy level. KL loss component for this hierarchy level, kli is defined as
$${\mathrm{kl}}_{i}=\alpha \times \mathop{\sum}\limits_{j,h,w}{\mathrm{KL}}_{i}[\,j,h,w].$$
(2)
With LC inputs, the spatial dimensions of the latent space tensors and therefore KLi do not decrease and the summing operation in this formulation leads to a higher value, owing to the larger number of summands (which are all non-negative). This gives unnecessarily high weight to the KL loss with respect to the likelihood loss component. We observed that this degrades the performance. To handle this, we center-cropped KLi to the shape they would assume if there were no LC inputs. Let \({\rm{KL}}_{i}^{\rm{cropped}}\) denote the appropriately center-cropped version of KLi. Our modified KL loss for \({\rm{denoi}}{\mathbb{S}}{\rm{plit}}\) becomes
$${\mathrm{kl}}_{i}=\alpha \times \mathop{\sum}\limits_{j,h,w}{\mathrm{KL}}_{i}^{\rm{cropped}}[\,j,h,w].$$
(3)
We encountered a very similar issue when working with volumetric data. In that case, pixel-wise KL divergence is a four-dimensional tensor C × Z × Hi × Wi. As we work with larger and larger Z, the summation in equation (2) would increase since the summation would be on all 4 dimensions. This again leads to giving more weight to the KL loss component against the likelihood loss, thus rendering the performance inferior. Note that this affect will become more severe when we increase the number of z frames in the input. In other words, adding more information in the input was not beneficial. To handle this, we separately took care of the extra z dimension by taking the average along this dimension. The resultant 3D tensor is then passed to equation (3) to compute the KL loss component. Note that there is an additional dimension of batch size which we have not mentioned in the above explanation. That is because KL loss is computed separately for every element in the batch.
Hyperparameters used during training
We use PyTorch package for creating our training and evaluation pipelines. We use a batch_size of 32, max_epoch of 400 and learning_rate of 0.001. We use Adamax optimizer and ReduceLROnPlateau as the learning rate scheduler with lr_scheduler_patience set to 150. During training, we use 16-bit precision. We use two LC inputs in our Deep-LC configuration (multiscale_lowres_count = 3). Please refer to our code ( for more details. All experiments were conducted using code hosted at however, we developed with the objective of providing user-friendly code with easier adaptability to custom datasets.
Additional experiments
\({{{\bf{Micro}}}}{{{\mathbb{S}}}}{{{\bf{plit}}}}\) versus PICASSO
In this section, we compare the performance of \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) with PICASSO19 (also Extended Data Fig. 8). For semantic unmixing k channels \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) needs as input a single superimposed image from the microscope whereas PICASSO needs k images from the microscope, which correspond to k spectrally overlapping fluorophores. Due to this mismatch in the data requirement, a direct comparison is not feasible. So, to compare \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) with PICASSO, we generate synthetic inputs using our HT-LIF24 dataset. However, we argue that our way of generation does not degrade the performance of PICASSO but instead, it should be easier for PICASSO to predict on this data as compared to the real data.
As fluorophores can be ordered according to the wavelength of the maximum intensity in their emission spectra, we first define such an order of our structure types. Next, for generating every channel of the input for PICASSO, we define three weights and take the weighted average of the three structures using these weights. The weights are set according to the order of the structures set above. For example, for the first channel, the weight given to the second structure will be higher than the weight given to the third structure. We generate two sets of weights, one being harder than the other. In the hard case, the dominant structure type is given 1.5 times more weight than the next dominant type and three times more weight than the least dominant type. In the easy version, the dominant structure type is given 2.5 times more weight than the next dominant type and five times more than the least dominant type. Once the input channels are generated, we add Gaussian noise of σ = 500 to each channel independently. We also train \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) on this data with the same Gaussian noise applied on top of the data. In Extended Data Fig. 8, we show the results on one random frame. In Supplementary Tables 7 and 8, we show the quantitative results.
Training Mode I versus Training Mode II (how important are spatial correlations?)
In this experiment, we inspect the performance drop between data acquisition types I and II. In cells, the location of different structures is often quite co-related with one another. For example, actin is mostly concentrated on the cell periphery whereas the nucleus is typically found at the center of the cell. In acquisition modes I and II, input is created by summing the crops from individual channels. In acquisition type I, as the channels are independently acquired, summing the crops of these different channels will generate an input patch where the naturally occurring colocation property cannot be preserved. In acquisition type II, since both channels are concurrently acquired, the inputs are created from summing the crops of different channels with each crop taken from the same location in the micrographs and therefore these inputs preserve the naturally occurring colocation property. In this experiment we quantify the benefit of using this colocation information.
We work with the HT-T24 dataset, which falls under Training Mode III. We train three models. In the first model, we create the input using Training Mode I (we create the input by picking target patches from the same location and therefore maintain the spatial co-relation). In the second model, we use Training Mode II, meaning that we pick crops from random locations from the different channels and use them to create the input. This model naturally does not have access to naturally occurring spatial co-relation information in its training data. The third model is trained using Training Mode III. We evaluate all three models on the held-out test set where the inputs have spatial co-relation preserved and are not synthetic (they are imaged from the microscope). In Supplementary Table 4, we find that the first model outperforms the second by 1.3 CARE-PSNR and 0.009 \({\mathbb{M}}{\rm{icroMS}}{\mbox{-}}{\rm{SSIM}}\). Naturally, the model trained with Training Mode III is best and outperforms Training Mode I by 0.6 CARE-PSNR and 0.004 \({\mathbb{M}}{\rm{icroMS}}{\mbox{-}}{\rm{SSIM}}\).
Training Mode I versus Training Mode III (summed versus acquired inputs)
Here, we quantify how much the performance degrades if, during training, the input is created by simply summing the two channels as compared to input coming directly from the microscope. We find that while there is indeed a performance drop as can be seen in Supplementary Table 4, the drop is not detrimental. This experiment shows the utility of our approach in the case when synthetic input is used for its training but for evaluation, inputs coming directly from the microscope are used.
\({{{\bf{Micro}}}}{{{\mathbb{S}}}}{{{\bf{plit}}}}\) enables a more effective use of the available photon budget
By filtering fewer photons: traditional multiplexed imaging relies on emission filters that selectively pass photons from one fluorophore at a time to minimize spectral overlap. As a consequence, a substantial fraction of emitted photons is discarded, and relaxing the filters leads to bleedthrough artifacts. \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) changes this trade-off because multiple structures can be imaged in the same acquisition. This allows microscopists to use substantially broader emission filters, collecting photons from several fluorophores simultaneously, without introducing the ambiguities that would otherwise arise in a multichannel setting.
To roughly quantify this advantage, we analyzed three fluorophores from the HT-LIF24 dataset (DAPI, FITC and TRITC), using their emission spectra (downloaded from fpbase.org) normalized as probability mass functions. We compared the fraction of photons transmitted by conventional multicolor filter configurations to the photons captured when using a broad, highly permissive filter suitable for \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\). Across three representative scenarios, with emission filter thresholds chosen to (1) maximize photon collection; (2) reduce bleedthrough by 25%; and (3) reduce bleedthrough by 50%, with conventional multiplexed imaging being respectively 22%, 34% and 55% fewer photons efficient than the results obtained using \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) (Extended Data Fig. 7).
In practical terms, this means that even in bleedthrough-optimized multiplexed imaging, each channel discards a large fraction of emitted photons, whereas \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) can reclaim much of this loss by aggregating photons from multiple fluorophores in a single measurement.
By enabling gentler imaging due to denoising: our method, next to performing semantic unmixing, also performs unsupervised denoising. As denoising improves the SNR of images it is applied to, microscopists can acquire the raw data more gentle, accepting a lower initial SNR3. To illustrate this on a concrete example, we assessed the similarity of a biological structure imaged at various exposure times between 2 ms and 20 ms with very high SNR data of the same regions of interest acquired at 500-ms exposure time. More concretely, we have conducted these experiments on the three-channel data (Nucleus, Microtubules and Kinetocore) of the HT-LIF24 dataset. As shown in Supplementary Table 11 (bottom), even the denoised 2-ms exposure micrographs lead to a considerably higher quality with regard to the 500-ms images than even the 5-ms raw data, for channel 1 even compared to the 20-ms raw acquisitions, suggesting at least a threefold to tenfold reduction of the required photon budget and therefore enabling users of \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) to image considerably more gentle to reach the same quality required for downstream processing.
Statistics and reproducibility
For both qualitative and quantitative evaluations, we generate 50 predictions per input using the trained \({\rm{Micro}}{\mathbb{S}}{\rm{plit}}\) model, running it 50 times. Predictions employ tiled inference with overlap: each test input frame is divided into overlapping 64 × 64 patches; the 50 predictions per patch are averaged, and the central 32 × 32 region is extracted. These regions from overlapping patches are then stitched together to form a full-frame prediction. The same approach applies to 3D models. Please refer to the supplementary material for more details. Thus, metrics in Table 1 (with standard errors in Supplementary Table 12) and qualitative results in Figs. 1d–f, 3a and 4 all use these 50-prediction averages.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.