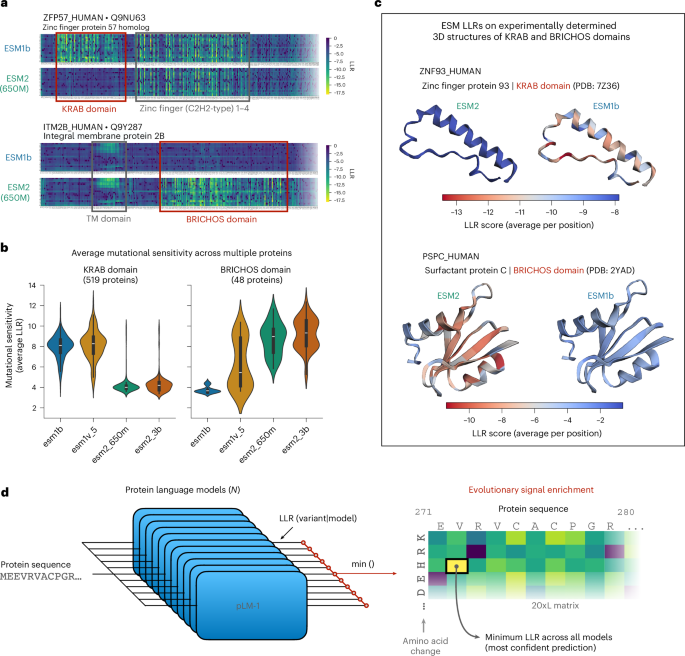
Enriching for evolutionary indicators inside the ESM household of fashions
Scoring variant results
ESM fashions are pretrained with the masked language modeling goal to foretell possibilities of amino acids being at specific positions in protein sequence. Based mostly on this property, we comply with the earlier work21 to attain the impact of a mutated sequence (G^v) by evaluating it with the corresponding WT sequence (v^) utilizing LLR, as follows:
$$mathrmv(v^|v^v)=mathopvlimits_vlog p(_={v_^G}|_v)-log p(v_v={v_^{mathrmv}}|{x}_{backslash t}),$$
the place T denotes the set of mutated positions, (x^{mathrm{wt}}_t) and (x^{mathrm{mt}}_t) denote the wild-type and mutant amino acids at place t, and (p({x}_{t}=alpha|{x}_{backslash t})) denotes the chance of amino acid (alpha) predicted by the mannequin at place t conditioned on the remainder of the sequence. Intuitively, by way of variant impact, the LLR rating represents the benign diploma of a mutation in comparison with the reference sequence. Increased scores point out extra benign and vice versa. Following the WT marginal chance scheme21 for inference effectivity, we carry out a single ahead cross on the WT sequence to attain the chances and calculate the rating for all mutations.
Evolutionary sign enrichment and the ESMIN dataset
Given a set of pretrained ESM fashions, for every mutation, we calculate LLR scores for all fashions and take the minimal LLR rating (that’s, most pathogenic or damaging prediction throughout all fashions) to create a pseudo-LLR rating for the mutation. This strategy selects the mannequin with the utmost confidence within the WT residue relative to a given substitution for that place, successfully prioritizing fashions which have recognized the corresponding residue as a part of a mutationally delicate, evolutionarily conserved motif. Following the above course of, we create the ESMIN dataset with 11 ESM fashions (ESM2: 8M, 35M, 150M, 650M and 3B; ESM1b; and 5 ESM1v fashions) (Supplementary Desk 1), utilizing 20,284 manually reviewed human protein sequences from UniProt (February 2022)52. For proteins longer than 1,022 amino acids (most context window of ESM1b and ESM1v fashions), we comply with a sliding window strategy6 to section them into 1,022-residue home windows with an overlap of 511 residues. For every protein, we compute scores for all potential (19times L) missense variants (the place L is the protein size), leading to ~197 million variants in complete, throughout 2.11 × 104okay segments. We additional filter out noninformative protein segments (that’s, segments whose ESMIN scores are all comparable in a variety near zero), by first computing the minimal ESMIN rating throughout all variant positions (besides the primary) for all segments after which filtering out segments whose minimal ESMIN rating is increased than the ninety fifth percentile of the distribution. This ends in a complete of 19,600 segments throughout 18,683 proteins with a complete of 191.3 million distinctive variant scores. The ensuing ESMIN scores vary from −31.2 to 7.02, with the rating of the WT variants being 0.
Most confidence co-distillation and coaching of VESM fashions
We develop a co-distillation framework to enhance all particular person sequence-only fashions inside the ESM household (Supplementary Desk 1). Simplifying notation from the earlier part, we let (V) denote the set of all potential mutations within the human proteome and (F(G)) be the corresponding log chance ratio assigned to the variant (v) in (V) by mannequin (G). The primary iteration for co-distillation of (n) fashions ({G}_{1},ldots ,{G}_{n}) utilizing MSE because the loss operate, is given by
For variant (v) in (V) and mannequin ({G}_{i}), i = 1, .‥, n:
({mathrm{Loss}}_{i}) = MSE((F({G}_{i})), (mathop{min ,}limits_{s}F({G}_{s})))
backpropagate(({mathrm{loss}}_{i}), mannequin ({G}_{i})).
To keep away from coaching (n) fashions in parallel and to bypass the necessity to recompute (mathop{min ,}limits_{s}F({G}_{s})) scores in every iteration, we think about a simplified strategy that precomputes and fixes (mathop{min ,}limits_{s}F({G}_{s})triangleq) ESMIN((v)) for all iterations. On this case, co-distillation is carried out by the distillation of the precomputed ESMIN scores to every particular person mannequin that contributed to its computation, that’s, by setting within the above, ({mathrm{loss}}_{i}) = MSE((F({G}_{i})), ESMIN((v))). Lastly, for higher coaching convergence and to take care of co-distilled mannequin scores in an identical vary with respect to their pretrained variations, we shift the ESMIN goal scores for every mannequin to match the empirical imply of (F({G}_{i})) throughout all variants (v) in (V). These shifts are calculated as soon as utilizing the LLR scores from the bottom ESM fashions and are stored fixed throughout coaching.
Parameter-efficient coaching
We freeze the low-level layers of the pretrained ESM fashions and fine-tune solely: (1) the final hidden layer and (2) the language mannequin head. The trainable parameters account for a small fraction of the overall variety of parameters, for instance, roughly 3.3% and a pair of.8% of the overall parameters for ESM2 (650M) and ESM2 (3B) fashions respectively. All fashions had been skilled with precisely the identical settings for 5 epochs. Following commonplace follow for fine-tuning pretrained encoders53,54, we scale the training price based mostly on coaching period and use AdamW with a price of 4 × 10−4, which is obtained by adjusting a regular 1 × 10−3 price to a five-epoch schedule. AdamW β are set to (0.9, 0.999) and weight decay is 0.01, and we apply a cosine scheduler with a 5% warmup. We randomly cut up the proteins utilizing a small set of 200 segments for validation and the remaining (19,434 segments) for coaching.
Computing sources
All experiments had been performed on a single NVIDIA H100-SXM GPU (81.5-GB VRAM) per run. The workstation has 4xH100-SXM, however no mannequin/information parallelism used for coaching. For all coaching runs we used: Python (3.12.3), PyTorch (2.3.0), CUDA (12.1), cuDNN (8.9.7) and blended precision (FP16). All different packages used are included within the ‘environment.yml’ file positioned in our github repository ( Coaching of the most important ESM-3B mannequin on our system with the above settings (final layer + head; 85.4 million trainable parameters) took ~18 h with peak VRAM measured at ~77 GB. For comparability coaching of the 35M-parameter mannequin took solely 35 min. See Supplementary Desk 2 for extra particulars.
Assessing the generalization capabilities of the co-distilled fashions
To judge the generalization functionality of our co-distilled fashions, we conduct an ablation research by various the proportion of protein sequences used for coaching. First, we exclude from the coaching pool all human proteins sharing greater than 30% sequence similarity with any protein in our analysis benchmarks (2,400 proteins in Balanced ClinVar and 101 proteins in DMS organismal health and exercise assays). To do that, we merged the coaching and benchmark sequences and clustered them with MMseqs255,56 (simple cluster; launch 13-45111) utilizing –min-seq-id 0.30, –cov-mode 0 (protection relative to the longer sequence), -c 0.8 (≥80% aligned size) and –cluster-mode 1 (connected-component). We then eliminated any coaching sequence that coclustered with a benchmark member. This filtering step excludes 5,279 sequences from the total coaching set of 18,683 human proteins. From the remaining 14,355 proteins, we create nested subsets of lowering measurement (50%, 10%, 5% and 1% of all sequences) to evaluate mannequin efficiency with restricted coaching information. Subset sampling was repeated 5 instances with unbiased random seeds. For every subset, we fine-tuned three ESM2 fashions (8M, 35M and 150M) utilizing a 90%/10% coaching/validation cut up and following the identical coaching settings described above.
Positive-tuning on downstream duties utilizing VESM mannequin embeddings
To evaluate the impact of our co-distillation framework on the protein illustration capabilities of ESM fashions, we consider the efficiency of the embeddings from each the bottom and co-distilled variations of ESM2-650M (VESM2) on 9 downstream duties throughout 4 classes: (1) operate prediction: thermostability prediction job from FLIP57 that predicts the thermostability stage of proteins, steel ion binding prediction58, (2) localization prediction: subcellular localization59 with two location classes (DeepLoc-2, binary classification) and ten location classes (DeepLoc-10, multiclass classification), (3) protein–protein interplay prediction: human protein–protein interplay (HumanPPI) prediction59,60, (4) mutational impact prediction: adeno-associated virus health57, β-lactamase exercise59, fluorescence prediction and protein stability duties from TAPE61. We comply with the identical prepare/legitimate/check splits offered by SaProt14.
We freeze the embeddings from each fashions and prepare a (head) neural community constructed on the highest of ESM’s embedding. For Thermostability, HumanPPI, MetalIonBinding, DeepLoc-2 and DeepLoc-10, we use the ESMClassification module and the identical fine-tuning framework as in SaProt14. For the remainder of the duties, we design an identical two-layer community on high of the ESM’s embedding that may be a sequence of a linear layer (from embedding measurement to embedding measurement) and a LayerNorm layer, adopted by a GELU activation and a linear projection layer. We prepare the community utilizing the AdamW optimizer with β = (0.9, 0.98) and a cosine linear scheduler with studying price of 1 × 10−4. We use MSE loss for regression duties and cross entropy loss for classification duties. For all duties we repeat the coaching thrice with completely different random seeds and report the imply and std of every mannequin’s efficiency.
A number of rounds of co-distillation
We chosen the highest 4 co-distilled fashions from the primary spherical (ESM2-3B, ESM1b, ESM2-650M and ESM1v5) and carried out two further rounds of co-distillation (rounds 2 and three; Fig. 3b). The co-distillation course of was stored the identical as spherical 1, with the one distinction being that per-variant LLR scores throughout the taking part fashions had been aggregated by averaging as a substitute of taking the minimal. In spherical 2 we co-distilled these 4 fashions on the identical set of human proteins utilized in spherical 1. For the third spherical, to extend studying capability we repeated averaging co-distillation utilizing a dataset completely composed of nonhuman proteins (see under). For all rounds, coaching continued from every mannequin’s previous-round checkpoint.
Coaching setting
To extend mannequin capability whereas retaining effectivity, we fine-tune the pinnacle and the final three embedding layers. This raises the variety of trainable parameters shut to eight% for ESM2 (3B). We lowered the training price to 1 × 10−4 to proceed coaching from the earlier spherical. Weight decay stays 0.01 however is utilized solely to major weight matrices (excluding biases and normalization parameters), following greatest practices to mitigate forgetting and preserve coaching stability62. The identical cosine scheduler with 5% warmup is used.
Nonhuman protein sequences
To generate the nonhuman dataset we began from a UniProtKB reviewed export (downloaded 20 February 2025) and first compiled the set of human protein-family names by parsing the ‘protein families’ subject of Homo sapiens data. We then retained solely nonhuman entries that (1) had an InterPro accession, (2) had been lower than 1,022 amino acids lengthy and (3) whose circle of relatives label was not within the human set (entries missing a household label weren’t excluded by this step). To take away redundancy, remaining sequences had been ranked by an proof rating derived from UniProt the ‘protein existence’ subject (the place 1 is the protein stage, 2 is the transcript stage and three is the homology), with a +0.5 penalty if the household label was lacking, and the top-ranked sequence per InterPro accession was stored. This course of yielded a dataset of 23,803 nonhuman proteins that was used each within the closing spherical of co-distillation, in addition to in all downstream information distillation steps (see under).
Multiround ablation research
We ablated the aggregation operator utilized in every spherical, denoting a schedule as agg1–agg2–agg3 (for instance, min–avg–avg means minimal aggregation in spherical 1, common in rounds 2 and three). We evaluated three aggregation mixtures: (1) avg–avg–avg, (2) min–min–min and (3) min–avg–avg, throughout all three rounds, reporting intermediate outcomes the place, min–avg for instance denotes the corresponding results of the second spherical (Prolonged Information Fig. 4). We notice that to comply with the identical setting we used for our major outcomes, spherical 1 co-distillation was carried out by aggregating LLRs throughout all 11 fashions, not solely the 4 used for the next rounds. Throughout each benchmarks, and all 4 fashions evaluated, utilizing most confidence in spherical 1 was important for attaining the strongest downstream efficiency. Averaging throughout a number of rounds was not capable of additional enhance efficiency. After utilizing minimal aggregation in spherical 1, the selection between averaging and minimal had a smaller impact, with averaging in rounds 2 and three attaining the perfect general outcomes (Prolonged Information Fig. 4b).
Distilling VESM-3B into 650M-, 150M- and 35M-parameter fashions
To broaden accessibility throughout compute budgets, we distilled the converged mannequin from spherical 3 (VESM-3B) into the bottom (non-co-distilled) ESM2 backbones at 650M, 150M and 35M parameters. First, VESM-3B was used to compute goal LLR scores on the union of the human and nonhuman protein units described above, after which every ESM2 mannequin was skilled on these goal LLRs with an MSE loss, as described in our co-distillation framework. For this information distillation step, we fine-tuned the total fashions for ESM2-35M and ESM2-150M, and for the bigger ESM2-650M, we fine-tuned the output head and the final 5 embedding layers (~100 million parameters). We used the identical optimization setup as within the multiround coaching: AdamW with β (0.9, 0.999) and a cosine studying price scheduler with 5% warmup. The educational charges had been set as follows: 1 × 10−3 for ESM2-35M, 1 × 10−3 for ESM2-150M and 1 × 10−4 for ESM2-650M. The ensuing fashions are known as VESM-35M, VESM-150M and VESM-650M.
Distilling VESM-3B into ESM3
Having established our co-distillation framework utilizing sequence-only fashions from the ESM household, we subsequent prolong this strategy to the sequence part of a multimodal protein mannequin. Particularly, we apply our framework to the ESM335 mannequin (esm3-sm-open-v1; 1.4B parameters; which integrates a number of modalities together with sequence, construction, solvent accessibility, secondary construction (SS8) and useful annotations. Our goal is to fine-tune solely the sequence-related modules of ESM3, whereas preserving the remaining parts mounted at their pretrained weights.
For coaching, we assemble input-output pairs utilizing protein sequences and VESM-derived LLRs, to distill the VESM (3B) mannequin into the sequence module of ESM3. To extend coaching variety, we use each units of human and nonhuman protein sequences described above. We notice that no 3D construction or some other modality apart from sequence is used as enter throughout coaching. Since ESM3 makes use of the identical head for construction conditioned and sequence solely inference, we preserve the pinnacle parameters frozen throughout coaching and fine-tune solely the final 5 embedding layers of the mannequin. We use the identical optimization setup as above (information distillation, ESM2 fashions) with studying price 5 × 10−5.
Throughout inference, we allow each sequence and 3D construction inputs to the fine-tuned mannequin. The structural enter is processed utilizing ESM3’s default configuration (pretrained weights), ensuing within the VESM3 mannequin described within the Primary. We notice that using 3D construction as enter to VESM3 throughout inference is elective and that the fine-tuned mannequin retains the capability to simply accept all eight enter modalities supported by ESM3. On this work, we solely evaluated efficiency of VESM3 utilizing 3D construction as enter. To acquire VESM++ scores, we common the LLRs from VESM3 and VESM-3B, that’s, LLR(VESM++) = (LLR(VESM3) + LLR(VESM-3B))/2.
Inference runtime
All VESM fashions developed on this work inherit the computational effectivity of their corresponding base fashions inside the ESM household. To offer steering with respect to compute sources required for utilizing our fashions in follow, we performed a set of runtime experiments reporting peak VRAM (GB), throughput (seq s−1) and latency (ms) throughout three completely different sequence lengths (brief, 254 amino acids; medium, 510 amino acids; and lengthy, 1,022 amino acids). Supplementary Desk 3 supplies the total outcomes for the 35M-, 650M- and 3B-parameter fashions (sequence-only VESM), in addition to VESM3 with sequence + construction enter. We additionally included ESM2-15B for reference. Batch measurement was set to 16 for all experiments and for every metric we reported the typical over steady-state home windows after warmup, with commonplace deviation throughout two runs.
For instance, we will use the above outcomes to match runtime estimates for calculating the LLRs of all potential single missense mutations in a set of two.5 × 104 protein sequences, the place each is ~1,000-amino-acids lengthy (that’s, ~450 million variant impact scores in complete). For this reference dataset, VESM-3B would want ~1 h of inference time, whereas consuming lower than 16 GB of VRAM. VESM-650M would want ~20 min and the smallest sequence-only VESM mannequin (35 million parameters) would full this job in round 3 min. For comparability, VESM3 would additionally want ~40 min however would require a a lot bigger VRAM reminiscence footprint measured at ~46 GB. We notice that each one of our experiments had been carried out on our native workstation (4xH100-SXM GPU) so whereas absolute wall-clock time estimates could differ when carried out on different techniques/configurations, the relative efficiency throughout fashions and the height VRAM necessities shall be extra straight comparable with the estimates offered above.
Datasets
ProteinGym ClinVar
The dataset consists of 62,656 missense variants masking 2,525 human genes. To keep away from information circularity, we excluded supervised strategies and meta-predictors which have been skilled on medical labels (ClinPred, VEST4, REVEL, MetaRNN, BayesDel and MutationTaster) and strategies with greater than 10,000 lacking values. We then stored all variants for which all strategies have out there predictions. This resulted in a dataset with 52,637 variants (26,897 pathogenic and 25,740 benign) throughout 2,227 genes that we used for our analysis. We additionally generated a subset of this dataset (known as Balanced ClinVar), by subsampling majority class labels to match the variety of pathogenic and benign variants for every protein. This balanced model of ClinVar features a complete of 27,468 variants throughout 2,400 proteins.
Full ClinVar dataset obtain
To check with AlphaMissense we downloaded the newest launch of ClinVar (March 2025 model) and computed MAFs for all variants utilizing gnomAD v4. The ClinVar dataset was filtered to incorporate all hg38 SNVs with predicted results on RefSeq NM ids. To map between hg38 coordinates, RefSeq and UniProt accessions, we used the newest RefSeq annotation launch (GCF_000001405.40-RS_2024_08) and the UniProt ID mapping server (out there at uniprot.org/id-mapping). For high quality management we stored variants annotated with at the very least 1 ‘Gold Star’, leading to a dataset of 151,600 missense variants. For these variants we added MAF computed from gnomAD_v4 utilizing the AF column, as MAF = min(AF,1-AF). For variants not included in gnomAD v4, we set MAF = 0. We additionally computed MAF from gnomAD v2 (v2.1.1, liftover_grch38) and annotated variants which have gnomAD v2 MAF >1 × 10−5 as ‘used for training AlphaMissense’. Lastly, we mapped precomputed AlphaMissense scores utilizing hg38 genome coordinates from AlphaMissense_hg38.tsv.gz (downloaded from After filtering out lacking values, we obtained a dataset of 142,951 variants (46,247 pathogenic and 96,704 benign) throughout 2,685 genes.
ProteinGym DMS
The DMS benchmark consists of 217 DMS assays masking 696,311 single variants. The assays are categorized by taxon (96 human, 40 eukaryote, 50 prokaryote and 31 virus) and coarse choice kind (77 organismal health, 43 exercise, 66 stability, 18 expression and 13 binding assays). For our evaluations we additional mix them into construction associated assays (stability, expression and binding, 97 complete) and performance associated assays (health and exercise, 120 complete). We used the offered experimental scores for all DMS assays aside from health and exercise measurements of CALM1, TPK1, UBC9, RASH, TADBP, SYUA and SRC. For these assays we utilized a change of the shape x → abs(x − WT), the place WT represents the WT measurement (both 0 or 1, relying on the assay). This adjustment displays the truth that, for these assays, variants scoring increased than the WT are usually related to deleterious results, as mentioned within the authentic research63. This preprocessing step improved efficiency throughout all evaluated strategies, excluding the structure-based mannequin ProSST19, whose efficiency on TADBP dropped considerably, from 0.54 to 0.08 (Prolonged Information Fig. 6b). Though we had been unable to pinpoint the reason for this discrepancy, it’s unlikely to be associated to structural options, because the areas assayed in TADBP are intrinsically disordered64.
Protein construction information for analysis
For ProteinGym DMS, we used the 3D buildings offered by the benchmark. For ProteinGym ClinVar, we used 3D construction offered by AlphaFold Protein Construction Database ( for two,300 sequence-matched proteins and generated 3D construction for the remaining proteins utilizing the AlphaFold2 script from ColabFold65 (https://github.com/sokrypton/ColabFold).
Comparisons with state-of-the-art PLMs and VEP strategies
We in contrast VESM fashions with 25 fashions on ClinVar and 39 fashions on the DMS benchmark. These baselines are the newest or broadly used PLMs and VEP strategies, together with PLMs skilled on unaligned sequences (for instance, ESM, ProGen2 and Tranception), structure-based fashions (SaProt14, ESM335, ProSST19 and ISM45), MSA-based fashions (PoET20, EVE36, TranceptEVE18, GEMME37, VespaG38 and RSALOR61) and different VEP strategies that use homology-based (for instance, SIFT39 and PROVEAN40) or population-based (for instance, PrimateAI13 and CADD42) approaches. See Supplementary Desk 4 for an in depth description.
For the ProteinGym benchmark, we obtained precomputed scores from ProteinGym for all strategies besides RSALOR61, ProSST, SaProt, ESM3, ISM, VespaG and ESMC. Amongst these fashions, ISM scores are at present not reported within the benchmark. Different fashions don’t have scores for the ClinVar benchmark. Moreover, we needed to make sure that all structure-based fashions use the identical set of 3D buildings for analysis. Due to this fact, we compute variant impact scores for all the above fashions, following the inference scripts offered within the corresponding repositories. For the comparability with AlphaMissense7, we used the precomputed scores offered by the authors (https://zenodo.org/records/10813168).
Efficiency analysis on ClinVar
We evaluated efficiency on ClinVar utilizing the world underneath the receiver working attribute curve (AUC) as our major metric. We reported each international AUC (calculated throughout all variants) and common per-gene AUC (calculated individually for every gene with at the very least ten annotated variants). We bootstrapped the worldwide AUC calculation in Fig. 4a, by randomly sampling 50 label-balanced units of 6,000 variants (3,000 benign and three,000 pathogenic). The plot reveals the estimated imply and commonplace deviation.
Prediction accuracy versus variant annotation
For this evaluation, the scores of every methodology had been first calibrated by performing logistic regression on a small set of ClinVar variants (n = 1,000), randomly sampled to include an equal variety of benign and pathogenic labels. This step ensures that each one the scores are comparable between 0 and 1, with 0.5 being the classification threshold. Then we diverse the classification threshold symmetrically to exclude variants that every methodology is most unsure about (with scores round 0.5). For every threshold, we computed the prediction accuracy, outlined as the typical accuracy of appropriately annotating each benign and pathogenic variants, in addition to the overall variety of variants that every methodology was capable of annotate (that’s, those that weren’t excluded by the edge). Total, this course of was repeated ten instances by randomly sampling the calibration set, and the 2 portions (± 2 commonplace deviations across the imply) had been plotted in opposition to one another within the determine.
Comparability with AlphaMissense/MAF filtering
We in contrast VESM fashions with AlphaMissense throughout completely different MAF thresholds (1 × 10−5, 1 × 10−4, 1 × 10−3, 1 × 10−2 and 1 × 10−1) to evaluate the influence of inhabitants frequency on mannequin efficiency. For every threshold, we filtered the ClinVar dataset (Feb 2025 launch) to incorporate solely variants under the desired MAF worth and calculated the AUC for every mannequin, by randomly sampling 100 label-balanced units of 20,000 variants (10,000 benign and 10,000 pathogenic) from genes that include each labels. Even on the lowest MAF threshold (1 × 10−5) the filtered ClinVar dataset remained nicely balanced with a complete of 18,145 benign and 40,046 pathogenic variants throughout 2,559 genes. We additionally carried out a particular comparability on the subset of ClinVar variants not utilized in AlphaMissense coaching, by eradicating gnomAD v2 variants with MAF > 1 × 10−5. This resulted in a set of 31,967 pathogenic and 15,392 benign variants throughout 1,510 genes. We emphasize that this filtering step relies on MAF calculated based mostly on gnomAD v2 and doesn’t exclude 4,818 variants which can be reported in gnomAD v4 with MAF >1 × 10−5. As above the imply AUC and commonplace deviation was calculated over 100 bootstraps however with 10,000 (as a substitute of 20,000) variants sampled in every iteration to account for the smaller measurement of the filtered dataset. On this dataset, we additionally computed binary classification metrics by first calibrating predictions for all fashions as in Fig. 4c (logistic regression, n = 1,000 variants) and utilizing 0.5 because the classification threshold. The reported metrics are estimated by sampling the calibration set 50 instances.
Efficiency analysis on DMS information
We evaluated mannequin efficiency on the ProteinGym DMS benchmark utilizing the weighted common of Spearman correlations between mannequin predictions and experimental measurements, following the scripts and scoring tips outlined within the ProteinGym github repository17 ( We in contrast all VESM fashions (3B, 650M, 150M, 35M, VESM3 and VESM++) in opposition to 39 different strategies (see ‘Comparisons’ in Supplementary Desk 4) throughout 4 datasets: (1) human DMS assays measuring health and exercise (Fig. 5a), (2) all DMS assays measuring health and exercise (Fig. 5b,c), (3) all DMS assays measuring binding, stability and expression (Fig. 5b) and (4) nonhuman DMS assays measuring health and exercise (Fig. 5d).
To evaluate consistency between human DMS and ClinVar benchmarks, we calculated the correlation between mannequin performances on the 2 datasets, contemplating solely strategies evaluated throughout each (Fig. 4a, Fig. 5b). Along with common efficiency, we computed a rank rating by sorting strategies in ascending order from 1 to 43 based mostly on their weighted common Spearman correlations (Fig. 5b). We additionally assessed pairwise win charges66 between fashions (Fig. 5c). For every pair of fashions, we computed the proportion of assays the place the primary mannequin achieved the next Spearman correlation than the second. Formally, the win price of mannequin ({m}_{1}) over ({m}_{2}), is outlined as (mathrm{win},mathrm{price}({m}_{1},{m}_{2})=frac{1}{N}{sum }_{i=1}^{N}1{{S}_{i}({m}_{1}) > {S}_{i}({m}_{2})}), the place ({S}_{i}(m)) denotes the Spearman correlation achieved by mannequin (m) on assay i, (N) is the overall variety of assays and (1{cdot }) is the indicator operate. We notice that though win charges had been utilized in earlier work66 to match between fashions with nonoverlapping units of VEP, right here we examine fashions on precisely the identical assays and variants.
UK biobank abstract statistics and the Genebass medical phenotyping benchmark
Genebass abstract statistics
We downloaded abstract statistics (gene-level impact measurement and affiliation P worth) for the highest pLoF gene burden associations with blood biochemistry biomarkers (pLoF SKAT-O P worth <1 × 10−6) from the Genebass internet portal ( together with solely genes with at the very least 25 missense variants. This resulted in a dataset of 332 gene–phenotype pairs throughout 186 genes and 27 biomarkers. For every gene–phenotype pair, we then downloaded the single-variant affiliation abstract statistics (174,073 impact sizes and P values throughout 80,245 distinct missense variants). Lastly, we annotated the corresponding gene-level impact sizes and P values for 153 out of 332 gene–phenotype pairs that had been recognized as vital by Missense SKAT-O (P worth <1 × 10−6).
Variant-level VESM rating regression
For every gene–phenotype pair we carried out a linear regression of the variant-level VESM predictions in opposition to the Genebass-derived single-variant impact sizes (β coefficients) and reported the standardized slope (Pearson correlation) as a proxy of the gene stage impact measurement estimate and the corresponding VESM regression derived P worth.
Scientific phenotyping benchmark
To create a benchmark based mostly on Genebass abstract statistics we needed to make sure that the included gene–phenotype pairs contained detectable indicators utilizing missense variants alone. We subsequently additional filtered the dataset to incorporate solely (1) gene–phenotype pairs that had been recognized as considerably related by Missense SKAT-O (P worth <1 × 10−10) and (2) missense variants that had been individually related to the phenotype (single-variant affiliation P worth <0.05). This resulted in a dataset of 103 gene–phenotype pairs throughout 61 genes and 25 phenotypes. We evaluated strategies on this dataset by making use of the identical linear regression strategy described above to predictions derived from VESM (3B, 650M, 150M and 35M), VESM3 and VESM++, AlphaMissense, ESM1b, ESM2 (650M) and the log10-transformed MAF of the variants (Fig. 6b). To make sure robustness of the benchmarking outcomes we additionally evaluated the above strategies on datasets derived utilizing completely different selections for the filtering thresholds (Prolonged Information Fig. 7a-b).
License
We notice that the predictions and weights of the VESM3 and VESM++ fashions are derived from fine-tuning the ESM3-small-open mannequin developed by EvolutionaryScale which is obtainable underneath a noncommercial license settlement (see for detailed license choices for these fashions).
Reporting abstract
Additional data on analysis design is obtainable within the Nature Portfolio Reporting Abstract linked to this text.