TxPert architecture
TxPert forecasts how genes will respond to perturbations (represented as vectors in ℝⁿ) based on a collection of perturbation identifiers (P ⊆ 𝒫 := {1, …, N}) and a baseline cellular state derived from a control gene expression profile (x ∈ 𝒳 ⊂ ℝⁿ). This control profile has been carefully matched to the target cell in terms of cell line and experimental batch to minimize technical variability. Here, n ∈ ℕ denotes the number of genes measured in the experiment, while N ∈ ℕ refers to the total number of distinct perturbations observed across the dataset. These perturbation identifiers allow the model to retrieve corresponding node embeddings from a biological knowledge graph (KG) encoding gene–gene (or gene product–gene product) interactions. These embeddings are then fused with the basal state representation to generate the predicted perturbed expression profile.
To integrate the baseline cellular state with perturbation information, TxPert first maps both into a shared latent space: the control expression profile x is transformed into a latent vector s ∈ ℝᵈ, and each perturbation p ∈ P is mapped to a latent embedding zₚ ∈ ℝᵈ, where d ∈ ℕ is the chosen dimensionality of the latent space. The model then applies a latent shift strategy: a learned decoder gφ predicts the final perturbed state by operating on the sum of the basal latent vector and the aggregated perturbation embeddings, i.e., ŷ = gφ(s + ∑p∈P zₚ). Training is performed by minimizing the mean squared error (MSE) between the true expression profile y and the prediction ŷ:
$$mathcal{L}(mathbf{y}, widehat{mathbf{y}}) = frac{1}{n} | mathbf{y} – widehat{mathbf{y}} |_2^2$$
This additive formulation naturally extends to single, double, and higher-order multi-perturbation scenarios of order m := |P| due to its compositional structure. As an alternative, more expressive fusion mechanisms could employ transition functions (s′ = Tψ(s, z)) to model sequential application of perturbations, enabling dynamic tracking of cellular state changes—e.g., ŷ = gφ(Tψ(…Tψ(s, zp₁)…, zpₘ)). To obtain the latent representations s and zₚ (for p ∈ P), TxPert uses two dedicated encoder modules: the basal state encoder and the perturbation encoder, described in detail below.
Basal state model
The basal state encoder captures intrinsic biological properties of the cell—such as its identity, cell-cycle stage, and other baseline phenotypic characteristics—by compressing the control gene expression vector x ∈ ℝⁿ into a compact, low-dimensional embedding s = fbasal(x) ∈ ℝᵈ. Through hyperparameter optimization, a multi-layer perceptron (MLP) was found to yield the best performance for predictions involving unseen perturbations and dual-perturbation combinations (see Supplementary Note 1 and Supplementary Fig. 7 for cross-cell-type results). The MLP provides a direct, deterministic mapping from high-dimensional gene expression data to a fixed-size latent space. Its architecture strikes a balance between simplicity and expressiveness: it is computationally efficient while still capable of capturing the complex nonlinear relationships inherent in transcriptomic data.
Basal information matching and aggregation
A critical challenge in modeling the basal state lies in accurately aligning the control sample with the perturbed target cell. Since experimental conditions—including protocols and platforms—can differ significantly across datasets, we address this by randomly selecting a control sample that shares the same cell line and dataset or experimental protocol as the perturbed condition. We also investigate a more refined strategy called basal state matching, in which the control is chosen to closely match the batch-related metadata (e.g., sequencing batch or library preparation method) of the perturbed sample. Because multiple suitable controls may exist, we randomly draw one from this matched set. Additionally, we explore basal state averaging, where instead of relying on a single control, we compute the mean expression profile across all compatible controls for a given cell line and/or batch. This yields a more robust and stable estimate of the true baseline state. In our experiments, both matching and averaging strategies consistently enhanced model accuracy.
Encoders
In addition to using MLPs that operate directly on raw gene-expression profiles, we also tested embeddings from several state-of-the-art transcriptomics foundation models as alternative inputs for representing the basal state. Specifically, we evaluated scGPT and scVI, both pre-trained on the Joung dataset39. We also considered a simplified variant that omits the basal state encoder entirely, using the raw expression vector x directly as the latent state (i.e., s := x). In this setup, the perturbation decoder learns a Δ-expression vector from the perturbation embeddings, which is then simply added to the control profile: ŷ = x + gφ(∑p∈P zₚ). This resembles the general baseline approach but replaces manually designed heuristics with a learned prediction of expression change. As expected, this configuration performs best in data-limited scenarios—for instance, when predicting perturbation effects in cell lines with no prior exposure to the tested perturbations.
Perturbation model
To learn meaningful representations of genetic perturbations, we leverage graph neural networks (GNNs) that incorporate biological knowledge graphs (KGs) encoding known gene–gene (or gene product–gene product) interactions. The GNN generates a matrix of node embeddings corresponding to each perturbation token: {1, …, N} ↦ Z ∈ ℝ^(N×d), where N ∈ ℕ is the number of perturbations relevant to the task at hand. Each row of Z represents the learned latent encoding of a specific perturbation, so zₚ ∈ ℝᵈ—the p-th row—serves as the perturbation embedding used in the latent shift step. Concretely, each perturbation p ∈ {1, …, N} is initially assigned a random input node embedding hₚ⁰ ∈ ℝ^(d₀), with d₀ ∈ ℕ. These initial embeddings are stacked into an input feature matrix H⁰ ∈ ℝ^(N×d₀). During training, these node features (1) are treated as learnable parameters updated via backpropagation and (2) are further refined through L rounds of message passing in the GNN: H^(ℓ) = LAYERθ_ℓ(H^(ℓ−1)) for 0 ≤ ℓ ≤ L, with the final embedding matrix Z set equal to H^(L). This design enables the model to encode perturbation effects based on their biological context within established interaction networks such as GO19, STRING18, or proprietary knowledge sources.
Real-world knowledge graphs are inherently imperfect: they may include spurious or incorrect interactions (false positives), lack known true connections (false negatives), and can originate from
Our data draws from a wide range of sources, each contributing different—and sometimes contradictory—viewpoints. Working with such intricate data requires carefully selected Graph Neural Network (GNN) architectures tailored to overcome these specific challenges.
To meet these needs, we adopt and refine two core GNN strategies. First, to manage unreliable or noisy connections in the graph, we employ attention-based models like the Graph Attention Network (GAT)40,41. GAT’s key strength lies in its ability to dynamically adjust the importance of neighboring nodes, which helps suppress less trustworthy links—a significant advantage over non-attention methods such as standard graph convolutions42 used in GEARS. Second, to tackle incomplete graph structures and capture long-range biological relationships, we use Graph Transformers (GTs), particularly Exphormer16,17. Unlike traditional GNNs limited to immediate neighbors, Exphormer can attend to distant nodes, enabling it to infer hidden connections and fill structural gaps in the network.
We also found it essential to leverage multiple Knowledge Graphs (KGs) that provide complementary biological insights relevant to our tasks. To harness this diversity, we explore architectures designed for synergistic learning across heterogeneous sources. This includes extending GAT into GAT-Hybrid—which applies node-level attention to weigh information from different KGs—and introducing Exphormer-MG, a provenance-aware variant of Exphormer. Additionally, we develop GAT-MLG, a multilayer extension of GAT that uses a supra-adjacency matrix to seamlessly integrate signals across multiple biological networks.
Supplementary Note 3 offers a thorough technical account of the graph representation learning methods used to encode gene perturbations, the proposed model variants, and the strategies for fusing complementary information from multiple KGs. In our experiments, the GNN-generated embeddings for perturbed genes were combined with a basal (unperturbed) state representation to forecast the resulting gene expression profile. This setup allowed us to systematically compare how different GNN approaches—leveraging attention, flexible connectivity, and multigraph integration—perform when learning from the inherent complexity of biological knowledge graphs.
Training and evaluation framework
Algorithm 1
TxPert training algorithm
Require: Perturbed cells (mathcal{Y}), control cells (mathcal{X}), biological prior graph G = (V, E) with perturbations (mathcal{P} subset V)
Ensure: Minimize mean squared error (MSE) between predicted and actual perturbed cell measurements
1: Randomly initialize input perturbation embeddings ({mathbf{h}_v^0}_{v in V})
2: for each training step do
3: Sample a batch of perturbed cell profiles: ({mathbf{y}_i}_{i=1}^B subset mathcal{Y})
4: Sample matched control cells from the same experimental batches: ({mathbf{x}_i}_{i=1}^B subset mathcal{X})
5: Enhance perturbation embeddings using the graph prior: ({mathbf{z}_v}_{v in V} leftarrow mathtt{GNN}(G, {mathbf{h}_v^0}_{v in V}))
6: for each sample in the batch do
7: Encode control cells into a basal latent representation: bi ← MLPbasal(xi)
8: Identify perturbations (P_i subset mathcal{P}) linked to target yi
9: Fuse control and perturbation embeddings: (widehat{mathbf{z}}_i leftarrow mathtt{COM}(mathbf{b}_i, {mathbf{z}_p : p in P_i}))
10: Decode into predicted perturbed expression profile: (widehat{mathbf{y}}_i leftarrow mathtt{MLP}_{text{dec}}(widehat{mathbf{z}}_i))
11: Compute per-sample loss: (mathcal{L}_i leftarrow mathtt{MSE}(widehat{mathbf{y}}_i, mathbf{y}_i))
12: end for
13: Aggregate batch loss: (mathcal{L} leftarrow sum_{i=1}^B mathcal{L}_i)
14: Backpropagate gradients and update model parameters
15: end for
Data splits
The dataset was partitioned into training, validation, and test sets by grouping samples according to perturbation identity. This ensured that entirely unseen perturbations were held out in both validation and test sets, with target proportions of 0.5625, 0.1875, and 0.25, respectively. For the cross-cell-type task, the test set consisted of a cell type not seen during training; only control cells from this type were included in training, and both seen and unseen perturbations were evaluated within it. As an exception, for the doubles perturbation task on the Norman dataset, we used the predefined data splits from the GEARS framework.
Hyperparameters for each model were tuned using the validation set’s Pearson Δ score. All reported results are based solely on test set performance.
Metric definitions
Unless otherwise noted, all metrics are computed as weighted averages—specifically, the mean of per-perturbation means across all cells receiving that perturbation.
Expression value representations and Δ
When not otherwise specified, gene expression values (mathbf{x} in mathcal{X} cup mathcal{Y}) are processed as log1p-transformed, library-size-normalized counts (with a target library size of 4,000). For a raw count vector (mathbf{x}_{text{raw}} in mathbb{R}^n), the transformation is defined as:
$$mathbf{x} := log left(1 + 4000 cdot frac{mathbf{x}_{text{raw}}}{|mathbf{x}_{text{raw}}|_1} right).$$
An alternative representation, called Δ, centers expression values relative to batch-matched controls. For a perturbed cell expression profile (mathbf{y}_i in mathcal{Y}) from cell line c and batch b, the Δ value is computed as:
$$delta_i := mathbf{y}_i – overline{mathbf{x}}_{c,b},$$
where (overline{mathbf{x}}_{c,b}) denotes the average expression of control cells (mathbf{x} in mathcal{X}) from the same batch b and cell line c.
Pearson Δ
Adapted from the “Pearson correlation (Δ expression)” metric in the GEARS study, Pearson Δ measures the correlation between predicted and observed log fold changes relative to batch-matched control means:
$$text{Pearson}Delta(p) := mathrm{Pearson}(widehat{delta}_p, delta_p),$$
where (widehat{delta}_p) and δp represent the predicted and true Δ expression vectors for perturbation p, respectively.
of the prediction and ground truth, respectively, averaged across replicates of a given perturbation (pin {mathcal{P}}). For clarity, we define this and subsequent metrics for individual perturbations (pin {mathcal{P}}), though equivalent versions can be applied to groups of perturbations (Psubset {mathcal{P}}). The final performance score is obtained by averaging the results over all predicted perturbation effects.
Note that, exclusively for the GEARS model, we directly use the ‘Pearson correlation (Δ expression)’ value as computed in the GEARS codebase. In practice, we verified that the difference between this metric and our own ‘Pearson Δ’ is negligible compared to the differences observed across distinct models.
Retrieval
We employ two variations of the retrieval rank metric. These metrics evaluate how similar a prediction is to the ground truth not in absolute terms, but in comparison to other perturbations. These metrics are identical to the rank average defined in PerturBench10, except they measure similarity with a specific scale: a perfect match scores 1, random performance scores 0.5, and a perfectly anti-correlated prediction scores 0:
$$begin{array}{ll}mathrm{Retrieval}: & =displaystylefrac{1}{N}sum _{pin {mathscr{ mathcal P }}},mathrm{rank},({hat{delta }}_{p}), mathrm{rank},({hat{delta }}_{p}): & =displaystylefrac{1}{N-1}mathop{sum}limits_{{qin {mathscr{ mathcal P }}}atop {qne p}}{{bf{1}}}_{{mathrm{Pearson}({hat{delta }}_{p},{delta }_{p})ge mathrm{Pearson}({hat{delta }}_{p},{delta }_{q})}}.end{array}$$
In the ‘normalized’ retrieval setting, the total number of perturbations (N:=| {mathcal{P}}|) matches the original experimental design. For ‘fast retrieval’, to improve computational efficiency, we use a randomly seeded reference set containing only 100 perturbations, adding the query perturbation p if it is not already included (hence N ∈ {100, 101}). Analogous to Pearson Δ, we report the average performance across all perturbations.
Nonlearned general baseline
To establish a minimum performance threshold, we implement a simple, non-learned baseline model. This model predicts expression profiles based solely on the average values found in the training data. It uses an additive method that combines the following:
The average control expression profile for the specific test cell type.
The average change for that specific perturbation (if it has been seen before) or the overall average change for all perturbations (if it is unseen).
When multiple cell lines are included in the training data, we either calculate a weighted average based on the number of samples per cell line or use the average changes from the most similar cell line (nearest-cell-line baseline). Similarity is determined by the average correlation of shared perturbation Δ values between the test and candidate cell lines.
Consider a multiset of training samples (,mathrm{Train},subset {{mathcal{P}}}_{mathrm{train}}times {{mathcal{C}}}_{mathrm{train}}times {{mathcal{B}}}_{mathrm{train}}) that consists of combinations of perturbation(s), cell line(s), and technical batch effects. A corresponding multiset is defined for the test data. Consider a perturbation p such that (p, cp, bp) ∈ test, with cell line cp and batch effect bp. Implicitly, a mapping for (cp, bp) is linked with a set of control cell profiles in that same context.
If a matching entry exists in the training set: (p, c, b) ∈ train, we use:
$$begin{array}{ll}{widehat{{bf{y}}}}_{(p,{c}_{p},{b}_{p})} & = & {bar{{bf{x}}}}_{({c}_{p},{b}_{p})}+displaystylefrac{1}{| {(q,c,b)in ,mathrm{Train}:q=p}| }mathop{sum }limits_{{(q,c,b)in ,mathrm{Train}}atop{q=p}}{{bf{y}}}_{(p,c,b)}-{bar{{bf{x}}}}_{(c,b)}.end{array}$$
Otherwise, we apply the global average change (Δ) observed across all perturbations in the training set:
$${widehat{{bf{y}}}}_{(p,{c}_{p},{b}_{p})}={bar{{bf{x}}}}_{({c}_{p},{b}_{p})}+frac{1}{| ,mathrm{Train}| }mathop{sum }limits_{(q,c,b)in mathrm{Train}}{{bf{y}}}_{(p,c,b)}-{bar{{bf{x}}}}_{(c,b)}.$$
For multiple perturbations, this baseline first tries to use training samples that contain the exact same combination of perturbations. If unavailable, it splits the combination into individual components. Each component is then added sequentially to the control mean, using a local Δ estimate if one exists, or falling back to the global Δ otherwise.
Estimating Experimental Reproducibility: Split-Half Validation and Sample-Based Extension
Because Perturb-seq is a destructive assay, it is impossible to measure the same cell in both its perturbed and unperturbed states. This means we must evaluate the accuracy of distributional averages rather than individual cells. To estimate the experimental reproducibility, we use a split-half validation method:
For each unique combination of perturbation(s), cell line, and batch, we perform three steps:
-
1.
Randomly divide the test cells into two roughly equal groups.
-
2.
Compute the average expression profile for each group.
-
3.
Calculate the agreement between these two averages using various metrics.
Since the specific split can introduce randomness, we repeat this process across multiple iterations with different random seeds and report the overall average performance. This benchmark represents the experimental reproducibility—a concept analogous to accuracy in other areas of machine learning.
Consider a set of expression profiles ({mathcal{S}}subset {mathcal{Y}}) for a fixed perturbation, cell line, and batch (p, c, b) in the test set:
$$begin{array}{rcl}{{mathcal{S}}}^{{prime} } & subseteq & {mathcal{S}}:,| {{mathcal{S}}}^{{prime} }| approx | {mathcal{S}}| /2 {bar{{mathcal{S}}}}_{1} & = & frac{1}{| {{mathcal{S}}}^{{prime} }| }mathop{sum }limits_{{bf{y}}in {{mathcal{S}}}^{{prime} }}{bf{y}} {bar{{mathcal{S}}}}_{2} & = & frac{1}{| {mathcal{S}}backslash {{mathcal{S}}}^{{prime} }| }mathop{sum }limits_{{bf{y}}in {mathcal{S}}backslash {{mathcal{S}}}^{{prime} }}{bf{y}}.end{array}$$
We then calculate and report:
$$mathrm{Reproduce}(p,c,b)=mathrm{Metric}({bar{{mathcal{S}}}}_{1},{bar{{mathcal{S}}}}_{2}),$$
where the Metric function evaluates the agreement between the two halves, serving as an indicator of inherent biological and technical variability.
Here, metric stands for any one of our evaluation metrics—examples include Pearson Δ, Retrieval, or MSE.
In principle, split-half experimental reproducibility shouldn’t represent a hard upper limit on how well all models can perform during testing, since it works with a different test set—using only half the data for prediction and the other half for evaluation. That said, in practice, it serves as a strong (though still theoretically achievable) benchmark that models should aim to match or surpass.
Because split-half reproducibility tends to underestimate true reproducibility—due to using only half the biological replicates—we also developed a sampling-based method to estimate what reproducibility would look like using a full-size dataset. Here’s how it works: starting from the original count matrix, we compute a gene-level probability distribution (via a multinomial distribution and its maximum-likelihood estimate) for each perturbation within every batch. We then draw samples from these distributions to create two new datasets, each matching the size of the original in terms of total cell count but with resampled (stochastic) counts. These synthetic datasets undergo the same processing as the real data: log1p normalization and selection of highly variable genes (HVGs), after which we calculate experiment reproducibility just as before. For a side-by-side view of split-half versus sampling-based reproducibility, see Supplementary Table 4.
Data
Perturb-seq data sources
We test our method on multiple datasets, including CRISPRi (gene knockdown) experiments targeting ~2,000 essential genes in K562 and RPE1 cell lines from an earlier study13 (also used by GEARS4), as well as similar CRISPRi screens in Jurkat and HEPG2 lines from another prior work24. We also incorporate the Norman15 dataset, which includes 94 single and 110 double CRISPRa (gene overexpression) perturbations in K562 cells.
Graphs: sourcing and processing
The graphs we use to provide inductive bias fall into two broad categories: (1) expert-curated biological knowledge bases and (2) systematic, large-scale perturbation screens.
Curated graphs in category 1 include the GO graph (originally employed in GEARS), where nodes are linked based on high Jaccard similarity between their GO terms19, along with the STRING18 and Reactome43 interaction networks.
Graphs in category 2 come from comprehensive perturbation datasets like DepMap44 and Perturb-seq45. These resources map genetic perturbations to either morphological or transcriptomic changes, offering rich insights into how cells respond to genetic interference. To turn these datasets into graphs, we represent genes and cell lines as embeddings in a high-dimensional space and analyze their relationships computationally.
To build these graphs, we first calculate pairwise cosine similarity between every pair of genes (gi, gj) using their aggregated embeddings ({{bf{x}}}_{{g}_{i}}) and ({{bf{x}}}_{{g}_{j}}). The cosine similarity formula is:
$$
begin{array}{rcl}
text{cosine similarity}({{bf{x}}}_{{g}_{i}},{{bf{x}}}_{{g}_{j}}) & = & frac{{{bf{x}}}_{{g}_{i}} cdot {{bf{x}}}_{{g}_{j}}}{parallel {{bf{x}}}_{{g}_{i}} parallel parallel {{bf{x}}}_{{g}_{j}} parallel } \
& = & frac{sum_{k=1}^{n} {{bf{x}}}_{{g}_{i}k} {{bf{x}}}_{{g}_{j}k}}{sqrt{sum_{k=1}^{n} ({{bf{x}}}_{{g}_{i}k})^2} cdot sqrt{sum_{k=1}^{n} ({{bf{x}}}_{{g}_{j}k})^2}}
end{array}
$$
Where:
({{bf{x}}}_{{g}_{i}} cdot {{bf{x}}}_{{g}_{j}}) is the dot product of the two vectors.
parallel {{bf{x}}}_{{g}_{i}} parallel and parallel {{bf{x}}}_{{g}_{j}} parallel denote the Euclidean lengths (magnitudes) of the embedding vectors for genes gi and gj, respectively.
{{bf{x}}}_{{g}_{i}k} and {{bf{x}}}_{{g}_{j}k} are the k-th components of those embedding vectors.
We take the absolute value of cosine similarities because a strongly negative cosine value doesn’t inherently mean a weaker biological relationship—it shouldn’t directly translate to a negative edge weight in the graph.
We also leverage proprietary internal data from genome-wide perturbation screens, where we assess perturbation similarity using both imaging (microscopy) and transcriptomic profiling across multiple cell types.
Graph filtering settings were tuned through empirical testing. The best-performing setup involved keeping only the top 1% of edges by absolute weight for screen-derived graphs. For all other graph types, we additionally limited each target node to no more than 20 incoming edges. When edges were undirected, their direction was assigned arbitrarily.
Data understanding
Additional methodological details relevant to specific analyses are provided below.
Pharos knowledge rank
The Pharos database compiles various metrics that reflect how extensively studied or well-characterized individual genes are26. From this, we compute a composite knowledge rank by averaging the ranks of two scores: the PubMed mention count and the negative log novelty score from Pharos. This combined rank allows us to group genes into knowledge tiers and assess whether model performance varies by gene familiarity. Knowledge levels 0 through 3 correspond to the following ranges of this Pharos rank:
Level 0 (least characterized): 0–0.2
Level 1: 0.2–0.4
Level 2: 0.4–0.6
Level 3: 0.6–0.8
Level 3 (most characterized): 0.8–1.0
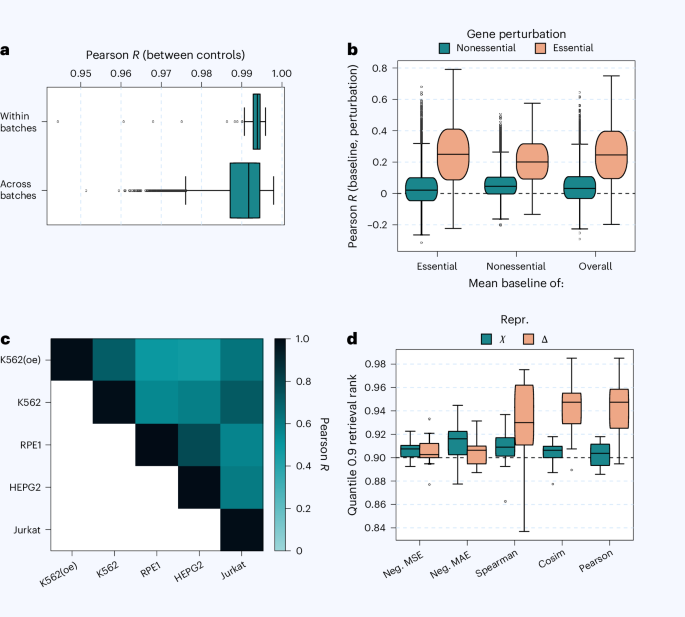
Within versus across
When examining correlations between control and mean baseline conditions, we distinguish between “within-context” and “across-context” correlations. All samples are first split randomly into two non-overlapping halves, A and B. Within-context correlation compares A and B within the same experimental context, while across-context correlation evaluates a randomly
To prevent samples from one batch from being improperly merged with samples from a different batch, entire batches were kept intact rather than divided—except in specific cases shown in Fig. 1a. In those across-batch comparisons, full batches were combined without splitting to ensure a cautious, conservative measure of variability between batches. For comparisons involving individual control cells, the data were split and then recombined. As for the baseline measurements, the change values (δ) from replicate perturbant cells were first combined (pre-aggregated) and then divided into two parts that contained distinct (non-overlapping) perturbations.
Functional enrichment
To create a clear biological summary of the actual gene expression shifts captured by our average baseline, we began by computing a combined “meta” baseline. This overall baseline was derived by averaging across all available cell types and datasets, using only genes that were present (expressed) across all sources. We then classified genes that showed a positive change greater than 0.05 (δ > 0.05) as upregulated, and those with a negative change less than −0.05 (δ < −0.05) as downregulated. Using these two gene sets, we performed functional enrichment analysis separately for each group, comparing them against the full set of genes in the shared dataset. This analysis was carried out using the GOATOOLS software package46.
Metric selection through retrieval
To avoid overwhelming readers with numerous similar or slightly conflicting performance measures, we conducted an initial “test of the test metrics” to identify the most informative ones. We built upon an evaluation approach introduced in earlier research12, adapting it to suit our specific data and goals. In essence, this method evaluates whether a given data representation and its associated metric can preserve enough detail to correctly identify and distinguish different perturbations when retrieving them across different biological contexts. For our purposes, we adjusted the method to operate directly on individual single-cell expression profiles—since these are both the input and output of our model—rather than on aggregated data. We applied this retrieval test across a core set of essential perturbations in four key cell types: K562, RPE1, HEPG2, and Jurkat. For each retrieval test, instead of combining cells, we randomly selected one representative cell per perturbation. We repeated this process three times for every possible pairing of cell types, resulting in a total of n = 3 × 6 = 18 independent estimates per perturbation. We report results at the 0.9 quantile to emphasize strongly active perturbants, though similar trends were seen at other thresholds.
It’s important to note that focusing on single cells meant we could not use the representation recommended in the prior study12, which relied on signed P values. To ensure this limitation didn’t compromise our findings, we ran a preliminary check using the exact setup from that earlier work. We were only able to replicate their high performance when making assumptions that would restrict the flexibility of our own data and training pipeline. Specifically, we found that the strong performance of the signed P value depended on fitting a single global model across all experimental contexts—which assumes a uniform estimate of gene-level variability everywhere.
While we prioritized representations and metrics widely used in perturbation-based transcriptomics studies, we recognize that count-based approaches were not included in this evaluation.
General
All model development and data analysis were conducted using Python 3.12 and PyTorch 2.6.0. Visualizations were generated using seaborn 0.13.2 and Matplotlib 3.10.8. In box-and-whisker plots, the central box spans the 25th to 75th percentiles (the interquartile range), with a line marking the median. Whiskers extend to the most distant data point within 1.5 times the interquartile range from the box edges.
Reporting summary
Additional details about the experimental design and methodology can be found in the Nature Portfolio Reporting Summary associated with this article.