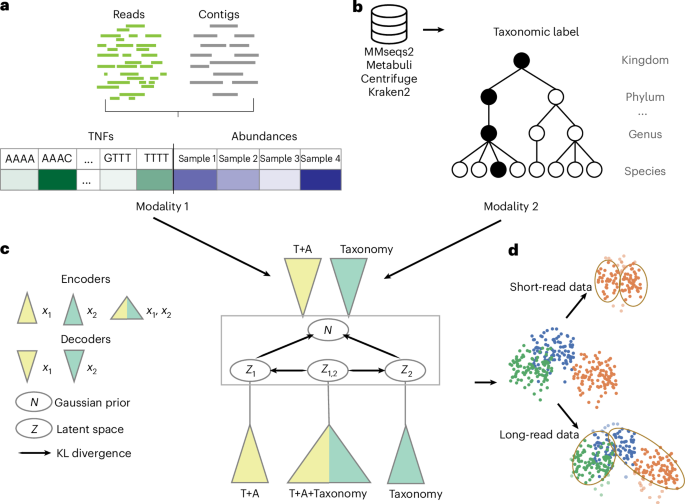
Bimodal VAE
The VAE is a generative mannequin performing variational inference over the latent variable z. The mannequin is formally outlined as p(x,z) = p(z)p(x|z). The intractable posterior q(z|x) and the conditional distribution p(x|z) are approximated by neural networks utilizing the ELBO-loss operate:
$$beginxx=q(z_zleft[log pleft(right)right]-mathrmzleft(qleft(proper)zzleft(0,Iright)proper)endz$$
(1)
The bimodal VAE extends the essential VAE by permitting coaching and inference on the dataset the place (1) the enter consists of two modalities and (2) a modality could be lacking for a number of samples. Thus, discover that, whereas VAMB is skilled on each TNFs and abundances, we don’t outline it as bimodal for the aim of this abstract, as each TNFs and abundances are current for all samples and could be transformed into one modality by concatenating the corresponding enter vectors.
Whereas the VAE approximates the posterior q(z|x) with a neural community encoder that takes x as an enter, the bimodal VAE extends this method by modeling q(z|x1,x2), q1(z|x1) and q2(z|x2), which exchange the one q(z|x). There are two decoders approximating distributions p(x1|z) and p(x2|z). Multimodal VAEs differ in (1) the best way they approximate q(z|x1,x2), q1(z|x1) and q2(z|x2) by neural networks and/or (2) the construction of the loss operate.
TaxVAMB implements the VAEVAE50 mannequin from the bimodal VAE household, which fashions q(z|x1,x2), q1(z|x1) and q2(z|x2) by corresponding neural networks. The next ELBO-like loss L is minimized:
$$beginzz_zz_[z_[log _(z_z)+log z_(z_z)] -mathrm(q(_z,z_)Vert p(z_z))-mathrmz(q(z_,z_)Vert p(z_z))] +{{mathbbz}}_{_{textz}(_z)}[{{mathbb}}_,z_zright)[log _{1}({x}_{1}z)]-mathrm{KL}(q({x}_{1})Vert p(z))] +{{mathbb{E}}}_{{p}_{textual content{paired}}({x}_{2})}[{{mathbb{E}}}_{q(z| {x}_{2})}[log {p}_{2}({x}_{2}z)]-mathrm{KL}(q({x}_{2})Vert p(z))] { {mathcal L} }_{1}{=}{{mathbb{E}}}_{{p}_{textual content{unpaired}}({x}_{1})}[{{mathbb{E}}}_{q(z| {x}_{1})}[log {p}_{1}({x}_{1}z)]-mathrm{KL}(q({x}_{1})Vert p(z))] { {mathcal L} }_{2}{=}{{mathbb{E}}}_{{p}_{textual content{unpaired}}({x}_{2})}[{{mathbb{E}}}_{q(z| {x}_{2})}[log {p}_{2}({x}_{2}z)]-mathrm{KL}(q({x}_{2})Vert p(z))] {mathcal L} {=}{ {mathcal L} }_{textual content{paired}}+{{mathcal{L}}}_{1}+{{mathcal{L}}}_{2}finish{array}$$
with DKL (p(x)||q(x)) being the Kullback–Leibler divergence between two likelihood distributions p(x) and q(x), outlined as:
$$start{array}{l}mathrm{KL}left(pleft(xright)mathrmqleft(xright)proper)={int }_{-infty }^{infty }pleft(xright)log left(frac{pleft(xright)}{qleft(xright)}proper){rm{d}}xend{array}$$
(2)
The coaching process consists of establishing the dataset with paired and unpaired samples. Let C be a listing of all contigs. Three copies of C, denoted as Cpaired, C1 and C2, are independently shuffled. The paired samples are ordered tuples (x1,x2) the place x1 is a concatenation of TNF vector and abundance vector (the enter of VAMB) and x2 is a taxonomy label vector and x1 and x2 correspond to the identical contig from the set Cpaired. An unpaired TNFs and abundances vector x1 corresponds to a contig from the record C1. An unpaired taxonomy label x2 corresponds to a contig from the record C2. The ahead cross follows the steps from Algorithm 1.
Algorithm 1Loss computation (ahead cross)
Require: Paired pattern (({x}_{1},{x}_{2})), unpaired pattern ({x{prime} }_{1}), unpaired pattern ({x{prime} }_{2})
1: (z{prime} sim qleft({x}_{1},{x}_{2}proper))
2: ({z}_{{x}_{1}}sim {q}_{1}left({x}_{1}proper))
3: ({z}_{{x}_{2}}sim {q}_{2}left({x}_{2}proper))
4: ({d}_{1}={D}_{mathrm{KL}}(q(z{prime} |{x}_{1},{x}_{2})parallel {q}_{1}({z}_{{x}_{1}}|{x}_{1}))+{D}_{mathrm{KL}}({q}_{1}({z}_{{x}_{1}}|{x}_{1})parallel p(z)))
5: ({d}_{2}={D}_{mathrm{KL}}(q(z{prime} |{x}_{1},{x}_{2})parallel {q}_{2}({z}_{{x}_{2}}|{x}_{2}))+{D}_{mathrm{KL}}({q}_{2}({z}_{{x}_{2}}|{x}_{2})parallel p(z)))
6: ({L}_{mathrm{paired}}=log {p}_{1}({x}_{1}z)+log {p}_{2}({x}_{2}z)+log {p}_{1}({x}_{1}|{z}_{{x}_{1}})) (+log {p}_{2}({x}_{2}|{z}_{{x}_{2}})+{d}_{1}+{d}_{2})
7: ({L}_{{x}_{1}}=log {p}_{1}({x{prime} }_{1}|{z}_{{x}_{1}})+{D}_{mathrm{KL}}({q}_{1}({z}_{{x}_{1}}|{x{prime} }_{1})parallel p(z)))
8: ({L}_{{x}_{2}}=log {p}_{2}({x{prime} }_{2}|{z}_{{x}_{2}})+{D}_{mathrm{KL}}({q}_{2}({z}_{{x}_{2}}|{x{prime} }_{2})parallel p(z)))
9: (L={L}_{mathrm{paired}}+{L}_{{x}_{1}}+{L}_{{x}_{2}})
Information preprocessing
The workflow of preprocessing the info is identical as in Taxometer (model 5.0.4)42 and VAMB (model 5.0.4)11. The artificial brief paired-end reads from every pattern have been aligned utilizing bwa-mem (model 0.7.15)73. BAM information have been sorted utilizing SAMtools (model 1.14)74. Contigs ≤ 2,000 bp have been eliminated for every dataset. The long-read datasets have been each sequenced utilizing Pacific Biosciences HiFi expertise. We assembled every pattern utilizing hifiasm-meta (model 0.3.1)3, mapped reads utilizing minimap2 (model 2.24)75 with the ‘-ax map-hifi’ setting after which continued with the identical workflow as with the brief reads.
Abundances and TNFs
The workflow of computing abundances and TNFs is identical as in Taxometer (model 5.0.4)42 and VAMB (model 5.0.4)11. Computation of abundances and TNFs was carried out utilizing the VAMB metagenome binning device11. To find out TNFs, tetramer frequencies of nonambiguous bases have been calculated for every contig, projected right into a 103-dimensional orthonormal area and normalized by z-scaling every tetranucleotide throughout the contigs. To find out the abundances of every pattern, we used pycoverm (model 0.6.0; The abundances have been first normalized inside the pattern by the full variety of mapped reads after which throughout samples to sum to 1. To find out absolute abundance, the sum of abundances for a contig was taken earlier than the normalization throughout samples. The dimensionality of the function desk was then Nc × (103 + Ns + 1), the place Nc is the variety of contigs and Ns is the variety of samples.
Community structure and hyperparameters
The encoder architectures for the concatenated vector of abundances and TNFs is identical as in Taxometer (model 5.0.4)42 and VAMB (model 5.0.4)11. The enter vector of dimensionality Nc × (103 + Ns + 1) was handed by 4 totally related layers ((103 + Ns + 1) × 512, 512 × 512, 512 × 512, 512 × 512) with leaky ReLU activation operate (detrimental slope 0.01), every utilizing batch normalization (ϵ 1 × 10−5, momentum 0.1) and dropout (P = 0.2).
The encoder community for the taxonomy labels had the enter dimensions of Nl, the place Nl is the variety of leaves within the taxonomic tree. The enter vector was handed by 4 totally related layers (Nl × 512, 512 × 512, 512 × 512, 512 × 512) with leaky ReLU activation operate (detrimental slope: 0.01), every utilizing batch normalization (ϵ 1 × 10−5, momentum 0.1) and dropout (P = 0.2).
The encoder community for the concatenation of the 2 modalities had the enter dimensions of (103 + Ns + 1) + Nl, the place Ns is the variety of samples and Nl is the variety of leaves within the taxonomic tree. The enter vector was handed by 4 totally related layers ((103 + Ns + 1) × 512, 512 × 512, 512 × 512, 512 × 512) with leaky ReLU activation operate (detrimental slope: 0.01), every utilizing batch normalization (ϵ 1 × 10−5, momentum 0.1) and dropout (P = 0.2).
The bimodal VAE has two decoder networks, one for every modality. Each of them observe the identical architectures because the corresponding encoders, with the enter vector having the dimensionality of the latent area and the output having the dimensionality of the corresponding modality.
For brief-read datasets, the community was skilled for 300 epochs with batch measurement 256 and latent area dimensionality 32. For long-read datasets, the community was skilled for 1,000 epochs with batch measurement 1,024 and latent area dimensionality 64. All fashions have been utilizing the Adam optimizer with studying charges set by D-Adaptation76. The mannequin was applied utilizing PyTorch (model 1.13.1)77 and CUDA (model 11.7.99) was used when working on a V100 GPU.
Hierarchical loss
The hierarchical loss is identical as in Taxometer (model 5.0.4)42. A phylogenetic tree was constructed for every dataset from the taxonomy classifier annotations for the set of contigs. Thus, the ensuing taxonomy tree T was a subgraph of a full taxonomy and the area of potential predictions was restricted to the taxonomic identities that appeared within the search outcomes. For the above experiments, we used a flat softmax loss. Let Nl be the variety of leaves within the tree T. The likelihoods of leaf nodes of the taxonomy tree have been obtained from the softmax over the community output layer with dimensionality 1 × Nl. The probability of an inner node was then a sum of likelihoods of its kids and computed recursively bottom-up. The mannequin output was a vector of likelihoods for every potential label. For the backpropagation, the detrimental log-likelihood loss was computed for all of the ancestors of the true node and the true node itself. Predictions have been made for all taxonomic ranges and, for every stage, the node descendant with the very best probability was chosen. If no node descendant had probability > 0.5, the predictions from this stage and the degrees under weren’t included within the output.
Taxonomic classifiers
We obtained the taxonomic annotations for contigs of all 9 short-read and two long-read datasets from MMseqs2 (model 17.b804f)33, Metabuli (model 1.1.0)78, Centrifuge (model 1.0.4.2)35 and Kraken2 (model 2.1.3)32. For MMseqs2, we used the mmseqs taxonomy command. For Metabuli, we used the metabuli classify command with ‘–seq-mode 1’ flag. For Centrifuge, we used the centrifuge command with ‘-k 1’ flag. For Kraken2, we used the kraken command with ‘–minimum-hit-groups 3’ flag. MMseqs2 and Metabuli have been configured to make use of GTDB model 220 because the reference database. Centrifuge and Kraken2 have been configured to make use of NCBI identifiers, launch 229. All of the taxonomic annotations have been first refined with Taxometer42 (model 5.0.4) with the default parameters (epochs 100, batch measurement 1,024). For datasets in Figs. 4 and 5, the MMseqs2 classifier configured with GTDB was used for all datasets; for the wheat phyllosphere dataset, we used Kraken2 (model 2.1.3) configured with NCBI.
Benchmarked binners
The Metabat (model 2.17-66-ga512006) ‘metabat’ command with the default parameters was used. Metadecoder (model 1.0.19) ‘coverage’, ‘seed’ and ‘cluster’ instructions have been used as described on GitHub ( The Comebin (model 1.0.4) ‘run comebin.sh’ command with default parameters was used. ‘Comebin (single)’ signifies the usage of Comebin in a single-sample mode. The SemiBin2 (model 2.2.1) ‘multi_easy_bin’ command was used with the flags ‘–engine gpu –separator C -t 20 –write-pre-reclustering-bins and –self-supervised’. VAMB, AVAMB and TaxVAMB have been run as part of the VAMB codebase (model 5.0.4), with the corresponding instructions ‘vamb bin default’, ‘vamb bin avamb’ and ‘vamb bin taxvamb’. The workflows can be found on GitHub ( The log information for the failed runs are additionally obtainable on GitHub (https://github.com/RasmussenLab/TaxVamb-Benchmarks/tree/main/log_files_for_crashed_runs).
Reclustering utilizing SCGs
Quick-read and long-read reclustering algorithms that used SCGs have been the identical as launched in SemiBin2 (ref. 21). The code was tailored from the SemiBin2 codebase ( and for the TaxVAMB codebase ( TaxVAMB used the identical 107 single-copy marker genes as utilized in SemiBin2 to estimate the completeness, contamination and F1 rating of each bin. Completeness for every bin was calculated as (frac{N}{107}), contamination was calculated as (frac{G-N}{G}) and F1 rating was calculated as (frac{2times mathrm{completeness}instances (1-mathrm{contamination})}{mathrm{completeness}+(1-mathrm{contamination})}), the place 107 is the variety of completely different SCGs in a bin and G is the full variety of sequences matching any SCG.
For the short-read datasets, ok-means-based reclustering of TaxVAMB and VAMB clusters was carried out. Bins the place a couple of marker gene of the identical form was current have been reclustered with the weighted ok-means technique utilizing the contigs containing the repeated marker gene because the preliminary centroids. This resulted in bins with lowered contamination. For the long-read datasets, the DBSCAN algorithm from Python library scipy (model 1.10.0) was used to carry out the clustering from scratch (the earlier clusters, made by TaxVAMB/VAMB, weren’t used). As in SemiBin2, DBSCAN was run with ϵ values of 0.01, 0.05, 0.1, 0.15, 0.2, 0.25, 0.3, 0.35, 0.4, 0.45, 0.5 and 0.55. From all ensuing bins, the most effective one (F1 rating) was recursively chosen and its contigs have been faraway from all of the remaining bins, after which the choice of the most effective bin was repeated. This was repeated till no extra bins fulfilled the factors for minimal high quality (completeness > 90%, contamination < 5%). One change that was made within the TaxVAMB long-read reclustering was that it carried out the described process per set of contigs assigned to the identical genus by the Taxometer refinements of the offered taxonomic annotations.
CAMI2 benchmarks
For brief-read benchmarking, we used 5 CAMI2 datasets: Airways (ten samples), Oral (ten samples), Pores and skin (ten samples), Gastrointestinal (ten samples) and Urogenital (9 samples), the assemblies of which have been sample-specific. The CAMI2 datasets include the next variety of genomes with nonzero abundance: Oral, 799; Pores and skin, 610; Urogenital, 254; Gastrointestinal, 268; Airways, 828. The distinctive variety of genomes per pattern is listed in Supplementary Desk 5. We benchmarked the next binners on the artificial CAMI2 toy human microbiome dataset: Metabat10, MetaDecoder79, COMEBin22, SemiBin2 (ref. 21), AVAMB28 and VAMB11. We used taxonomic labels from 4 taxonomic classifiers as an enter to TaxVAMB: MMSeqs2 (ref. 33), Metabuli77, Kraken2 (ref. 35) and Centrifuge78. AVAMB, VAMB and TaxVAMB bins have been postprocessed with reclustering utilizing SCGs. We used the variety of HQ bins and assemblies estimated utilizing BinBencher (model 0.3.0)54 as a metric. For the MAG taxonomic annotation experiment, we used CheckM2 (model 1.0.2)40. We benchmarked utilizing BinBencher (model 0.3.0)54 in opposition to a reference computed from the CAMI2 floor reality. The metrics used have been the numbers of HQ (outlined as recall ≥ 0.9, precision ≥ 0.95) assemblies or genomes. As outlined within the BinBencher paper, precision was counted because the variety of true optimistic mapping positions for every genome–bin pair, divided by the full variety of positions in a bin. For genomes, the recall was counted relative to the total size of the genome from which the reads have been simulated from, whereas, when counting assemblies, the recall was relative to the assembled a part of these genomes (that’s, the a part of the genomes lined by a contig that was used as enter to the binner). The variety of HQ genomes displays the MAG high quality relative to the underlying organic organism and, thus, relies upon extra on limitations of the dataset, whereas the meeting metric might higher mirror the methodological good points from utilizing a special algorithm.
Quick-read actual knowledge benchmarks
For benchmarking utilizing actual short-read datasets, we used the next: sea water with 5 samples80, bee hives with 18 samples81, forest soil with 12 samples82, rhizosphere with ten samples83, human saliva with 15 samples84 and vaginal microbiome with ten samples85. We assembled every pattern utilizing metaSPAdes (model 4.2.6)86 and mapped reads utilizing minimap2 (model 2.24)87 with the ‘-ax sr’ setting. We used taxonomic labels from 4 taxonomic classifiers as an enter to TaxVAMB: MMSeqs2 (ref. 33), Metabuli78, Kraken2 (ref. 32) and Centrifuge35. MMSeqs2 was evaluated with three databases: GTDB, TrEMBL88 (January 2025 launch) and Kalmari89 (model 3.7). For evaluating the standard (completeness and contamination) of the ensuing MAGs, we used CheckM2 (model 1.0.2)40. For detecting chimeric genomes, we ran GUNC (model 1.0.61)55 utilizing the ‘gunc run’ command. The numbers of sequencing reads for every dataset and pattern are listed in Supplementary Desk 6.
Lengthy-read benchmarks
For long-read benchmarking, we used a human intestine microbiome dataset with 4 samples and a dataset from anaerobic digester sludge with three samples90, each sequenced utilizing Pacific Biosciences HiFi expertise. We assembled every pattern utilizing hifiasm-meta (model 0.3.1)3, mapped reads utilizing minimap2 (model 2.24)87 with the ‘-ax map-hifi’ setting and, from there, proceeded as with the brief reads. For evaluating the standard (completeness and contamination) of the ensuing MAGs, we used CheckM2 (model 1.0.2)40. For detecting chimeric genomes, we ran GUNC (model 1.0.61)55 utilizing the ‘gunc run’ command.
Multisample scaling
For the experiment that evaluated the variety of bins given a special variety of samples, we used a short-read human intestine dataset with 1,000 samples from Almeida et al.58, in addition to our personal wheat phyllosphere dataset with 211 samples. For every dataset, we cut up all of the samples into three units of chunks: (1) chunks of 100 samples; (2) chunks of ten samples; and (3) chunks of 1 pattern. Every chunk was handled as an unbiased dataset. We then summed the ensuing variety of HQ bins inside every set of chunks. Taxonomic annotations have been carried out with MMseqs2 for the human intestine dataset from Almeida et al. and with Kraken2 for the wheat phyllosphere dataset.
Taxonomic annotation validation: ok-fold analysis
All contigs in a dataset that have been annotated by a classifier have been randomly divided into 5 folds. Taxometer was then skilled 5 instances, every time utilizing one fold as a validation set and the remaining 4 folds because the coaching set. Predictions have been generated for the 5 validation units after coaching on the remaining folds. The 5 validation units have been then concatenated to reconstruct the total dataset. This ensured that each contig acquired a prediction that was made with out prior data of its classifier annotation.
The analysis metric was outlined because the fraction of appropriately predicted contigs on the area stage (Micro organism, Archea, and so forth.) over all contigs, whereas accounting for the truth that some ground-truth annotations could also be lacking. In different phrases, the rating displays what number of predictions match the obtainable ground-truth annotations, normalized by the full variety of contigs within the dataset.(mathrm{Accuracy}=frac{mathrm{No}.,mathrm{of},mathrm{right},mathrm{predictions},left(mathrm{the place},mathrm{floor},mathrm{reality},mathrm{exists}proper)}{mathrm{complete},mathrm{quantity},mathrm{of},mathrm{contigs},mathrm{in},mathrm{dataset}}) the place an accurate prediction was when the Taxometer output matched the ground-truth classifier annotation. If no ground-truth annotation was obtainable for a contig, the contig was excluded from the numerator (as correctness can’t be decided) however nonetheless included within the denominator to mirror the truth that lacking values exist within the knowledge. This metric, thus, gives the general share of right predictions throughout the dataset. The command could be accessed within the codebase as vamb taxonomy_benchmark.
Bin annotations for CAMI2 MAGs
For taxonomic classification analysis in Supplementary Figs. 9 and 10, we used Kraken2 configured with the NCBI database and in contrast its efficiency to GTDBtk, which offered GTDB annotations. Somewhat than immediately evaluating these two annotation units, we evaluated each in opposition to ground-truth annotations from CAMI2 datasets. The unique ground-truth taxonomy annotations have been offered as NCBI identifiers as a part of the CAMI2 problem dataset. To determine ground-truth GTDB annotations for the CAMI2 datasets, we ran GTDBtk on the offered CAMI2 ground-truth genomes, acquiring full annotations right down to the species stage. We manually verified a number of genomes to make sure consistency with NCBI annotations. On condition that CAMI2 genomes are a part of public databases, now we have excessive confidence within the high quality of the GTDBtk annotations and, due to this fact, used them as floor reality. On this experimental design, Kraken2 annotations have been in comparison with the bottom reality offered by CAMI2, whereas GTDBtk bin annotations have been in contrast with GTDBtk ground-truth genome annotations.
Human intestine (irritable bowel syndrome) dataset: pattern assortment and processing
Human fecal samples have been collected from 4 wholesome people and 20 individuals with irritable bowel syndrome. In complete, 1–5 samples have been collected from every particular person, yielding a complete of 70 samples. DNA was extracted from fecal samples utilizing a bead-beat micro AX gravity package (A&A Biotechnology) in line with the producer’s directions and the extracts have been additional purified utilizing phenol–chloroform extraction and ethanol precipitation in line with a longtime protocol91. Samples have been sequenced by NovoGene, who ready PCR-based libraries and generated 150-nt paired-end sequencing knowledge on the NovaSeq 6000 platform. Sequencing reads have been quality-controlled and adaptor-trimmed utilizing trim_galore (model 1.15), which used cutadapt (model 3.6.9)92. The default high quality threshold (Phred rating: 20) was used however an additional 16 and 6 nt have been trimmed from the 5′ and three′ ends of the reads, respectively, as this setting was discovered to yield higher contiguity in assemblies in some benchmarking runs. BMtagger (model 1.1.0)93 was then used to take away doubtlessly human reads from the sequencing knowledge utilizing GRCh38.p13 as a reference database. Meeting was carried out utilizing SPAdes (model 3.15.4)94.
Wheat phyllosphere dataset: pattern assortment and processing
A complete of 24 subject plots of Triticum aestivum have been sampled by amassing composite samples of 30 flag leaves 9 instances between June 7 and July 14, 2022, at a subject trial in Ringsted, Denmark. The experimental design included three wheat cultivars, 4 replicates and two therapies, which have been unsprayed and sprayed with a fungicide. The samples have been washed in 100 ml of wash answer (0.9% NaCl + 0.05% Tween-80), vigorously shaken for two min, sonicated for two min, vigorously shaken once more for two min, filtered (10 µm) and centrifuged (4,000g, 15 min); the pellet resuspended in 1 ml of 1× PBS and saved at −20 °C till DNA extraction utilizing the FastDNA SPIN package (MP Biomedicals) for soil in line with directions, eluting in 100 µl of DES. DNA libraries have been construct utilizing the Illumina Nextera XT package (Illumina) however samples with <0.1 ng µl−1 DNA have been constructed with a onefold-diluted amplicon tagment combine, 20 PCR cycles and a better ratio of AMPure XP beads (Beckman Coulter)95. Libraries have been sequenced utilizing Illumina paired-end (2 × 150 bp) expertise (NovaSeq 6000 S4 model 1.5).
Wheat phyllosphere dataset: knowledge evaluation
Uncooked sequencing reads have been filtered utilizing fastp (model 023.2)96 with the choice ‘–trim_tail 1 –cut_tail –trim_poly_g –dedup –length_required 80’. High quality management of the filtered reads was assessed utilizing MultiQC (model 1.12)97. To take away reads originating from wheat or potential human contamination, the reads have been mapped to the reference sequences GCF_018294505.1, MG958554.1 and GCF_000001405.40 (GRCh38.p14). Mapping was carried out utilizing Bowtie (model 2.5.3)98. Paired reads the place each mates have been unmapped have been extracted utilizing SAMtools (model 1.18)73 with the ‘fastq -f 13’ possibility. Metagenomic assemblies have been generated for every pattern utilizing SPAdes (model 3.15.4)94 with the ‘–meta -k 21,33,55,77,99’ possibility. Meeting statistics have been computed utilizing QUAST (model 5.2.0)99. MAGs have been created with TaxVAMB utilizing Kraken2 taxonomic annotations based mostly on NCBI. The MAGs, for which the vast majority of contigs have been annotated as Eukaryotes, have been examined for completeness with BUSCO (model 5.8.2)67. Moreover, to construct taxonomic bushes, MAGs have been assigned the taxonomy utilizing GTDBtk (model 2.4.0) configured with the GTDB database model 220. Taxonomic bushes have been constructed utilizing ggtree (model 3.19)100, tidytree (model 0.4.6) and treeio R (model 4.4.1) libraries. A two-sample Mann–Whitney U-test was carried out on P. agglomerans abundances by splitting the samples into two teams: 143 samples from the sooner days (June 7, 10, 14, 17 and 21, 2022) and 103 samples from the later days (June 28 and July 4, 7 and 14, 2022) utilizing scipy (model 1.10.0).
Reporting abstract
Additional info on analysis design is offered within the Nature Portfolio Reporting Abstract linked to this text.