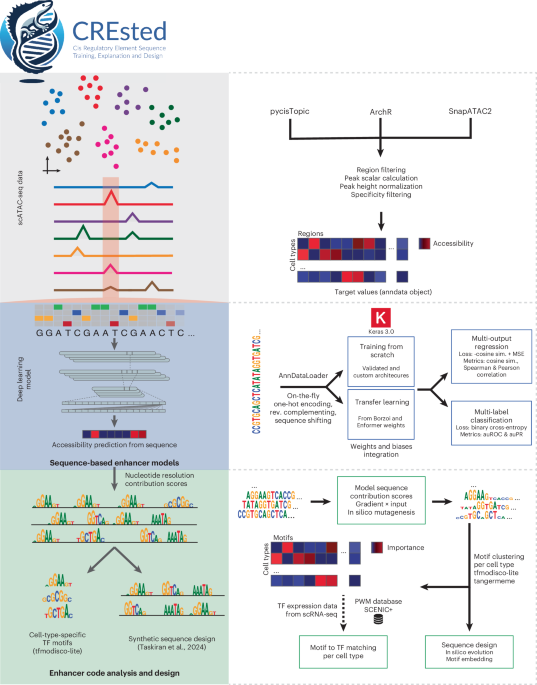
CREsted workflow
CREsted (RRID: SCR_026617) consists of 4 essential modules: knowledge preprocessing, mannequin coaching, enhancer code evaluation and enhancer design. CREsted could be discovered at and https://github.com/aertslab/CREsted/.
Knowledge preprocessing
CREsted preprocessing has two essential modes: cell-type-specific peak-height modeling and matter modeling34,36 (RRID: SCR_026618).
Peak-height preprocessing
A peak-height matrix is generated over consensus peaks utilizing CPM-normalized pseudobulk BigWig tracks per cell kind utilizing pybigtools79 (RRID: SCR_026627), producing the max, imply, sum or logarithm of the sum of both minimize websites or protection per cell kind. Optionally, peak widths could be resized whereas guaranteeing that they keep inside chromosome boundaries. By default, a peak width of two,114 bp is used and the height top is calculated from the middle 1,000 bp. The matrix is saved in an ‘anndata’ object32,80 (RRID: SCR_018209).
Peak normalization
A min–max normalization is utilized by utilizing cell-type-specific scaling elements based mostly on constitutive peaks16. Constitutive peaks are robotically recognized as these with excessive accessibility in a cell kind (high 1% by default) which might be nonspecific (({rm{Giniindex}} < {mu }_{{rm{Giniindex}}-{rm{s}}.{{rm{d}}.}_{{rm{Giniindex}}}}); ({mu }_{{rm{Giniindex}}}) and ({{rm{s}}.{rm{d}}.}_{{rm{Giniindex}}}) are the typical Gini index and customary deviation of the Gini index throughout all peaks, respectively) and the typical values of these peaks, per cell kind, are used as cell-type-specific scaling elements (Supplementary Observe 1).
Matter modeling preprocessing
Both a binary matrix (matter classification) or a likelihood matrix is generated over consensus peaks for every matter containing both a binary label indicating areas belonging to matters or the likelihood of every area for every matter. This matrix is saved in an anndata32,80 (RRID: SCR_018209) object.
Dataset splitting
Areas are break up into coaching, validation and check units both based mostly on an outlined chromosomal break up or by randomly shuffling areas following predefined proportions.
Mannequin coaching
CREsted implements coaching of multi-label fashions—predicting accessibility in a number of cell sorts from one sequence—from scratch, in each classification and regression settings, and by switch studying from Enformer10 (RRID: SCR_024805) and Borzoi15 (RRID: SCR_026619). Keras 3.0 (RRID: SCR_026159) is used because the mannequin coaching framework, permitting each TensorFlow (RRID: SCR_016345) and PyTorch (RRID: SCR_018536) backends.
Regression fashions
By default, the sum of the cosine similarity between predicted and ground-truth peak-height vectors and the MSE between the log-transformed predicted and ground-truth peak-height vectors is used because the loss perform. By default, the cosine similarity worth is dynamically weighted based mostly on the MSE magnitude. Different loss features carried out in CREsted embrace: cosine similarity, Poisson loss and multinomial loss. By default, the Adam optimizer is used with a studying fee of 1 × 10−3, which is decayed with an element of 4 after 5 epochs with no lower in validation loss, and stopping is utilized after ten epochs with no lower in validation loss.
Tremendous-tuning to cell-type-specific areas
Regression fashions could be fine-tuned on cell-type-specific peaks with a decrease studying fee than their base fashions. Cell-type-specific peaks are outlined as these with ({rm{Giniindex}} > {mu }_{{rm{Giniindex}}}+{rm{s}}.{{rm{d}}.}_{{rm{Giniindex}}}); ({mu }_{{rm{Giniindex}}}) and ({{rm{s}}.{rm{d}}.}_{{rm{Giniindex}}}) with ({mu }_{{rm{Giniindex}}}) and ({{rm{s}}.{rm{d}}.}_{{rm{Giniindex}}}) are the typical Gini index and customary deviation of the Gini index throughout all peaks, respectively.
Classification fashions
By default, the binary cross-entropy loss perform is used with an Adam optimizer and studying fee of 1 × 10−3. The educational fee is decayed with an element 4 after 5 epochs with no lower in validation loss and stopping is utilized after ten epochs with no lower in validation loss.
Mannequin architectures
A number of architectures are carried out in CREsted. By default, a dilated convolution neural community (CNN) is used for regression and an everyday CNN for matter classification.
Switch studying from pretrained supervised fashions
For switch studying from Enformer10 and Borzoi15, the fashions’ enter dimension is shrunk (Supplementary Observe 3), and they’re fine-tuned to foretell cell-type-specific peak heights in two rounds: one on consensus peaks and one other on cell-type-specific peaks with a lowered studying fee.
Enhancer code evaluation
Contribution rating calculations
Each gradient-based and prediction-based strategies are used for contribution calculation, impressed by the tfomics package deal37.
$${{rm{I}}{rm{G}}}_{i}(x,{x}^{{prime} })={int }_{alpha =0}^{1}frac{{rm{partial }}f({x}^{{prime} }+alpha (x-{x}^{{prime} }))}{{rm{partial }}{x}_{i}}{rm{partial }}alpha$$
(1)
$${{rm{I}}{rm{G}}}_{i}(x,{x}^{{prime} }=0)={int }_{alpha =0}^{1}frac{{rm{partial }}f(0+alpha (x-0))}{{rm{partial }}{x}_{i}}{rm{partial }}alpha ={int }_{alpha =0}^{1}frac{{rm{partial }}f({ax})}{{rm{partial }}{x}_{i}}{rm{partial }}alpha$$
(2)
$${mathrm{EI}{rm{G}}}_{i}(x)=frac{1}{m}mathop{sum }limits_{n=1}^{m}(mathrm{IG}({x}_{i},{S(x}_{i})))$$
(3)
With:
(x), the sequence to elucidate;
(x{prime}), the baseline sequence;
(f(x)), the mannequin to elucidate with;
(alpha), the progress alongside the combination path;
(Sleft(xright)), a perform randomly permuting x alongside the sequence axis; and
(m), the variety of permuted baselines to make use of.
In built-in gradients81 (IGs; equation (1)), gradients are computed alongside a path from a baseline to the enter sequence being defined. By default, a zeroed-out sequence is used as baseline (equation (2)) and the integral is approximated utilizing the Riemann integral over gradients for 26 steps (α), together with the baseline and ultimate sequence. In anticipated built-in gradients82 (EIGs; equation (3)), impressed by anticipated gradients83, the enter sequence is permuted 25 occasions (m; by default) and that is used because the baseline.
In silico saturation mutagenesis mutates every enter nucleotide to all different nucleotides and quantifies the impact of every mutation on the mannequin’s prediction.
TF-MoDISco evaluation
Contribution scores, for the related class(es), are calculated on cell-type-specific areas and tfmodisco-lite38,39 (RRID: SCR_024811) is carried out to acquire motifs from these areas. We offer a motif database34, which is matched to the discovered motifs in a report.
Sample clustering
Tomtom motif similarities are calculated for all patterns throughout TF-MoDISco runs per cell kind utilizing tangermeme40 (v.0.4.0; RRID: SCR_026620). Teams of patterns passing a sure similarity rating threshold are merged and represented by the sample with the best info content material. Single-instance patterns with an info content material under an outlined threshold are discarded.
Sample to TF matching
Recognized motif patterns are matched to TF candidates based mostly on a motif-to-TF database34, and the Pearson correlation between the significance vector of the motif sample over all cell sorts and the typical expression per cell kind is calculated. Solely TFs that go a correlation threshold (0.2 by default) are stored.
Enhancer design
Seed sequence technology
The enhancer design strategies in CREsted are impressed by Taskiran et al.41. Random DNA sequences of a given size are generated that observe the GC content material of genomic sequences. To this finish, the fraction of every nucleotide of genomic sequences is calculated for every place, and nucleotides are sampled in keeping with this position-dependent distribution.
Optimization perform
Two optimization features are carried out in CREsted. The default perform optimizes a weighted distinction between a goal class of curiosity and all different lessons (equation (4)).
$${rm{price}}_{{S_i}}=[{X}_{{ct}}({S}_{i})-{X}_{{ct}}({S}_{i-1})]-frac{1}{N}mathop{sum }limits_{bg}^{B}({w}_{{bg}}[{X}_{{bg}}({S}_{i})-{X}_{{bg}}({S}_{i-1})])$$
(4)
With:
({ct}), the goal class;
({S}_{i}), the designed sequence at iteration i;
({X}_{{ct}}(S)), the prediction rating of the mannequin on sequence (S) for cell kind ct;
(N), the entire variety of lessons;
(B), the set of lessons aside from the goal class; and
({w}_{{bg}}), the user-defined weight for sophistication ‘bg’ (defaults to 1).
One other perform optimizes the Euclidean distance between a predicted vector of chromatin accessibility to a user-defined goal one (equation (5)).
$${{rm{price}}}_{s}=sqrt{{sum }_{{ct}}^{C}{({X}_{{ct}}(s)-{t}_{{ct}})}^{2}}$$
(5)
With:
C, the set of cell forms of curiosity;
({X}_{{ct}}(s)), the prediction rating of the mannequin for cell kind ct on sequence s; and
({t}_{{ct}}), the goal prediction rating for cell kind ct.
Customized optimization features could be outlined by the consumer.
ISE
A set of seed sequences is optimized by making all potential nucleotide substitutions for every sequence and deciding on probably the most optimum substitution in keeping with an optimization perform. This course of is repeated for a user-defined variety of iterations41.
Motif embedding
A set of seed sequences is optimized by sequentially inserting a set of user-defined TFBSs in every sequence and deciding on probably the most optimum location in keeping with an optimization perform, whereas guaranteeing that the newly positioned binding website doesn’t overlap beforehand positioned binding websites41.
Enhancer fashions
Mannequin coaching
CREsted mouse cortex peak regression mannequin, DeepBICCN2
Mouse motor cortex was downloaded from ENCODE42 (GSE229169), and cut-site BigWigs (RRID: SCR_007708) had been generated per cell kind16. Preprocessing and coaching (pretraining and fine-tuning) had been carried out utilizing default parameters, besides utilizing the highest 3% of peaks per cell kind to calculate peak-normalization scaling elements. Areas of chromosomes 8 and 10 had been used for validation (56,064 consensus and 9,951 cell-type-specific peaks), 9 and 18 for check (49,936 consensus and eight,198 cell-type-specific peaks) and the remaining for coaching (440,993 consensus and 73,326 cell-type-specific peaks). Stochastic 3-bp sequence shift and reverse-complement augmentation had been used throughout coaching with a batch dimension of 256 for pretraining and 64 for fine-tuning.
gReLU mouse cortex peak regression mannequin
A default (6 × 106 parameters utilizing 512 convolutional filters) and huge (22 × 106 parameters utilizing 1,024 convolutional filters) multi-class regression mannequin was skilled with the gReLU package deal29 (v.1.0.3; RRID: SCR_026621), following the 3_train.ipynb tutorial pocket book. The identical cut-site BigWigs (RRID: SCR_007708) and consensus peaks from the Zemke et al. dataset42 and the identical prepare–validation–check splits and the identical peak-height scalars per cell kind as used for coaching the CREsted peak regression mannequin had been used.
CREsted human PBMC peak regression mannequin, DeepPBMC
Protection BigWigs (RRID: SCR_007708) had been downloaded from (ref. 84). Preprocessing and coaching (pretraining and fine-tuning) had been carried out utilizing default parameters, besides utilizing the highest 3% of peaks per cell kind to calculate peak-normalization scaling elements. Areas of chromosomes 8 and 10 had been used for validation (29,385 consensus and 6,070 cell-type-specific peaks), 9 and 18 for check (19,892 consensus and 4,303 cell-type-specific peaks) and the remaining for coaching (278,687 consensus and 51,644 cell-type-specific peaks). Stochastic 3-bp sequence shift and reverse-complement augmentation had been used throughout coaching with a batch dimension of 256 for pretraining and 64 for fine-tuning (utilizing 1 × 10−6 as the training fee).
CREsted most cancers cell line peak regression mannequin, DeepCCL
Protection BigWigs (RRID: SCR_007708) had been generated for the cell traces HepG2, GM12878, MM029, MM099, MM001, A172, M059J and LN229 utilizing deepTools85 (RRID: SCR_016366) and WiggleTools86 (RRID: SCR_001170) and peaks had been referred to as utilizing MACS87 (v.3; RRID: SCR_013291) and merged into consensus peaks utilizing pycisTopic34 (v.2; RRID: SCR_026618). Preprocessing and coaching (pretraining and fine-tuning) had been carried out utilizing default parameters, besides utilizing the highest 3% of peaks per cell kind to calculate peak-normalization scaling elements, defining cell-type-specific areas with coefficient of variation of the protection throughout all areas higher than 0.9, utilizing the DilatedCNN mannequin structure with Swish activation perform, a filter dimension of 11 for first convolutional layer and a dropout of 0.2, utilizing the PoissonLoss perform over log-transformed values and utilizing the Lion optimizer88. Areas of chromosomes 7, 8, 9 and 10 had been used for validations (80,674 consensus and 39,585 cell-type-specific peaks), 5 and 6 for check (48,268 consensus and 27,020 cell-type-specific peaks) and the remaining for coaching (285,790 consensus and 140,761 cell-type-specific peaks). Stochastic 5-bp sequence shift and reverse-complement augmentation had been used throughout coaching with a batch dimension of 512, a studying fee of 5 × 10−5 and a weight decay of 1 × 103 for pretraining, and a batch dimension of 64, a studying fee of 5 × 10−6 and a weight decay of 1 × 10−1 for fine-tuning.
ChromBPNet most cancers cell line mannequin
For every cell line, a separate ChromBPNet8 mannequin was skilled utilizing the directions supplied on the corresponding GitHub web page, utilizing the beforehand referred to as peaks and the identical prepare–validation–check break up because the CREsted mannequin. After coaching, the fashions had been ported to CREsted by holding solely the rely output head and integrating all eight fashions into one ensemble. Lastly, the ChromBPNet ensemble was fine-tuned to cell-type-specific areas in an identical approach to the DeepCCL coaching regime. The Pearson correlation coefficient calculated on log-transformed ground-truth peak heights and mannequin predictions (48,268 and 27,020 test-set samples for the bottom and cell-type-specific comparability, respectively) had been used to judge the efficiency of the fashions for every class.
CREsted glioma biopsy matter mannequin, DeepGlioma
BAM information had been downloaded from the European Genome-phenome Archive repository, beneath EGAS00001003845 and EGAD00001005314, transformed to fragment information and used as enter for pycisTopic36 (v.2; RRID: SCR_026618). Pseudobulk ATAC-seq profiles had been generated and peaks had been referred to as utilizing MACS87 (v.2; RRID: SCR_013291; q worth = 0.001), which had been merged into consensus peaks with 500-bp width. Barcodes with lower than 1,000 distinctive fragments, with transcription begin website enrichment under 5 or that had been acknowledged as doublets had been eliminated. Leiden clustering (decision of 0.4) was carried out, leading to 14 clusters that had been manually annotated based mostly on gene exercise. Clusters annotated as wholesome cells had been eliminated, and the identical pycisTopic workflow was rerun leading to 14,275 cells and 311,118 areas. Twenty-five matters had been chosen based mostly on matter metrics and cell-topic possibilities had been corrected for batch results utilizing Concord89 (v0.0.10; RRID: SCR_022798). Area-topic possibilities had been binarized utilizing Otsu thresholding. One matter (matter 3) included solely 21 areas and was excluded from the following evaluation. On the remaining 24 matters, a CREsted classification mannequin was skilled. Areas of chromosomes 8, 9 and 10 had been used for validation (33,229 peaks), 5 and 6 for check (24,889 peaks) and the remaining for coaching (192,256 peaks). Stochastic 50-bp sequence shift and reverse-complement augmentation had been used throughout coaching with a batch dimension of 128.
CREsted zebrafish peak regression mannequin, DeepZebrafish
Fragment file and cell-level metadata file (containing cell-type and growth timepoint annotations) had been downloaded from NCBI’s Gene Expression Omnibus (GEO; GSE243256). The information had been preprocessed utilizing SnapATAC2 (ref. 35; v.2.6.4; RRID: SCR_026622). Consensus peaks had been referred to as by first calling peaks per cell kind–timepoint mixture (snap.tl.macs3) adopted by snap.tl.merge_peaks utilizing default parameters. Subsequent, a cut-site consensus peak rely matrix was generated (snap.pp.make_peak_matrix) setting the counting_strategy choice to ‘insertion’. Lastly, a normalized matrix containing counts per cell kind–timepoint mixture was made (aggregate_X) utilizing the ‘RPM’ normalization technique. The ensuing matrix was used to coach (pretrain and fine-tune) the CREsted peak regression mannequin, utilizing default parameters, besides utilizing 1,024 convolutional filters. Areas of chromosomes 8 and 10 had been used for validation (70,993 consensus and seven,518 cell-type-specific peaks), 9 and 18 as check (77,948 consensus and seven,521 cell-type-specific peaks) and the remaining for coaching (793,273 consensus and 89,637 cell-type-specific peaks). Stochastic 3-bp sequence shift and reverse-complement augmentation had been used throughout coaching with a batch dimension of 128 for each coaching phases and studying fee of two × 10−6 for fine-tuning.
CREsted Borzoi switch studying
A CREsted-ported Borzoi mannequin (replicate 0) was initialized, with its enter dimension diminished to 2,048 bp, cropping layer disabled and ultimate head layers (Conv1D + Softplus) disregarded (Supplementary Observe 3). This base mannequin’s weights had been frozen for the ‘frozen Borzoi models’. The ‘Borzoi CNN models’ adopted the identical structure, besides that solely the convolutional tower was stored (Supplementary Observe 3). The trunk mannequin was then prolonged with Flatten, Dense and Softplus layers to foretell 19 pseudobulk lessons. Fashions had been skilled on the Zemke et al. dataset42, following the identical knowledge and coaching protocol as DeepBICCN2, aside from the two,048-bp enter dimension, a batch dimension of 32 and studying fee of 5 × 10−5 for full-model first-round fine-tuning (on all consensus peaks) and 1 × 10−5 for second-round fine-tuning (on cell-type-specific peaks) and frozen mannequin fine-tuning. The ‘frozen Borzoi models’ had been fine-tuned for one spherical solely on both all consensus peaks or cell-type-specific peaks, as a single Dense layer doesn’t profit from a number of rounds of fine-tuning.
For all fashions, the bottom validation loss epoch was stored as the ultimate mannequin.
Tremendous-tuning gLMs
HyenaDNA74 (RRID: SCR_027471, hyenadna-small-32k, and Nucleotide Transformer75 (RRID: SCR_027472, nucleotide-transformer-500m-1000g, fashions had been retrieved from the Hugging Face repository utilizing the Transformers library (v.4.54.1; RRID: SCR_027381) and had been fine-tuned for cell-type-specific chromatin accessibility. For each fashions, the final layer’s embeddings had been imply pooled, and a Dense layer adopted by Softplus activation was added and an enter of two,048 bp was used. As a substitute of one-hot encoding the enter, they had been handed by means of the respective fashions’ tokenizers. PyTorch Lightning (v.2.5.2; RRID: SCR_027468), mixed with CREsted (v.1.5.0) utilizing the PyTorch backend (v.2.6.0+cu124) had been used for mannequin coaching. Each fashions had been fine-tuned for 2 rounds utilizing the identical knowledge and loss perform because the Borzoi fine-tuning, stopping when their validation loss plateaued. The Nucleotide Transformer was fine-tuned with unique studying charges of 5 × 10−5 and 1 × 10−5 for the first-round and second-round fine-tuning, with a batch dimension of 4 and a most of 20 epochs. HyenaDNA was skilled with a studying fee of 1 × 10−5 for the primary spherical and 5 × 10−6 for the second spherical, a batch dimension of 32 and a most of fifty epochs.
Mannequin analysis
CREsted and gReLU efficiency comparability
The imply Pearson correlation between log-transformed ground-truth peak heights and mannequin predictions (49,936 consensus peaks and eight,198 cell-type-specific peaks) for every class (n = 19 lessons) had been used to judge mannequin efficiency, and a two-sided Welch’s t-test was used to check for vital variations between the correlation values per class throughout fashions. We report the t-statistic, levels of freedom and two-sided P worth, which had been corrected for a number of testing utilizing the Benjamini–Hochberg process.
Comparability of base-tuned, fine-tuned and scratch fashions
Three fashions had been skilled on the Zemke et al. mouse cortex dataset42: (1) completely on consensus peaks (base), (2) first on consensus peaks adopted by fine-tuning on cell-type-specific peaks (fine-tuned) and (3) completely on cell-type-specific peaks (scratch). Fashions had been evaluated on test-set areas and utilizing locus scoring. We skilled baseline fashions on all peaks to ascertain a generalizable basis.
Chromosomal break up benchmarking
Fashions had been skilled on the mouse cortex dataset42 utilizing ten totally different chromosomal splits to cowl all chromosomes within the check set at the very least as soon as following default CREsted coaching parameters. Utilizing all fashions, contribution scores had been calculated on 171 functionally validated, on-target, mouse and human genomic enhancers17 for the corresponding goal lessons. The pairwise Spearman correlation of their contribution rating throughout fashions of various chromosomal splits was evaluated.
Closed areas benchmarking
The mouse motor cortex coaching set42 was supplemented with a further 550,000 randomly chosen non-exonic, non-peak areas and along with the consensus peaks from this dataset was used to coach a CREsted peak regression mannequin (‘Nonpeakextended’ fashions) adopted by fine-tuning on cell-type-specific areas. Mannequin efficiency on each test-set peaks and test-set closed areas (outlined as areas with mixture sign equal to zero) had been evaluated for each the bottom and fine-tuned fashions.
Gene locus predictions
Gene loci (50 kb upstream of the gene’s transcription begin website to 25 kb downstream of the three’ untranslated area) had been scored utilizing a 2,114-bp sliding window with 100-bp shifts and, for overlapping positions, the typical prediction rating was used.
CREsted predictions on validated mouse cortex enhancers
A set of 171 functionally validated, on-target, mouse and human genomic enhancers17 had been evaluated utilizing DeepBICCN2. The sequences had been zero-padded to suit the two,114-bp mannequin enter. Equally, the height heights had been evaluated straight and values had been zero-padded to 1,000 bp to suit the mannequin’s output. Precision and recall had been calculated utilizing as floor reality the cell kind the place the enhancer is lively in (together with secondary cell kind if current) utilizing the related lessons.
For base Borzoi, the enhancers had been evaluated of their genomic context by centering on the enhancer sequence and utilizing the encompassing genomic DNA sequence to return to an enter dimension of 524,288 bp and scored for the related mouse cortex lessons taking as predicted values the middle 32 bins, similar to 1,024 bp. These Borzoi predictions (for 19 related lessons matched between Borzoi and DeepBICCN2) had been remodeled to acquire their counts with out soft-clipping (that’s, prediction scores > 96 develop into 96 + (prediction rating − 96)2). An extra peak-scaling part was utilized utilizing the typical prediction on housekeeping promoter areas (centered, and prolonged by genomic flankings) utilized in Supplementary Fig. 1 as scaling elements.
EBF1 degradation evaluation in B cells
Precise hits of CCCCTGGG, CCCTAGGG and TCCCTGGG had been recognized and thought of as potential EBF1 TFBSs and changed by N’s, leading to a zero enter. The TCF3 gene locus (20 kb upstream and 5 kb downstream) was evaluated utilizing ‘crested.tl.score_gene_locus’ and in comparison with chromatin accessibility knowledge of wild-type B cells and B cells after EBF1 degradation58.
CREsted predictions in opposition to matters derived from the cisTopic evaluation of human Gliomas
Pseudobulk matter BigWigs (RRID: SCR_007708) had been generated utilizing the ‘export_pseudobulk’ perform in pycisTopic36 (v.2; RRID: SCR_026618) after binarizing cell-topic distributions. DeepCCL predictions and topic-pseudobulk accessibility had been evaluated on the highest 10,000 differentially accessible areas per cluster by averaging the predictions and accessibility of GBM and melanoma MES-like cell traces, and making use of Spearman correlation to determine potential biopsy matters that share an identical MES-like regulatory panorama. Equally, the predictions and accessibility of LN229 had been used to seek out candidate OPC/NPC-like matters.
GBM CNV evaluation
epiAneufinder90 v.1.1.3 (RRID: SCR_026269) was utilized on to the rely matrix of 14,275 cells and 311,118 areas derived from the pycisTopic evaluation of glioma biopsy samples utilizing default parameters and a 100-kb window dimension. The ensuing dataframe was filtered to incorporate solely cells belonging to the cell state of curiosity (MES-like) or to the related matters (matter 8 and matter 21). Home windows had been then labeled as both impartial (>99.9% of cells labeled as regular) or CNV (>2% of cells labeled with both achieve or loss)
CREsted predictions on validated zebrafish enhancers
A set of 54 examined and lively enhancers (Supplementary Desk 6 in Solar et al.76) had been rescaled to 2,114 bp centered on the center of every enhancer and scored utilizing DeepZebrafish.
Comparisons of CREsted-trained fashions with base Borzoi
Pseudobulk BigWigs (RRID: SCR_007708) had been downloaded19 ( for chosen cell sorts similar to these of Zemke et al.42. To this finish, every peak was padded to 524,288 bp utilizing the mm10 genome sequence and zeros the place it fell outdoors chromosome boundaries and the core 32 bins (similar to 1,024 bp) had been used for the comparability.
Motif evaluation
Mouse cortex motif evaluation
Contribution scores for every cell kind had been calculated on the two,000 most cell-type-specific areas (obtained based mostly on the typical of chromatin accessibility and prediction rating per area) and ‘tfmodisco-lite’ was utilized utilizing the perform: crested.tl.modisco.tfmodisco(window = 1000, max_seqlets = 20000) adopted by clustering of patterns utilizing: crested.tl.modisco.process_patterns(sim_threshold = 4, trim_ic_threshold = 0.025, discard_ic_threshold = 0.2) and crested.tl.modisco.create_pattern_matrix(normalize = False, pattern_parameter = ‘seqlet_count_log’).
Human PBMC motif evaluation
Contribution scores, for every cell kind, had been calculated on the highest 1,000 most cell-type-specific areas (obtained based mostly on the typical of chromatin accessibility and prediction scores per area) and ‘tfmodisco-lite’ was utilized utilizing the perform: crested.tl.modisco.tfmodisco(window = 1000, max_seqlets = 20000) adopted by clustering of patterns utilizing crested.tl.modisco.process_patterns(sim_threshold = 4.25, trim_ic_threshold = 0.05, discard_ic_threshold = 0.2) and crested.tl.modisco.create_pattern_matrix(normalize = False, pattern_parameter = ‘seqlet_count_log’).
Human PBMC ChIP–seq comparability
ChIP–seq peaks and BigWigs (RRID: SCR_007708) had been downloaded from ENCODE for PAX5 (ENCFF827VVQ and ENCFF914QGY), EBF1 (ENCFF895MHN and ENCFF810XRY) and POU2F2 (ENCFF803HIP and ENCFF934JFA) and from ChIP-Atlas ( for GATA3 (SRX4705120), RUNX1 (SRX1492212), ETS1 (SRX015825), CEBPA (SRX097095) and SPI1 (SRX4001818). For every sample recognized within the PBMC motif evaluation, TFs had been assigned based mostly on Tomtom40 (v.0.4.0; RRID: SCR_026620) similarity with our motif database34. Precision and recall was calculated, utilizing a number of contribution rating thresholds, the place true positives point out seqlets inside a ChIP peak, false-positive seqlets that aren’t contained in the peaks and false-negative ChIP peaks with out seqlets.
Human PBMC UniBind comparability
Unibind56-predicted direct ChIP–seq peaks had been downloaded for: PAX5 and EBF1 in B cells ( and for GATA3, RUNX1 and ETS1 in CD4+ T cells ( and and for CEBPA and SPI1 in CD14+ monocytes ( and The overlap of these websites and TFBS situations from DeepPBMC, calculated utilizing the tangermeme40 (RRID: SCR_026620) recursive_seqlets perform with a P-value parameter of 0.05, was assessed. Overlaps had been solely thought of when at the very least 50% of the occasion overlapped with the Unibind website. This evaluation was carried out for each the highest 1,000 most cell-type-specific areas and all consensus peaks.
Human PBMC motif enrichment evaluation utilizing pycisTarget
A cisTarget motif database was generated on the PBMC consensus peaks by first making a fasta file with 1-kb background padding utilizing the command line utility create_fasta_with_padded_bg_from_bed.sh and scoring utilizing the v.10 SCENIC+34 motif assortment (RRID: SCR_026702; and utilizing the command line utility create_cistarget_motif_databases.py. Each utilities can be found on (RRID: SCR_027473). Subsequent, pycisTarget34 (v.1.1; RRID: SCR_026626) was run on the identical cell-type-specific peaks as used for the tfmodisco-lite evaluation (see ‘Human PBMC motif analysis’) utilizing default arguments.
Human PBMC motif enrichment evaluation utilizing pyChromVAR
The PBMC fragment matrix was processed utilizing pyChromVAR57 (v.0.0.4; RRID: SCR_027456). Peaks current in fewer than 50 cells and cells with fewer than 2,000 or greater than 15,000 detected peaks, or with fewer than 4,000 or greater than 40,000 complete counts, had been eliminated. Time period frequency–inverse doc frequency (TF-IDF) rely normalization was used and dimensionality was diminished by latent semantic indexing, excluding the primary part. Motifs retrieved from the JASPAR 2024 CORE (RRID: SCR_003030) vertebrate assortment had been recognized. Per-cell deviations (that’s, actions) of motifs had been computed with pyChromVAR utilizing default parameters. Motif enrichment was computed by Wilcoxon rank-sum testing of the motif deviation scores. Lastly, the ‘Human PBMC UniBind comparison’ evaluation was repeated utilizing the motifs recognized with pyChromVAR.
Comparability of contribution-based patterns and enriched motifs from pycisTarget and pyChromVar
A pairwise similarity matrix was calculated between all recognized motifs (each de novo by CREsted and enriched motifs discovered by pycisTarget and/or pyChromVar) utilizing TomTom from memesuite-lite (v.0.2; RRID: SCR_027429)91. t-SNE dimensionality discount was carried out on this matrix utilizing Scanpy (v.1.11.4; RRID: SCR_018139)92.
Contribution rating calculation comparability benchmark
Contribution scores had been calculated on the 171 in vivo-validated enhancers from Ben-Simon et al.17 for every enhancer’s goal class utilizing three strategies: IGs, EIGs and ISM (contemplating the utmost absolute impact per place). Pairwise comparisons had been evaluated utilizing Spearman and Pearson correlation. Moreover, the ‘Human PBMC UniBind comparison’, was rerun utilizing ISM as a substitute of EIGs.
GBM motif evaluation
For DeepCCL, contribution scores for every cell kind had been calculated on the highest 2,000 most cell-type-specific areas (based mostly on the typical of the chromatin accessibility and prediction rating per area) and tfmodisco-lite was carried out utilizing the perform: crested.tl.modisco.tfmodisco(window = 1000, max_seqlets = 20000), and the recognized patterns had been clustered utilizing crested.tl.modisco.process_patterns(sim_threshold = 3.5, trim_ic_threshold = 0.1, discard_ic_threshold = 0.2) and crested.tl.modisco.create_pattern_matrix(normalize = False, pattern_parameter = ‘seqlet_count’). The seqlet counts of the A172/M059J and the MM029/MM099 cell traces had been averaged and log-transformed (holding the unique signal of the contribution scores). Sample clusters had been manually annotated, disregarding nonmeaningful seqlets and seqlets with zero counts in each GBM and melanoma cell traces.
For DeepGlioma, contribution scores, per matter, had been calculated on the highest 2,000 areas of matters 8, 21, 25, 20, 14, 18 and 19, and tfmodisco-lite was carried out utilizing crested.tl.modisco.tfmodisco(window = 500, max_seqlets = 20,000).
For the mixed evaluation, contribution scores had been calculated for every cell line utilizing DeepCCL on the 1,000 most cell-type-specific areas (based mostly on the typical chromatin accessibility and prediction rating per area) and for every matter utilizing DeepGlioma on the highest 1,000 areas per matter. Moreover, the contribution scores of matter areas had been calculated for all cell traces utilizing DeepCCL. The Spearman correlation was used to check contribution scores for every area throughout fashions (taking the center 500 bp when evaluating the DeepCCL and DeepGlioma contributions) and the median correlation coefficient throughout areas was used to acquire a similarity rating between all cell traces and biopsy matters on a nucleotide contribution stage. Lastly, sample clustering was carried out utilizing crested.tl.modisco.process_patterns(sim_threshold = 3.5, trim_ic_threshold = 0.1, discard_ic_threshold = 0.2) and crested.tl.modisco.create_pattern_matrix(normalize = False, pattern_parameter = ‘seqlet_count_log’).
Zebrafish motif evaluation
Contribution scores had been calculated for all designed enhancers for the lessons: ‘Slow muscle cells’, ‘Fast muscle cells’, ‘Cardiac muscle’ and ‘Endothelial’ of growth stage 72 hpf and ‘tfmodisco-lite’ was run (crested.tl.tfmodisco utilizing default parameters). Muscle patterns had been clustered utilizing the features crested.tl.modisco.process_patterns and parameters sim_threshold = 4.25, trim_ic_threshold = 0.05 and discard_ic_threshold = 0.15 and visualized in a warmth map utilizing the perform clustermap_with_pwm_logos utilizing parameter importance_threshold = 4.
Enter dimension benchmark
Mannequin coaching
Fashions with enter sizes of 132 bp, 264 bp, 528 bp, 1,057 bp, 2,114 bp and 4,228 bp had been skilled on the PBMC dataset. First, the information had been preprocessed as described beneath ‘CREsted human PBMC peak regression model, DeepPBMC’. The variety of dilation layers was tailored based mostly on the enter dimension (the place the utmost variety of dilation layers was decided utilizing ({rm{outputwidth}}={rm{dilationfactor}}occasions ({rm{kernelsize}}-1)+1); and deciding on the maximal variety of layers minus 1 that also had output width ≥ 0) or zero-padding (by setting padding parameter to ‘same’) was used to bypass convolutional layers with dimensions lower than or equal to zero. This resulted in a complete of 12 base fashions that had been skilled (278,687 consensus peaks; batch dimension 128) and fine-tuned (51,644 cell-type-specific peaks; batch dimension of 64; studying fee of 1 × 10−6).
Comparability of seqlet areas and recognized patterns
Contribution scores had been calculated for the highest 2,000 most cell-type-specific areas per cell kind (crested.pp.sort_and_filter_regions_on_specificity on the typical of the particular and predicted cell-type accessibility in keeping with every mannequin), and seqlets had been recognized utilizing the tangermeme40 (RRID: SCR_026620; v.0.4.0) perform recursive_seqlets utilizing default parameters. Lastly, patterns had been recognized and in contrast throughout fashions utilizing TF-MInDi93 (RRID: SCR_027436; v.1.0.0).
Zebrafish enhancer design
Zebrafish cardiac and physique muscle enhancer design
Cell-type lessons of any timepoint annotated a: ‘Slow muscle cell’, ‘Slow muscle cells’, ‘Fast muscle cells’, ‘Heart’, ‘Heart field’ or ‘Cardiac muscle’ had been thought of and the typical over all constructive enhancers (see ‘CREsted predictions on validated zebrafish enhancers’) of the utmost prediction rating over all lessons was calculated (‘avg_pos_prediction_score’). A value perform was outlined that minimizes the Euclidean distance between the mannequin’s prediction rating and a goal vector the place lessons similar to ‘Cardiac muscle’ equals ‘avg_pos_prediction_score’ and different cell sorts are zero. Equally, for physique muscle a goal vector was used the place lessons similar to ‘Slow muscle cell’, ‘Slow muscle cells’ or ‘Fast muscle cells’ equals ‘avg_pos_prediction_score’ and the opposite cell sorts are zero. Three units of enhancers had been designed to be lively in each cell sorts, with various ranges of prediction scores: cardiac1/physique1, with equal prediction rating in cardiac and physique muscle cells; cardiac1/physique0.5, the place the prediction rating in physique muscle cells is half the prediction rating in cardiac muscle cells; and vice versa for cardiac0.5/physique1. 200 random DNA sequences had been initialized with comparable nucleotide content material throughout the height area as consensus peaks and had been optimized utilizing ISE (Supplementary Desk 1).
Zebrafish endothelial enhancer design
The identical method as was used to design enhancers focused for both cardiac or physique muscle was used to design endothelial cells. Courses similar to ‘Notochord’, ‘Endothelial’ and ‘Otic placode’ had been thought of, and a goal vector the place Endothelial was set to ‘avg_pos_prediction_score’ and the others to zero was used (Supplementary Desk 1).
Enhancers choice for experimental validation
From the 200 generated sequences for every job (see above), 5 sequences had been sampled at random per job and the nucleotide contributions had been visually inspected. Primarily based on this guide inspection, 3 of 5 randomly sampled sequences had been examined for every job.
Human most cancers cell traces
Tradition of human GBM cell traces
Cells had been cultured following routine cell tradition procedures, as described in, for instance, A172 cells (American Kind Tradition Assortment (ATCC), CRL-1620; RRID: CVCL_0131) had been cultured in DMEM (Thermo Fisher, 11965-092) medium supplemented with 10% FBS (Thermo Fisher, 17479-633), and 1% penicillin–streptomycin (Life Applied sciences, 15140122). MO59J cells (ATCC, CRL-2366; RRID: CVCL_0400) had been cultured in DMEM/F12 (Thermo Fisher, 11320-033) medium supplemented with 10% FBS, 1% penicillin–streptomycin and 0.05 mM non-essential amino acids (Thermo Fisher, 11140-068). LN229 cells (ATCC, CRL-2611; RRID: CVCL_0393) had been cultured in DMEM (Thermo Fisher, 11965-092) medium supplemented with 5% FBS and 1% penicillin–streptomycin. All cell traces had been passaged twice per week with Trypsin-EDTA 0.05% (Thermo Fisher, 11580-626).
Bulk ATAC-seq
Cells had been handled with Trypsin-EDTA 0.05% and washed with DPBS (Thermo Fisher, 14190-169). Cell viability and focus had been assessed by the LUNA-FL Twin Fluorescence Cell Counter. Per cell line, 80,000 cells had been used to carry out ATAC-seq, following the OmniATAC-seq protocol94. After centrifugation at 500g for five min at 4 °C, the supernatant was changed with 50 µl ice-cold ATAC-seq lysis buffer (10 mM Tris-HCl, 10 mM NaCl, 3 mM MgCl2, 0.1% IGEPAL CA-630, 0.1% Tween-20, 1% BSA and 0.01% digitonin), and the cells had been combined by pipetting. After incubation for five min on ice, 1 ml of wash buffer (20 mM Tris-HCl, 20 mM NaCl, 6 mM MgCl2, 0.1% Tween-20 and 1% BSA) was added to the lysed cells and combined gently by inverting thrice. After centrifugation at 500g for five min at 4 °C, supernatant was eliminated and the nuclei had been resuspended in 50 µl ATAC combine (10 mM Tris-HCl, 10% dimethylformamide, 5 mM MgCl2, 1× DPBS, 0.1% Tween-20, 0.01% digitonin and three.75 ng µl−1 Tn5 enzyme) and incubated for 1 h at 37 °C. Transposed DNA was purified utilizing a QIAGEN MinElute purification column and eluted in 15 µl elution buffer (Qiagen). The ATAC-seq library was accomplished by amplifying the transposed DNA in a complete quantity of fifty µl PCR combine (15 µl purified DNA, 25 µl NEBNext Excessive Constancy PCR Grasp Combine (NEB), 2.5 µl FWD and a pair of.5 µl REV primer) with the next program: 72 °C for five min, 98 °C for 30 s, adopted by ten cycles of 98 °C for 10 s, 63 °C for 30 s and 72 °C for 1 min). The ultimate libraries had been subjected to 0.4–1.2× double-sided Ampure purification and eluted in 20 µl elution buffer (Qiagen).
Sequencing
ATAC-seq libraries had been sequenced on an Illumina NovaSeq X system utilizing 51 cycles for learn 1 (ATAC paired-end mate 1), 8 cycles for index 1 (pattern index 1), 8 cycles for index 2 (pattern index 2) and 51 cycles for learn 2 (ATAC paired-end mate 2).
Knowledge processing
FASTQ information for HepG2 (RRID: CVCL_0027) and GM12878 (RRID: CVCL_7526) had been downloaded from ENCODE53,95 ( RRID: SCR_006793) beneath accession numbers ENCLB324GIU and ENCLB907YRF, respectively. The standard of those reads along with the newly generated ones for A172, M059 and LN229 was evaluated utilizing FastQC (v.0.11.9, RRID: SCR_014583) and any adaptors had been trimmed utilizing the fastq-mcf command from ea-utils (v.1.1.2.779, RRID: SCR_005553). Reads had been mapped to the hg38 genome utilizing Bowtie2 (ref. 96; v.2.4.4, RRID: SCR_016368), and duplicates had been eliminated utilizing Picard (v.2.27.1, RRID: SCR_006525). Peaks had been referred to as utilizing MACS3 (ref. 87; v.3.0.0b1, RRID: SCR_013291), and mixture genome protection was generated with the bamCoverage command of deepTools85 (3.5.0, RRID: SCR_016366). The protection of replicates was averaged utilizing WiggleTools (v.1.2.11, RRID: SCR_001170). For the melanoma cell traces, fragment information had been downloaded from the GEO beneath accession quantity GSE210745. Pseudobulk ATAC-seq profiles had been generated per cell line as proven within the SCENIC+ tutorial34 and peaks had been referred to as for every pseudobulk. From these, we chosen MM001, MM029 and MM099 for our evaluation. The ensuing peaks from all eight cell traces (that’s, the melanoma, GBM and ENCODE ones) had been merged right into a consensus set utilizing the ‘get_consensus_peaks’ perform of pycisTopic36 and had been used along with the eight protection information to coach the DeepCCL mannequin. ChIP–seq peaks for JunB, c-Jun and c-Fos in MM099 had been downloaded from the GEO utilizing identifier GSE159965 and for ZEB1 by means of As well as, we retrieved ChIP–seq peaks for TEAD1 in MSTO cells and TEAD4 in SK-MEL-147 by means of ReMap97 beneath accession numbers GSE68170 and GSE94488, respectively.
Authentication of newly generated cell line knowledge
The identification of all cell traces on which new knowledge had been generated on this examine was confirmed based mostly on the protection of single-nucleotide polymorphisms by the ATAC-seq knowledge, in comparison with publicly accessible knowledge on these cell traces (downloaded from ENCODE for A172 (ENCSR932KWJ) and M059J (ENCSR000EPG) and from the Sequence Learn Archive for LN229 (SRX15782182)) utilizing the instrument NGScheckmate98.
Enhancer reporter assays
In vivo validation of zebrafish enhancers
An in depth protocol describing enhancer reporter plasmid cloning, egg microinjection and imaging is obtainable on protocols.io by way of
Enhancer reporter plasmid
Artificial enhancer DNA sequences had been ordered from Twist Bioscience with fixed flanking DNA sequences: 5′ finish ‘ATATACCCTCTAGAGTCGAA’ and three′ finish ‘GATTACCCTGTTATCCCTAA’. These flanking areas had been used for PCR amplification of the sequences utilizing primers containing overhangs which might be homologous to the goal plasmid.
Fwd: 5′ TTAGGGATAACAGGGTAATCGCGAATTGGGTACCGGGC 3′
Rev: 5′ CTTTCAACAAGCCCGAAAGATCTTCTGGAAGCCTCCAGTGAATT 3′
KAPA HiFi HotStart ReadyMix (Roche) with primer and template concentrations of 300 µM and 0.1 ng µl−1, respectively, had been used and the PCR setup was: 95 °C for 3 min, 20 cycles of 98 °C for 20 min, 65 °C for 15 min, 72 °C for 15 min, and a ultimate elongation step of 72 °C for two min. Subsequent, the goal plasmid, Tol2-ISceI-ZSP:EGFP;cryaa:mCherry-ISceI-Tol2 (Addgene, 194518), was linearized by restriction digestion utilizing BglII and XhoI (at 0.5 U µl−1 and 1 U µl−1, respectively, in rCutSmart buffer, with 7 µg plasmid, 2-h incubation at 37 °C), purified (NucleoSpin Gel & PCR cleanup equipment, Macherey-Nagel) and mixed with purified enhancer DNA fragments in a NEBuilder response (NEBuilder enzyme combine from New England Biolabs, 7 fmol linearized plasmid, 65 fmol DNA fragment, 45-min incubation at 50 °C). Then, 2.5 µl of the response was remodeled into 20 µl of Stellar chemically competent micro organism (thaw cells on ice for 15 min, add plasmid, maintain 30 min on ice, 45 s warmth shock at 42 °C, 5 min on ice), which had been incubated in a single day at 37 °C on carbenicillin plates. Single colonies had been then grown in 10 ml LB medium, and plasmids had been remoted utilizing QIAprep Spin Miniprep Equipment (Qiagen) and eluted in 2 × 25 µl. All plasmid preps had been sequenced utilizing Sanger sequencing to substantiate right insertion within the goal plasmid with out sequence alterations. Enhancers C1, B1, F1, G1 and G2 include level mutations that ought to not have an effect on enhancer exercise in keeping with the mannequin prediction.
Egg microinjection
Plasmid DNA along with Tol2 mRNA had been injected (at concentrations of 30 ng µl−1 and 40 ng µl−1, respectively) in one-cell stage zebrafish eggs (of the wild-type AB pressure), and so they had been grown at 28.5 °C till 48 hpf. Zebrafish with profitable injection had been chosen based mostly on purple fluorescence within the eye on an Olympus SZX16 widefield fluorescence microscope. Fish had been anesthetized with 0.02% tricaine (MS-222 Ethyl 3-aminobenzoate methanesulfonate, Sigma) and mounted in 1% low-melting-point agarose (Invitrogen) on fluorodish (FD3510-100, World Precision Devices). All zebrafish breeding was authorised by the Moral Committee for Animal Experimentation of KU Leuven (ECD-000) and all experiments had been carried out on embryos youthful than 5 days after fertilization.
Imaging
Zebrafish embryos (48 hpf) had been imaged for GFP fluorescence on a spinning-disk confocal microscope consisting of a Nikon Ti2 physique and Crest X-Gentle V2 spinning disk utilizing a ×10 Plan Apo Lambda lens with numerical aperture of 0.45 and a ×20 Plan Apo Lambda lens with numerical aperture of 0.8. The Lumencor Spectra III utilizing the inexperienced channel was used to excite GFP fluorescence, utilizing excitation filter Semrock FF01-378/474/554/635/735-25 and dichroic mirror FF409/493/573/652/759-Di01-25×36. Semrock FF01-515/30-25 was used as an emission filter. NIS-element (RRID: SCR_014329) V6 software program was used to manage the microscope.
Guide evaluation of enhancer cell-type-specific enhancer exercise
For every job, enhancer reporter assays had been carried out in a number of zebrafish embryos (see figures for numbers) and in three replicates (that’s, totally different injection days and clutches). Cell-type-specific exercise of every enhancer was assessed utilizing entire embryo fluorescence microscopy, independently by three researchers, at a single developmental timepoint (48 hpf). Solely the cell-type specificity of every enhancer was assessed, not the extent of exercise, which isn’t potential because of the nature of our setup (that’s, no management for the copy variety of enhancer–reporters which might be current in every cell—both episomaly or built-in within the genome).
Reporting abstract
Additional info on analysis design is obtainable within the Nature Portfolio Reporting Abstract linked to this text.