Moral statements
All procedures involving human members complied with all related moral laws and have been performed in accordance with the Declaration of Helsinki. All datasets have been collected below institutional evaluate board approval (Approval No.2023-1048 by the Second Affiliated Hospital of Zhejiang College Faculty of Drugs, Approval No.IIT-20230612-0118-01 by Hangzhou First Folks’s Hospital, Approval No.2024-KLS-013-02 by Zhejiang Provincial Hospital of Conventional Chinese language Drugs, Approval No.2024-033 by Ningbo Medical Middle Lihuili Hospital, Approval No.IRB-20230357-R by Ladies’s Hospital Faculty of Drugs, Zhejiang College, Approval No.2023-0548 by Sir Run Run Shaw Hospital, Approval No.2024-002 by XiangFu Neighborhood Well being Service Middle). All information have been de-identified earlier than mannequin growth. Written knowledgeable consent was obtained from all members prior to review participation.
Research inhabitants
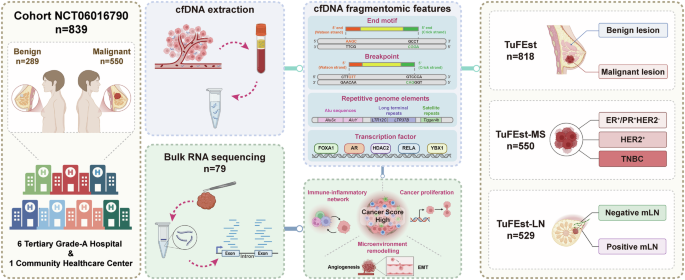
This research was designed as a potential multicenter case-control research (NCT06016790) that repeatedly enrolled 839 members, together with 550 BC sufferers and 289 benign controls from seven Chinese language medical establishments: the Second Affiliated Hospital of Zhejiang College Faculty of Drugs (SAHZU), Hangzhou First Folks’s Hospital (HZFH), Zhejiang Provincial Hospital of Conventional Chinese language Drugs (ZJTCM), Ningbo Medical Middle Lihuili Hospital (LHLI), Ladies’s Hospital Faculty of Drugs Zhejiang College (WHZJU), Sir Run Run Shaw Hospital (SRRSH), and Xiangfu Neighborhood Well being Service Middle (XFCHC). Eligible members have been females aged 18-70 years with pathologically confirmed diagnoses of both: (1) major invasive breast carcinoma or carcinoma in situ, or (2) benign breast lesions (fibroadenoma/adenosis/fibrocystic hyperplasia). All members offered their plasma samples. Exclusion standards included being pregnant/lactation, prior malignancies (5-year window), suspected however unconfirmed malignancies throughout the previous 12 months, latest blood transfusions (30-day window), lively mastitis, and different investigator-determined contraindications.
Pattern assortment and processing
Peripheral blood (8–10 mL) was collected in specialised cfDNA tubes (Ardent BioMed Cat. # BY10240301) throughout routine blood exams or earlier than surgical procedure. Samples have been subjected to twin centrifugation (1,600 × g → 16,000 × g, 10 min every at 4 °C) for plasma isolation, adopted by speedy storage at −80 °C. Frozen plasma was batch-shipped on dry ice to the central laboratory (OmixScience Analysis Institute, Hangzhou, China) for processing.
cfDNA assays and whole-genome sequencing
cfDNA was extracted from a median plasma quantity of 1 mL utilizing the VAMNE MagUltra circulating cell-free DNA isolation equipment (Vazyme Biotech, Nanjing, China, Cat. # N913) in line with the producer’s protocol, adopted by exact quantification utilizing a Qubit 4.0 fluorometer (Thermo Fisher Scientific, Lenexa, KS, USA) and storage at −80 °C. For library preparation, 5–20 ng of the extracted cfDNA was processed with the VAHTS Common DNA library prep equipment for Illumina V3 (Vazyme Biotech, Nanjing, China, Cat. # N610) and barcoded utilizing VAHTS multiplex oligos set 5 for Illumina (Vazyme Biotech, Nanjing, China, Cat. # N322). The standard and integrity of the libraries have been evaluated utilizing an Agilent 2100 Bioanalyzer (Agilent Applied sciences, Santa Clara, CA, USA) to verify that the libraries met the required requirements for optimum fragment measurement distribution. The ultimate libraries have been sequenced on the NovaSeq X Plus platform (Illumina, San Diego, CA, USA) utilizing paired-end 150 bp reads, with a median protection depth of two× for the cfDNA samples.
Uncooked WGS information underwent rigorous high quality management starting with FastQC (model 0.12.1, analysis, adopted by high quality filtering and adapter trimming utilizing Cutadapt (model 4.5, and Ktrim (model 1.4.1, The processed reads have been aligned to the human reference genome (GRCh37) utilizing BWA-MEM (model 0.7.17, with default settings, making certain that solely high-quality reads with a Phred rating above the edge have been retained. Additional analyses have been restricted to uniquely mapped reads that have been freed from PCR duplicates. These mapped reads have been subsequently sorted and listed utilizing Samtools (model 1.9, and duplicates have been recognized and eradicated utilizing Picard (model 2.18.29, The distribution of fragment sizes and related genomic options was inferred based mostly on the coordinates of mapped learn pairs, following established protocols utilizing Picard. To determine large-scale epigenetic variations in cfDNA fragmentation throughout the genome, which may be detected with low-coverage WGS, the hg19 reference genome was segmented into non-overlapping 5 Mb bins.
Fragmentomic function identification
We systematically quantified the frequencies of 4-bp finish motifs and 6-bp breakpoint motifs, normalizing complete motif occurrences to unity inside every class to allow comparative analyses. For the 6 bp motifs, we characterised the three bp sequences flanking cfDNA 5’ breakpoints (each upstream and downstream) and established genome-wide distribution profiles. To keep up consistency, the general frequency of all 6 bp breakpoint motifs was normalized to 1. We additional organized the motif options into an I × J matrix, the place every row corresponds to a participant recognized by their pattern ID. This matrix format permits for environment friendly interpretation and comparability of motif patterns throughout the cohort, resulting in deeper insights into cfDNA motif distributions.
Regional fragment measurement analyses employed 100 kb genomic bins utilizing Deeptools (model 3.1.2, optimized to steadiness decision with ample learn depth (~25,000 reads/bin). Every bin contained quick (S, 100–150 bp) and lengthy (L, 151-220 bp) fragments. To make sure information accuracy, areas overlapping the ENCODE blacklist or the hg19 hole monitor from the UCSC Genome Browser have been excluded, as these areas sometimes present poor mappability. This method yielded sturdy S/L ratios for the fragmentation sample analyses. By specializing in fewer, higher-quality bins, we improved the reliability of our fragment-size distribution analyses.
Alu repeats analyses of cfDNA
We categorized all parts from the gene and pseudogene households as switch RNA (tRNA), sign recognition particle RNA (srpRNA), small nuclear RNA (snRNA), small cytoplasmic RNA (scRNA), and ribosomal RNA (rRNA). Likewise, we grouped parts from the DNA, retroposon, and Rolling Circle (RC) households into the transposable component class. This led to eight major teams for evaluation: LINE, SINE, Medium Repeat (MER), DNA_TcMar-Tigger (DNA Transposable parts), Lengthy Terminal Repeat (LTR), satellites, transposable parts, and RNA parts. In our cfDNA function analyses and validation, we centered on essential parts, together with Alu sequences, lengthy terminal repeats (LTRs), RNA parts, and transposable parts. By this systematic classification, we adopted a radical method to grasp cfDNA options, that are important for investigating their roles in BC. The TuFEst algorithm integrates these repeat parts with the N-index and E-index and incorporates TF options to foretell non-cancerous and cancerous standing.
Machine studying and deep studying algorithm mannequin
On this research, TuFEst was applied as a binary classification mannequin50. It outputs a “cancer score” between 0 and 1, representing the likelihood {that a} plasma cfDNA pattern originates from a person with most cancers. A cutoff of 0.5 was utilized to this rating to categorise samples as “cancer” or “non-cancer.” Though the rating is a surrogate measure of the relative burden of tumor-derived DNA and the identify suggests its long-term potential for quantification, the present software and validation give attention to classification efficiency.
A GLM with a binomial distribution and a logit hyperlink operate, equal to logistic regression, was developed for binary classification. The mannequin utilized fragmentomics options as predictors, with the result variable coded as 1 for malignant and 0 for benign standing. To deal with potential pattern dependencies, group data was integrated utilizing SampleID. The analytical pipeline included function standardization (StandardScaler) adopted by logistic regression with a most of 1000 iterations. Mannequin efficiency was assessed by means of 10-fold stratified cross-validation with shuffling (random state = 42) to make sure reproducibility whereas sustaining class steadiness throughout folds.
A SVM with a linear kernel was applied for classification. The mannequin employed standardized fragmentomics options, with malignant standing encoded as 1 and benign as 0. Pattern identifiers have been retained for group-level evaluation. The pipeline consisted of function scaling adopted by SVM classification with likelihood estimation enabled. Efficiency was evaluated utilizing 10-fold stratified cross-validation, with metrics together with accuracy, AUC, sensitivity, specificity, PPV, and NPV. Chance thresholds at specificity ranges of 0.9 and 0.95 have been derived from ROC evaluation to assist scientific decision-making. All predictions and possibilities have been systematically recorded for aggregated evaluation.
An XGBoost classifier was applied utilizing a reproducible pipeline (random state = 42). Fragmentomics options have been standardized previous to mannequin coaching, with binary outcomes indicating malignant (1) or benign (0) standing. The mannequin was evaluated utilizing 10-fold stratified cross-validation, with complete metrics recorded for every fold. Medical determination thresholds have been established by figuring out likelihood values equivalent to 0.9 and 0.95 specificity ranges from ROC curves. All predictions and pattern metadata have been saved for subsequent evaluation, and the ultimate pipeline was saved for potential future use.
A MLP neural community was developed with fastened random state (42) for reproducibility. Options have been standardized earlier than being enter to the MLP classifier. The mannequin was evaluated utilizing 10-fold stratified cross-validation, with efficiency metrics together with accuracy, AUC, sensitivity, specificity, PPV, and NPV. Chance thresholds for particular specificity ranges (0.9 and 0.95) have been decided from ROC evaluation to outline scientific determination boundaries. The skilled pipeline, together with preprocessing and classification steps, was preserved for potential software, and general discriminative capability was validated by means of aggregated ROC evaluation.
A GBM was applied utilizing scikit-learn’s GradientBoostingClassifier. The mannequin was skilled on standardized fragmentomics options and evaluated through 10-fold stratified cross-validation (random state = 42). Every fold yielded likelihood estimates and efficiency metrics, together with accuracy, AUC, sensitivity, specificity, PPV, and NPV. Medical thresholds have been recognized by setting specificity ranges at 0.9 and 0.95 in ROC evaluation. Predictions, true labels, and group identifiers have been saved for mixed evaluation, and the ultimate mannequin was saved for potential deployment.
A SEM built-in predictions from 5 base learners-SVM, GLM, GBM, XGBoost, and MLP-using a logistic regression meta-learner. The pipeline included function standardization and was evaluated by means of 10-fold stratified cross-validation. Efficiency metrics have been calculated for every fold, and likelihood thresholds have been established at specificity ranges of 0.9 and 0.95. The ensemble demonstrated robust discriminative capability (AUC = 0.943), with all predictions and pattern data systematically recorded. The entire pipeline was saved to assist potential scientific implementation.
For tumor detection and classification, we screened six distinct machine studying algorithms to construct the fashions: GLM, SVM, XGBoost, MLP, GBM, and SEM. These fashions have been rigorously skilled and evaluated utilizing bootstrapping mixed with 10-fold cross validation. Through the 10-fold cross-validation course of, the coaching set was randomly divided into 10 subsets, with 9 subsets used for coaching and the remaining one used for testing, biking 10 instances to make sure that every subset was used for testing as soon as. We solely used coaching cohort samples from benign people and most cancers sufferers, coaching classifiers within the machine studying algorithms with options similar to motif function frequencies, and producing fashions to foretell most cancers scores for every pattern. Notably, all validation datasets remained untouched throughout mannequin coaching. The most cancers rating ranged from 0 to 1, with the next rating indicating a better likelihood of most cancers. After the analysis, we chosen the best-performing mannequin for the downstream analyses.
These fashions have been subsequently utilized to the validation datasets to generate most cancers prediction scores for every validation pattern and assess the mannequin efficiency. We in contrast the AUC values of the completely different fashions within the validation cohort and the sensitivity/specificity at a set specificity threshold within the inside validation cohort.
Three specialised fashions have been developed for various scientific functions. The TuFEst mannequin was designed to tell apart between benign people and most cancers sufferers utilizing a stacked ensemble that includes function data, together with TF binding web site protection, to reinforce predictive accuracy. The TuFEst-MS mannequin employs a GLM technique to categorise three BC subtypes: ER+/PR+, HER2+, and TNBC. This classification functionality is especially essential for predicting the subtype of metastatic lesions in sufferers with BC. The TuFEst-LN mannequin utilized GLM to foretell whether or not a affected person had mLN, particularly for analyzing sufferers with radiological proof of mLN however no surgical affirmation, or these with discordant imaging-pathological outcomes. All fashions underwent rigorous coaching utilizing 10-fold cross validation, with prediction scores calculated for every pattern. The fashions have been assessed utilizing the AUC-ROC metric to make sure their accuracy and reliability in sensible functions.
Choice and analyses of differential options
We utilized the wilcox.take a look at() operate to carry out the Wilcoxon rank-sum take a look at, assessing the statistical significance of variations in varied options throughout completely different classifications. The parameter actual = FALSE signifies using an approximate computational technique. Options with a p-value lower than 0.05 have been chosen to make sure statistical significance and get rid of the affect of random elements. Moreover, we utilized a fold change threshold better than 1, retaining solely options that exhibited vital variations between the 2 teams. Subsequently, the highest and backside 30 options have been recognized, and the median ratio between the benign and malignant teams was calculated for every function. Lastly, a standardized function heatmap was generated utilizing the pheatmap() operate from the pheatmap package deal, whereas the dimensions() operate was utilized for Z-score normalization, enhancing visualization and facilitating additional analyses.
Genome-wide TF analyses
ChIP-seq peaks from 5620 experiments have been downloaded from the ReMap 2020 database ( deciding on the CRMs (Cis-Regulatory Modules) information sort, which features a complete of 1,732,560 peaks.
For every autosomal peak, the height heart was designated as place 0, and the imply protection was computed for every pattern inside a ± 1,000 bp window relative to the height heart. The processed information have been then integrated as a function into the mannequin coaching pipeline to tell apart between benign people and sufferers with most cancers.
A sequence of filters have been utilized to make sure ample sequencing depth throughout benign and most cancers samples, retaining solely peaks with learn counts >25, learn protection ≥0.9, and fold change ≤0.5 or ≥2.
To research the organic processes related to the recognized TFs, we carried out KEGG pathway enrichment analyses utilizing the clusterProfiler package deal in R. TFs have been first mapped to Entrez IDs, and the KEGG enrichment evaluation was utilized to determine considerably enriched pathways. Pathways have been ranked based mostly on their adjusted p-values, and the enriched pathways have been visualized in a bubble chart utilizing ggplot2, with bubble measurement representing gene depend and coloration indicating p-value significance.
Estimation of the tumor immune microenvironment
To comprehensively profile the tumor immune microenvironment, we employed two computational deconvolution strategies: CIBERSORT51 and xCell52, which have been used to quantify the relative proportions of distinct tumor-infiltrating immune cell populations from the majority transcriptomic information. As well as, the ESTIMATE algorithm53 was utilized to calculate immune and stromal scores, reflecting the general infiltration ranges of immune and stromal parts throughout the tumor tissue.
Gene enrichment and pathway analyses
Gene Set Enrichment Evaluation (GSEA) was performed utilizing the “clusterProfiler” R package deal54 to determine considerably enriched hallmark gene units. GSVA was carried out utilizing the “GSVA” R package deal55 to compute sample-wise enrichment scores for particular gene units. Differentially expressed genes between the Most cancers Rating-Excessive and Most cancers Rating-Low teams have been recognized utilizing the “limma” R package deal, with thresholds set at |log₂(fold change)| >1 and P < 0.05.
Statistics & reproducibility
For statistical evaluation, a ROC curve was generated utilizing the pROC package deal (v. 1.18.5). Primarily based on the true positives (TP), true negatives (TN), false positives (FP), and false negatives (FN) for most cancers prediction, we calculated the sensitivity [TP/(TP + FN)], specificity [TN/(Tn + FP)], constructive predictive worth (PPV) [TP/(TP + FP)], unfavorable predictive worth (NPV) [TN/(Tn + FN)], and accuracy [(TP + TN)/(TP + FP + Tn + FN)]. These metrics have been computed utilizing the confusion matrix operate of the caret package deal (v. 6.0.94) in R (v. 4.4.1). The preProcess operate (v.1.2.2) from the caret package deal was used for function standardization. Moreover, the confusion matrix algorithm was employed to check ROC curves, offering a complete analysis of the diagnostic take a look at efficiency and making certain correct differentiation between benign and malignant instances. No statistical technique was used to predetermine pattern measurement. No information have been excluded from the analyses. Pattern processing for cfDNA library preparation and sequencing was randomized by case/management standing to reduce batch results. The Investigators weren’t blinded to allocation in the course of the experiments or throughout final result evaluation.
Reporting abstract
Additional data on analysis design is offered within the Nature Portfolio Reporting Abstract linked to this text.