How shut can an open mannequin get to AlphaFold3-level accuracy when it matches coaching information, mannequin scale and inference finances? ByteDance has launched Protenix-v1, a complete AlphaFold3 (AF3) copy for biomolecular construction prediction, launched with code and mannequin parameters below Apache 2.0. The mannequin targets AF3-level efficiency throughout protein, DNA, RNA and ligand constructions whereas conserving the complete stack open and extensible for analysis and manufacturing.
The core launch additionally ships with PXMeter v1.0.0, an analysis toolkit and dataset suite for clear benchmarking on greater than 6k complexes with time-split and domain-specific subsets.
What’s Protenix-v1?
Protenix is described as ‘Protenix: Protein + X‘, a basis mannequin for high-accuracy biomolecular construction prediction. It predicts all-atom 3D constructions for complexes that may embrace:
- Proteins
- Nucleic acids (DNA and RNA)
- Small-molecule ligands
The analysis group defines Protenix as a complete AF3 copy. It re-implements the AF3-style diffusion structure for all-atom complexes and exposes it in a trainable PyTorch codebase.
The challenge is launched as a full stack:
- Coaching and inference code
- Pre-trained mannequin weights
- Information and MSA pipelines
- A browser-based Protenix Internet Server for interactive use
AF3-level efficiency below matched constraints
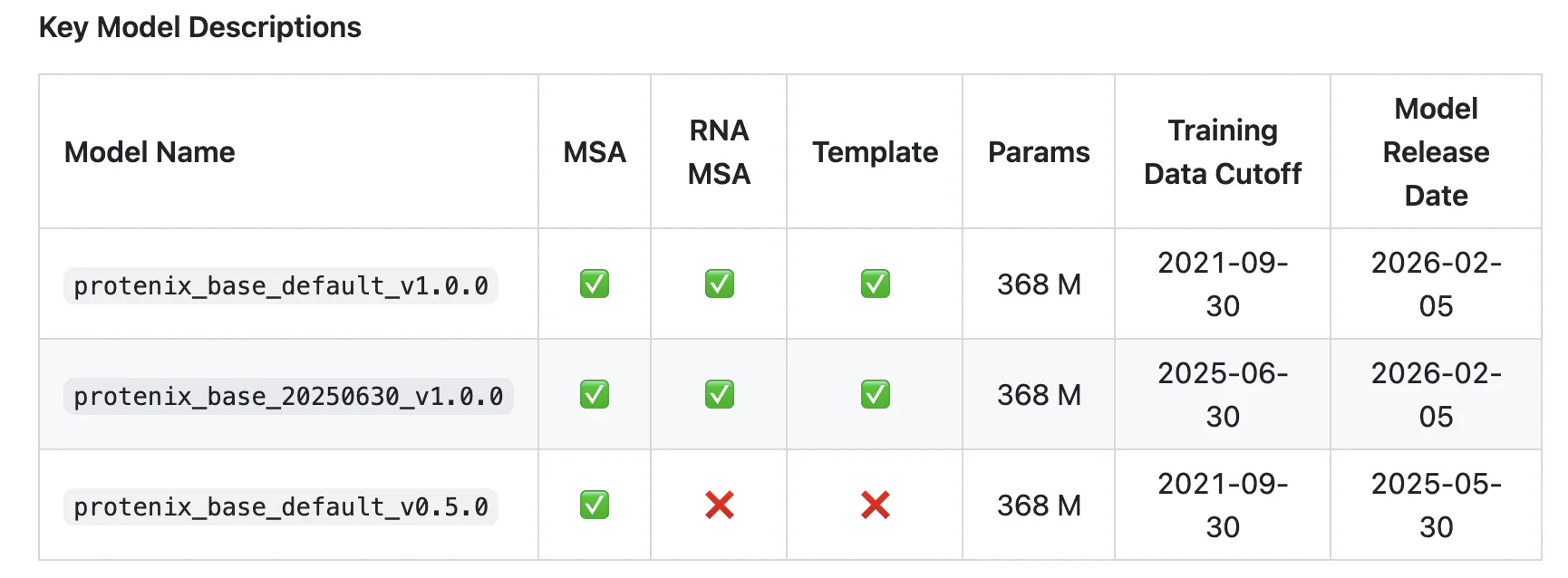
As per the analysis group Protenix-v1 (protenix_base_default_v1.0.0) is ‘the primary absolutely open-source mannequin that outperforms AlphaFold3 throughout numerous benchmark units whereas adhering to the identical coaching information cutoff, mannequin scale, and inference finances as AlphaFold3.‘
The necessary constraints are:
- Coaching information cutoff: 2021-09-30, aligned with AF3’s PDB cutoff.
- Mannequin scale: Protenix-v1 itself has 368M parameters; AF3 scale is matched however not disclosed.
- Inference finances: comparisons use related sampling budgets and runtime constraints.
On difficult targets akin to antigen–antibody complexes, growing the variety of sampled candidates from a number of to lots of yields constant log-linear enhancements in accuracy. This provides a transparent and documented inference-time scaling conduct slightly than a single mounted working level.
PXMeter v1.0.0: Analysis for 6k+ complexes
To assist these claims, the analysis group launched PXMeter v1.0.0, an open-source toolkit for reproducible construction prediction benchmarks.
PXMeter supplies:
- A manually curated benchmark dataset, with non-biological artifacts and problematic entries eliminated
- Time-split and domain-specific subsets (for instance, antibody–antigen, protein–RNA, ligand complexes)
- A unified analysis framework that computes metrics akin to complicated LDDT and DockQ throughout fashions
The related PXMeter analysis paper, ‘Revisiting Construction Prediction Benchmarks with PXMeter,‘ evaluates Protenix, AlphaFold3, Boltz-1 and Chai-1 on the identical curated duties, and exhibits how totally different dataset designs have an effect on mannequin rating and perceived efficiency.
How Protenix matches into the broader stack?
Protenix is a part of a small ecosystem of associated tasks:
- PXDesign: a binder design suite constructed on the Protenix basis mannequin. It stories 20–73% experimental hit charges and 2–6× greater success than strategies akin to AlphaProteo and RFdiffusion, and is accessible through the Protenix Server.
- Protenix-Dock: a classical protein–ligand docking framework that makes use of empirical scoring features slightly than deep nets, tuned for inflexible docking duties.
- Protenix-Mini and follow-on work akin to Protenix-Mini+: light-weight variants that scale back inference price utilizing architectural compression and few-step diffusion samplers, whereas conserving accuracy inside a couple of p.c of the complete mannequin on commonplace benchmarks.
Collectively, these parts cowl construction prediction, docking, and design, and share interfaces and codecs, which simplifies integration into downstream pipelines.
Key Takeaways
- AF3-class, absolutely open mannequin: Protenix-v1 is an AF3-style all-atom biomolecular construction predictor with open code and weights below Apache 2.0, focusing on proteins, DNA, RNA and ligands.
- Strict AF3 alignment for honest comparability: Protenix-v1 matches AlphaFold3 on vital axes: coaching information cutoff (2021-09-30), mannequin scale class and comparable inference finances, enabling honest AF3-level efficiency claims.
- Clear benchmarking with PXMeter v1.0.0: PXMeter supplies a curated benchmark suite over 6k+ complexes with time-split and domain-specific subsets plus unified metrics (for instance, complicated LDDT, DockQ) for reproducible analysis.
- Verified inference-time scaling conduct: Protenix-v1 exhibits log-linear accuracy positive aspects because the variety of sampled candidates will increase, giving a documented latency–accuracy trade-off slightly than a single mounted working level.
Take a look at the Repo and Strive it right here. Additionally, be at liberty to observe us on Twitter and don’t neglect to affix our 100k+ ML SubReddit and Subscribe to our Publication. Wait! are you on telegram? now you may be part of us on telegram as properly.




