Genome-wide profiling of histone modifications in mouse embryonic stem cells
To systematically examine the regulatory patterns of univalent and bivalent histone modifications, we in contrast the sign intensities and chromosomal distribution preferences of various histone marks. Amongst them, H3K4me3 displayed globally increased enrichment ranges in mESCs in comparison with the 2 repressive modifications, H3K27me3 and H3K9me3. These marks have been comparatively evenly distributed throughout autosomes, whereas fewer peaks have been noticed on intercourse chromosomes, suggesting that the institution of those modifications is basically secure throughout the genome (Fig. 1A). To additional dissect the combinatorial patterns of H3K4me3 with both H3K27me3 or H3K9me3, we categorized the peaks into monovalent, bivalent, and trivalent classes. Nearly all of peaks have been marked by a single histone modification, whereas co-modified areas—these carrying two or extra marks—constituted a smaller fraction of the full (Fig. 1B). Notably, areas concurrently marked by H3K4me3, H3K27me3, and H3K9me3 (trivalent domains) accounted for about 1% of H3K4me3 peaks, 8% of H3K27me3 peaks, and three% of H3K9me3 peaks.
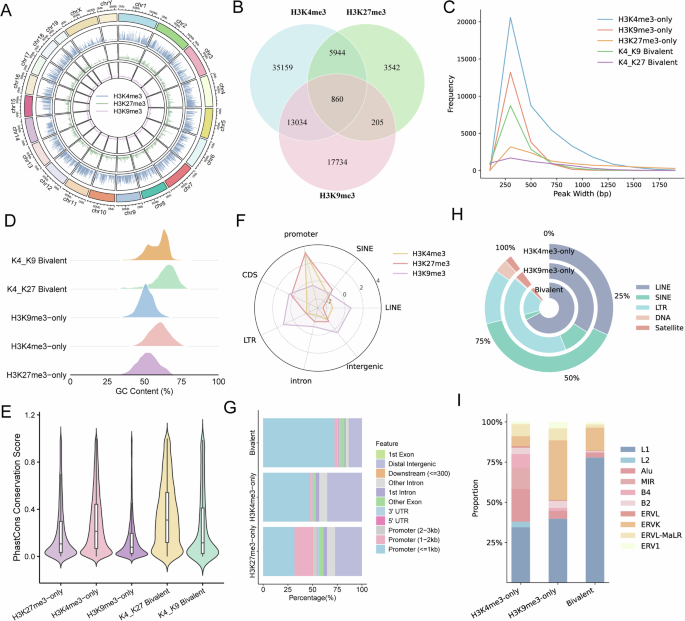
A Circos plot exhibiting the H3K4me3, H3K27me3, and H3K9me3 peak alerts throughout all chromosomes. B Venn diagram illustrating the genomic overlap amongst H3K4me3, H3K27me3, and H3K9me3 peaks. The noticed three-way overlap is considerably increased than anticipated below a random genomic null mannequin (imply ± SD = 2.7 ± 1.7; empirical p < 0.01), indicating a non-random co-occurrence of histone modification patterns. C Distribution of peak widths for monovalent (H3K4me3-only, H3K27me3-only, H3K9me3-only) and bivalent modifications. D GC content material distribution of genomic areas related to totally different histone modifications. E PhastCons conservation scores for monovalent and bivalent peaks, reflecting evolutionary constraint. F Radar plot exhibiting enrichment rating (log ratio of noticed/random) of H3K4me3, H3K27me3, and H3K9me3 peaks throughout varied genomic parts, together with promoters, coding sequences (CDS), introns, intergenic areas, and main transposable factor (TE) courses (LINE, LTR, and SINE). G Genomic annotation of H3K4me3-only, H3K27me3-only, and bivalent peaks throughout useful options (e.g., promoters, exons, introns, downstream areas). H Proportional overlap of histone modification peaks with main TE courses, together with LINE, SINE, LTR, DNA, and satellite tv for pc parts. I TE household composition (e.g., L1, B2, ERVK, and Alu) related to H3K4me3-only, H3K9me3-only, and bivalent peaks.
We subsequent investigated the underlying DNA sequence traits of monovalent versus bivalent peaks, specializing in peak width, GC content material, and conservation. Peak size evaluation revealed that almost all areas spanned 200 to 1000 bp, with bivalent peaks being barely narrower than monovalent ones (Fig. 1C). Moreover, bivalent peaks exhibited considerably increased GC content material and conservation scores, suggesting that these co-modified areas are preferentially positioned in conserved genomic parts (Fig. 1D, E).
Bivalent domains are sometimes discovered at promoter areas, as exemplified by the overlap of H3K4me3 and H3K27me3 enrichment at gene promoters (Fig. 1F, G). In distinction, H3K9me3 confirmed predominant enrichment inside lengthy terminal repeat (LTR) and lengthy interspersed nuclear factor (LINE) repetitive parts (Fig. 1F), according to its established position in retrotransposon silencing29. We additionally evaluated the enrichment of repetitive DNA parts inside H3K9me3/H3K4me3 bivalent domains. The outcomes confirmed that H3K9me3-marked bivalent areas have been considerably enriched in repetitive parts in mESCs, with H3K9me3/H3K4me3 domains significantly enriched for LINE/L1 parts (Fig. 1H, I).
Bivalent modifications coordinately regulate gene expression
To elucidate the results of distinct histone modification (HM) patterns on gene expression, we quantitatively assessed the sign intensities of H3K4me3, H3K27me3, and H3K9me3 at trivalently marked genes and correlated these with corresponding transcriptional ranges. Our evaluation revealed that elevated H3K4me3 alerts are strongly related to excessive gene expression, whereas enrichment of H3K27me3 or H3K9me3 correlates with diminished expression. These findings counsel that inside co-marked genomic areas, H3K4me3 seemingly capabilities because the principal activation mark, whereas H3K27me3 and H3K9me3 seem to contribute predominantly to repressive regulatory outcomes (Fig. 2A).
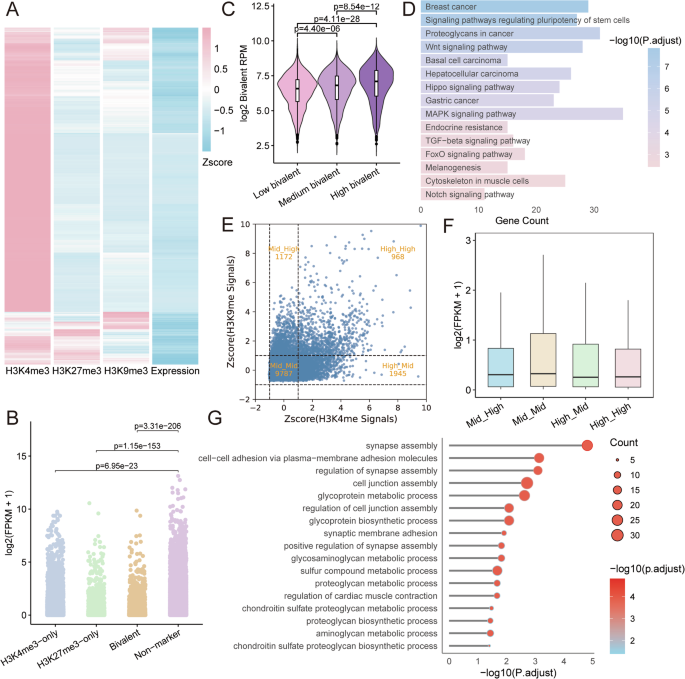
A Heatmap exhibiting the Z-score-normalized sign intensities of H3K4me3, H3K27me3, and H3K9me3, together with expression ranges of co-marked genes in mESCs. B Gene expression ranges (log₂(FPKM + 1)) related to the 4 chromatin states. n = variety of genes; two-sided Wilcoxon rank-sum check. C Bivalent histone modification RPM alerts throughout bivalent genes with low, medium, or excessive expression ranges. n = variety of genes; two-sided Wilcoxon rank-sum check. D Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment of extremely expressed bivalent genes (FDR < 0.05). E 9-quadrant scatter plot of Z-score-normalized H3K4me3 and H3K9me3 alerts at overlapping peaks, exhibiting 4 chromatin teams based mostly on sign ranges: Excessive-Excessive, Mid-Excessive, Excessive-Mid, and Mid-Mid. F Gene expression ranges related to the signal-defined chromatin teams in Fig. 2E. G GO organic course of (BP) enrichment of H3K4me3/H3K9me3 Excessive-Excessive bivalent genes (FDR < 0.05).
We additional analyzed the expression patterns of genes related to the 2 varieties of bivalent modifications (H3K4me3/H3K27me3 and H3K4me3/H3K9me3), and carried out useful enrichment analyses. Genes marked by H3K4me3/H3K27me3 bivalency exhibited comparatively low expression ranges, falling between these marked by H3K4me3-only and H3K27me3-only areas, however skewed nearer to the H3K27me3-only profile, indicating a basic tendency towards transcriptional repression (Fig. 2B). We additional categorized bivalent genes into excessive, medium, and low expression teams and noticed a optimistic relationship between bivalent mark depth and gene expression, with extremely expressed genes exhibiting the strongest bivalent alerts (Fig. 2C). Persistently, a genome-wide correlation evaluation revealed a weak however statistically vital optimistic affiliation between bivalent sign depth and gene expression (Spearman’s ρ = 0.186, p = 9.33 × 10−³⁴; Supplementary Fig. S1), in settlement with earlier research30,31. KEGG pathway enrichment evaluation of extremely expressed bivalent genes revealed vital affiliation with the “breast cancer” pathway, implying that some bivalent genes might contribute to oncogenesis. For instance, the bivalent gene POU4F1 has been proven to be important for selling and sustaining basal-like breast most cancers (BLBC), making it a possible therapeutic goal32 Bivalent genes are enriched in majority of the signaling pathways associated to the pluripotent upkeep and improvement, corresponding to signaling pathways regulating pluripotency of stem cells, TGF-beta, Wnt, MAPK, FoxO and Hippo signaling pathways33 (Fig. 2D).
Subsequent, we tried to outline bivalent genes with totally different sign intensities by overlapping H3K4me3 and H3K9me3 Chromatin Immunoprecipitation sequencing (ChIP-seq) peaks. The overlapping areas have been categorized into 4 teams based mostly on sign energy: Excessive-Excessive, Mid-Excessive, Excessive-Mid, and Mid-Mid. These teams exhibited a linear distribution sample, suggesting a sure diploma of co-regulation between H3K4me3 and H3K9me3 (Fig. 2E). By integrating RNA-seq knowledge, we discovered that genes positioned in areas with each excessive H3K4me3 and excessive H3K9me3 alerts (Excessive-Excessive) exhibited comparatively low expression ranges (Fig. 2F). Bivalent domains are recognized to endure dynamic transitions throughout cell destiny specification. For example, modifications in bivalent marks are implicated within the differentiation of neural stem cells34. Gene Ontology (GO) evaluation of H3K4me3/H3K9me3-associated genes revealed enrichment in neurodevelopmental processes corresponding to synapse meeting—a essential step throughout ESC differentiation into mature and useful neurons (Fig. 2G).
Interpretable mannequin of bivalent histone modifications pushed by sequence options
In embryonic stem cells, bivalent domains marked by H3K4me3 and H3K27me3 are preferentially enriched at promoters containing CpG islands3 and exhibit robust associations with particular DNA sequence options, corresponding to CpG-rich parts and transcription issue binding websites35. A earlier research recognized an enrichment of the TCCCC sequence motif at bivalent promoters in each mouse and human ESCs36. However, the sequence determinants that encode or direct the formation of bivalent histone modifications stay largely elusive. To discover whether or not bivalent chromatin areas harbor distinctive and predictive sequence options, we employed 5 supervised machine studying algorithms in addition to deep studying fashions to categorise bivalent versus monovalent areas. Particularly, overlapping bivalent peaks (200-1000 bp in size) have been chosen for sequence function extraction. For machine studying fashions, we used the broadly adopted ok-mer method, which has confirmed efficient in capturing brief sequence patterns related to chromatin state21. Insights from earlier analysis have been leveraged to information the number of parameter ok, and a variety of ok values from 3 to eight was adopted on this research19,37,38. For deep studying fashions, sequences have been encoded utilizing one-hot encoding, enabling the fashions to mechanically study latent sequence patterns (Fig. 3).
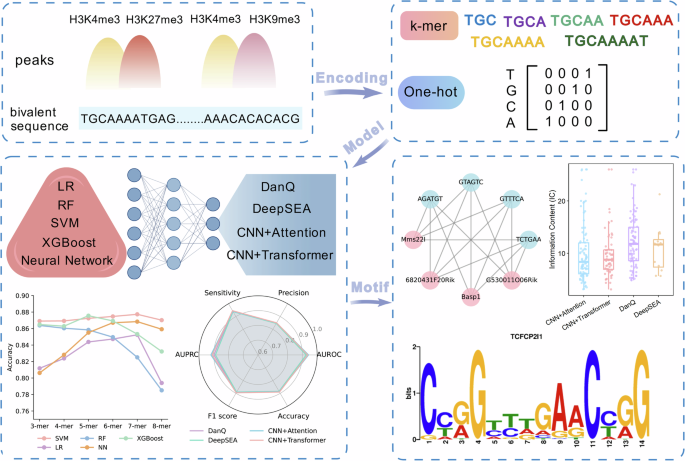
This diagram summarizes the computational workflow employed on this research. First, genomic areas marked by H3K4me3, H3K27me3, and H3K9me3 have been extracted and reworked into numerical representations utilizing both ok-mer options or one-hot encoding. Subsequent, a set of machine-learning fashions (logistic regression, SVM, random forest, XGBoost, and neural networks) and deep-learning architectures (DanQ, DeepSEA, CNN+Consideration, and CNN+Transformer) have been educated to tell apart bivalent from non-bivalent sequences. Mannequin efficiency was assessed utilizing accuracy, AUROC, AUPRC, sensitivity, precision, and F1-score. Lastly, motifs realized by the deep fashions have been interpreted through activation mapping and visualized by motif logos and transcription-factor interplay networks.
We first constructed a three-class classification mannequin to tell apart amongst H3K4me3-only, H3K9me3-only, and H3K4me3/H3K9me3 bivalent areas. Primarily based on comparative evaluation between Lasso regression and Random Forest for function choice, we adopted Random Forest, which recognized 797 informative 6-mer options (Supplementary Fig. S2). Analysis throughout totally different ok-mer sizes revealed that every one fashions carried out finest at ok = 6 (Fig. 4A), according to prior machine studying research on genomic sequence prediction18,20,39,40. Amongst all fashions examined, Assist Vector Machine (SVM) yielded the best classification efficiency, as indicated by ROC curves and total accuracy (Fig. 4B, C). The confusion matrix of the SVM mannequin confirmed robust classification capability, significantly for H3K4me3-only areas (Fig. 4D). To exclude the chance that the noticed predictive efficiency arose from random correlations, we additional evaluated a randomized null mannequin wherein sequence options have been shuffled whereas class labels have been preserved. Beneath this management, all fashions exhibited chance-level efficiency (AUROC ≈ 0.5), in distinction to the clear diagonal enrichment noticed when utilizing actual genomic sequences (Supplementary Fig. S5), indicating that the predictive alerts captured by the fashions are genuinely encoded in DNA sequence options. Utilizing the same technique, we additional distinguished H3K4me3/H3K27me3 bivalent areas from monovalent ones. Lasso regression was used to pick out 616 related sequence options. Optimum mannequin parameters have been decided through grid search with 5-fold cross-validation. Amongst all examined classifiers, the XGBoost-6mer mannequin achieved one of the best predictive efficiency (Supplementary Fig. S3). Apparently, H3K4me3/H3K9me3 bivalent areas have been categorized extra precisely than H3K4me3/H3K27me3 areas, indicating that they could harbor extra distinct sequence determinants.
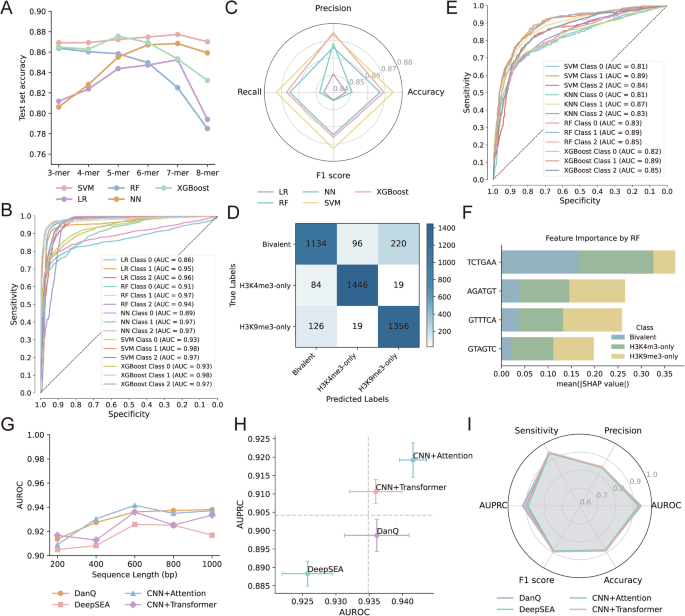
A Prediction check set accuracy of various machine studying fashions (SVM, Random Forest, XGBoost, Logistic Regression, Neural Community) throughout varied ok-mer sizes (3-mer to 8-mer) in classifying H3K4me3-only, H3K9me3-only, and bivalent areas. B ROC curves illustrating the efficiency of every mannequin on the check set in distinguishing bivalent from monovalent (H3K4me3 or H3K9me3) areas. C Radar plot summarizing precision, recall, F1 rating, and accuracy for every machine studying mannequin on the check set. D Confusion matrix of the SVM classifier exhibiting prediction efficiency throughout three courses: H3K4me3-only, H3K9me3-only, and bivalent. E ROC curves of check set predictions utilizing fashions educated on 4 consultant enriched sequence options derived from bivalent areas. F SHAP-based function interpretation of the Random Forest mannequin, indicating the contribution of 4 particular sequence options (ok-mers) to distinguishing bivalent, H3K4me3-only, and H3K9me3-only areas. G AUROC efficiency of deep studying fashions (DeepSEA, DanQ, CNN+Consideration, CNN+Transformer) educated on enter sequences of various lengths (200–1000 bp). H Comparability of AUROC and AUPRC scores of deep studying fashions utilizing 600 bp enter sequences. I Radar chart summarizing total efficiency metrics (accuracy, precision, sensitivity, F1 rating, AUROC, and AUPRC) of deep studying fashions on the check set.
Given the big variety of sequence options recognized by machine studying, mannequin interpretation stays difficult resulting from function complexity and potential redundancy. To deal with this, we carried out HOMER enrichment evaluation to prioritize core sequence motifs with potential organic relevance (Supplementary Fig. S6B). For the H3K4me3/H3K9me3 bivalent modifications, 4 key motifs have been considerably enriched. We retrained classification fashions utilizing solely these core options and noticed that every one fashions maintained strong predictive efficiency, with AUC values reaching roughly 0.87. Amongst them, the Random Forest (RF) mannequin persistently outperformed others, suggesting that these core sequence motifs are ample to seize important info for chromatin state classification (Fig. 4E). SHAP (SHapley Additive exPlanations) evaluation revealed that the motif “TCTGAA” exhibited the best optimistic contribution to the prediction of bivalent areas, indicating its robust affiliation with bivalent chromatin states. In distinction, the remaining three motifs predominantly contributed to the classification of H3K9me3-only areas. These findings counsel useful divergence amongst sequence motifs, the place distinct brief DNA sequences preferentially mark totally different histone modification states (Fig. 4F). These findings point out that totally different brief DNA motifs preferentially affiliate with distinct chromatin states.
Equally, 5 enriched motifs have been recognized for H3K4me3/H3K27me3 bivalent domains. As anticipated, the efficiency was barely diminished in comparison with fashions educated with the total function set, but the simplified mannequin gives improved interpretability and highlights biologically related sequence patterns (Supplementary Fig. S4). Primarily based on SHAP values, the motifs TCACAG, TTCAAA, and AGGGCT exhibited the best function significance, serving as key indicators for distinguishing H3K4me3/H3K27me3 bivalent areas from monovalent counterparts (Supplementary Fig. S4C). Collectively, these outcomes spotlight that particular brief DNA sequence motifs not solely carry predictive energy for chromatin state classification but in addition present mechanistic insights into the sequence determinants underlying bivalent histone modifications.
To achieve deeper insights into the sequence determinants regulating bivalent chromatin areas, we constructed deep studying fashions to foretell H3K4me3/H3K9me3 bivalency. We systematically evaluated the impact of various enter sequence lengths (200 bp, 400 bp, 600 bp, 800 bp, and 1000 bp) on mannequin efficiency. General, most fashions exhibited improved AUROC values with growing sequence size. Particularly, the DanQ, CNN+Consideration, and CNN+Transformer fashions achieved optimum efficiency when educated on sequences of 600 bp or 1000 bp (Fig. 4G), suggesting that these lengths are well-suited for this binary classification process. Amongst all evaluated fashions, the CNN+Consideration structure persistently demonstrated one of the best predictive efficiency, adopted by the DanQ and CNN+Transformer fashions, based mostly on complete analysis metrics (Fig. 4H, I). These outcomes counsel that deep studying fashions are able to capturing higher-order and context-dependent sequence options related to bivalent chromatin, with interpretable architectures additional facilitating motif discovery.
Evaluation of sequence regulatory grammar underlying bivalent histone modifications
To elucidate the regulatory grammar of distinct bivalent histone modifications, we carried out a comparative evaluation of sequence options related to H3K4me3/H3K9me3 and H3K4me3/H3K27me3 areas. Histone modification degree was reported to be carefully associated to the general GC content material of the corresponding area41, we categorized the recognized motifs into three teams based mostly on their GC content material: GC-rich, AT-rich, and impartial motifs. Most motifs have been GC-rich, whereas a smaller proportion have been AT-rich (Fig. 5A). Earlier research have implied the position of specific DNA sequences (or motifs) in setting the boundary of histone modification area or opening chromatin to permit transforming enzymes to bind DNA42. We subsequent examined the positional distribution of the chosen motifs inside bivalent sequences. Every bivalent area was divided into ten equal bins, with the outermost bins representing the sides and the central bins representing the middle. We noticed that motifs related to H3K4me3/H3K27me3 bivalency have been predominantly enriched on the edges of bivalent areas, whereas these linked to H3K4me3/H3K9me3 bivalency have been largely categorized as impartial. These outcomes point out that the 2 varieties of bivalency exhibit distinct motif positional preferences inside bivalent domains (Fig. 5B; Supplementary Fig. S6A).
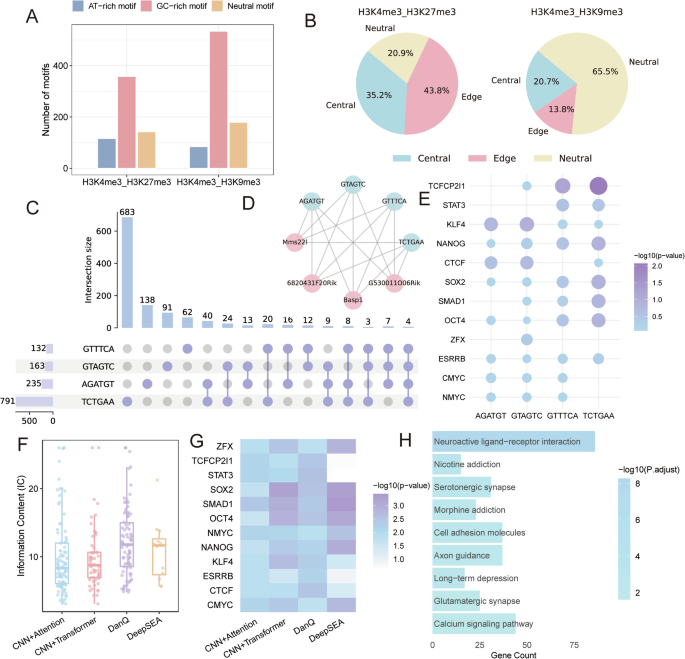
A Grouped bar plot exhibiting the variety of GC-rich and AT-rich 6-mer motifs recognized in H3K4me3/H3K9me3 and H3K4me3/H3K27me3 areas. B Pie charts exhibiting the positional preferences of motifs (central, edge, impartial; particulars in Motif Evaluation) inside bivalent peaks. C UpSet plot of bivalent genes related to 4 enriched motifs in H3K4me3/H3K9me3 areas. D Chord diagram exhibiting motif-gene associations in H3K4me3/H3K9me3 areas. E Bubble plot of transcription components considerably enriched for the 4 consultant motifs. F Boxplot of data content material (IC) scores for motifs recognized by CNN+Consideration, CNN+Transformer, DanQ, and DeepSEA fashions. G Heatmap of transcription issue matches for motifs predicted by every deep studying mannequin. H KEGG pathway enrichment evaluation of bivalent genes related to DanQ-predicted motifs (FDR < 0.05).
Subsequent, we additional examine potential transcription components (TFs) related to these sequence options, we used the TomTom device to match all recognized motifs with a mouse motif database. Pluripotency-related transcription components, together with STAT3, ESRRB, KLF4, OCT4, SOX2, and SMAD1, have been considerably enriched in each varieties of bivalent areas (Supplementary Fig. S6), suggesting shared regulatory packages in embryonic stem cells. Notably, the important thing motif TCTGAA that distinguishes H3K4me3/H3K9me3 bivalent areas was primarily matched to the TF TCFCP2l1 (Fig. 5E). These findings counsel that the DNA sequences co-occupied by H3K4me3/H3K27me3 or H3K4me3/H3K9me3 are doubtlessly regulated by pluripotency components in embryonic stem cells. We additional mapped the recognized sequence options of H3K4me3/H3K9me3 bivalent modifications to particular peak areas, annotated the corresponding genes, and recognized 4 bivalent genes containing all 4 key motifs (Fig. 5C, D). These peaks exhibited intermediate ChIP-seq sign intensities between H3K4me3-only and H3K9me3-only areas (Supplementary Fig. S6E), supporting the notion of a definite chromatin state.
To interpret the realized sequence patterns, we utilized a sequence alignment-based interpretability technique (SABM) to H3K4me3/H3K9me3-labeled areas. This technique identifies subsequences that activate convolutional neurons and stacks place weight matrices (PWMs), analogous to conventional estimation strategies43. We used info content material (IC) to judge every mannequin’s skill to establish useful sequence motifs, with the DanQ mannequin reaching the best IC values, indicating stronger motif definition on this subtype (Fig. 5F). Utilizing the TOMTOM device, we recognized the transcription components related to motifs realized by totally different fashions. These transcription components are able to binding not solely to the promoters of pluripotency-related genes in embryonic stem cells, but in addition to their enhancer areas, thereby sustaining the pluripotent state (Fig. 5G). Subsequently, we utilized the FIMO device to find the precise positions of every motif (motif websites), and annotated the corresponding bivalent genes. KEGG pathway enrichment evaluation of those bivalent genes revealed vital enrichment within the Neuroactive ligand–receptor interplay pathway, which is carefully related to nervous system improvement and stem cell differentiation (Fig. 5H).
In abstract, whereas H3K4me3/H3K9me3 and H3K4me3/H3K27me3 bivalent areas share widespread pluripotency-related regulatory components, they differ in base composition, motif positioning, TF specificity, and related organic pathways. These findings counsel that distinct sequence grammars underlie the 2 courses of bivalent chromatin, reflecting their divergent regulatory roles in embryonic stem cells.