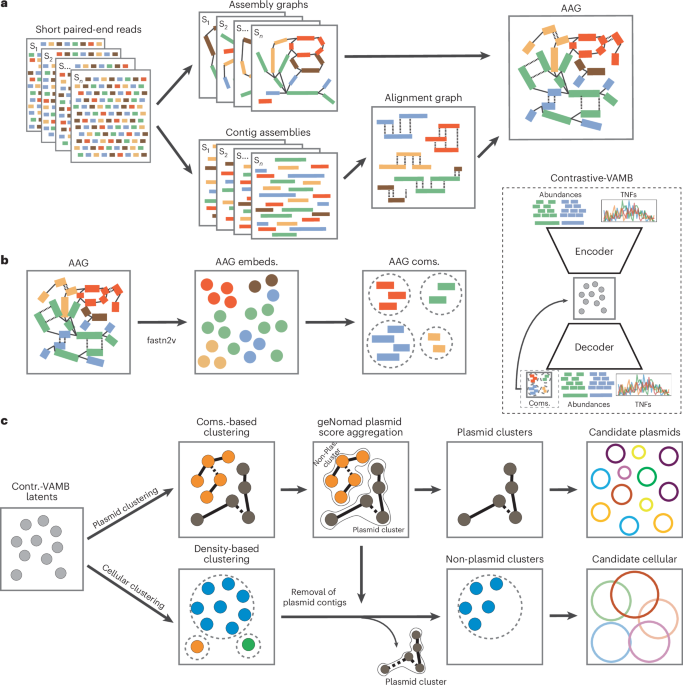
Overview of PlasMAAG
The inputs to the PlasMAAG pipeline are a set of reads per pattern. Reads are assembled with metaSPAdes (model 3.15.5)42, creating an meeting graph and contigs for every pattern. The contigs throughout all samples are concatenated collectively to create the contig catalog. Reads are mapped to the catalog with minimap2 (model 2.24)43 and SAMtools (model 1.18)44, creating per-sample BAM recordsdata. The alignment graph is generated by aligning the contigs throughout samples with the Nationwide Middle for Biotechnology Info (NCBI) fundamental native alignment search instrument (BLAST; model 2.15.0)45. The meeting and alignment graphs are merged into the AAG. Then, fastnode2vec (model 0.05)35, an optimized model of node2vec46, is used to embed native AAG context of every contig into an embedding house. The okay-mer composition and abundance options of contigs are embedding utilizing a VAE, the place an extra loss time period is added, which penalizes distance between contigs of the identical neighborhood. Utilizing the VAE embeddings, communities are expanded, merged and purified. The geNomad instrument (model 1.8.0)36 is used to separate plasmid from nonplasmid contigs; communities of plasmid contigs are extracted as separate bins, whereas the remaining contigs are extracted in bins utilizing a clustering algorithm.
Benchmark datasets
We primarily based our benchmark datasets on the prevailing CAMI2 human microbiome toy dataset, the CAMI2 marine dataset and the CAMI2 pores and skin human microbiome dataset with addition of phage reads from 280 phage genomes from the Metagenomic Intestine Virus (MGV) database47 (Supplementary Desk 5). All CAMI2 datasets are publicly accessible short-read metagenomic benchmarks generated below the CAMI initiatives37,48. The human dataset consists of 5 simulated environments: airways, gastrointestinal, oral, pores and skin and urogenital, designed to imitate human microbiome communities37, whereas the CAMI2 marine dataset represents a sea atmosphere48. For every pattern, we simulated 5 Gbp of two× 150-bp Illumina-like reads from the supply genomes utilizing the CAMISIM software program49. In comparison with the unique CAMI2 datasets, we needed to make a number of adjustments. The unique dataset didn’t present meeting graphs; thus, we assembled the reads utilizing metaSPAdes (model 3.15.5) and mapped the ensuing contigs again to the CAMI2 supply genomes or the MGV phage genomes to find out their origin utilizing minimap2, accepting hits with an identification > 97% and a question protection > 90%. As a result of this method initially led to many unmapped or ambiguously mapping contigs due to overeager learn correction within the assembler, we resimulated the reads utilizing wgsim ( with zero sequencing errors and a default fragment measurement of 270 (imply)26 and disabled error correction within the assembler. Furthermore, CAMI2 thought-about plasmids to be a part of their mobile host genome with the identical abundance, which might inhibit our abundance-based binning method. As an alternative, we simulated plasmids as distinct genomes with their very own abundance proportional to the host abundance, modulated by a Gaussian random variable that includes a plasmid copy quantity mannequin, as carried out beforehand17. For the samples containing phages, phage abundances have been sampled from a lognormal distribution with imply of 1 and s.d. of 4. Lastly, CAMI2 didn’t include reads simulated from throughout the sides of the underlying round sequences, which prevents meeting graph cycles and hobbles graph peeling-based approaches similar to that utilized by SCAPP. We made positive to incorporate such reads and allow round contigs by simulating reads spanning the junction between the beginning and finish positions of the genomes. Our reassembled CAMI2 human and marine datasets have been, like the unique CAMI2 datasets, composed of ten samples for all datasets besides urogenital, which had 9. Our datasets had the next variety of contigs after retaining solely contigs bigger than 2 kbp: airways, 35,537; gastrointestinal, 80,325; oral, 94,251; pores and skin, 31,181; urogenital, 29,550; marine, 150,565; pores and skin with phages, 34,979.
Meeting graph edge weighting
Meeting graphs have been extracted from the assembly_graph_after_simplification.gfa file from metaSPAdes and transformed right into a NetworkX (model 3.4.2)50 directed graph, with contigs represented as nodes and hyperlinks between segments in contigs represented as edges. To counterpoint the meeting graph sign for binning, graph edges have been weighted with the normalized linkage metric, which depends on the variety of hyperlinks between any segments from every pair of contigs, normalized by the size of the contigs. For a pair of contigs ci, cj, the variety of hyperlinks connecting these contigs n_linksij and the contig lengths lc, normalized linkage is expressed as
$$mathrm{normalized},{mathrm{linkage}}_{{c}^{i}{c}^{j}}=frac{{nmathrm{_links}}_{{ij}}}{min ({l}^{{c}^{i}},{l}^{{c}^{j}})}$$
Alignment graph edge weighting
After meeting, contigs shorter than 2,000 bp have been discarded, as described beforehand24. Contigs have been aligned all towards all utilizing NCBI BLAST utilizing the ‘blastn’ command with ‘-perc_identity 95’, solely conserving between-sample hits, alignment identification ≥ 98.0% and alignment ≥ 500 bp. We additionally eliminated alignments between sequences that contained massive sections that didn’t align due to sequence variety, as we wished the alignments to characterize shared sequences throughout samples. The remaining set of alignments after filtering was outlined as ‘restrictive’ alignments. From the alignments, we created an alignment graph with contigs as nodes and alignments as edges. Edges have been weighted with the normalized alignment metric to mirror the alignment certainty. For a pair of contigs ci, cj, alignment identification id, alignment size L and contig size lc, normalized alignment is expressed as
$$mathrm{normalized},{mathrm{alignment}}_{{c}^{i}{c}^{j}}=frac{mathrm{id}}{100}frac{min (L,{l}^{{c}^{i}},{l}^{{c}^{j}})}{min ({l}^{{c}^{i}},{l}^{{c}^{j}})}$$
AAG neighborhood extraction with fastnode2vec
Meeting and alignment graphs share no edges as a result of their edges join solely within-sample and between-sample contigs, respectively. This allowed us to trivially merge the graphs by including the sides from one graph into the opposite, thus creating the AAG. Thus, the AAG is a single graph that accommodates the nodes and edges from the person meeting graphs and the alignment graph. We aimed to leverage this built-in graph to reinforce the binning sign, whereas conserving all particular person nodes (that’s, contigs) intact and unchanged. To extract communities from the AAG, we first ran fastnode2vec on the AAG to acquire contig embeddings. We created a brand new graph by linking contigs inside a cosine distance of 0.1 in embedding house, after which we outlined every linked part to be a contig neighborhood. We optimized the fastnode2vec hyperparameters and clustering radius to generate pure communities on the genome degree, operating a small grid search over the resimulated CAMI2 airways dataset. The embedding dimensions, stroll size, variety of walks, window measurement, p and q parameters from fastnode2vec have been set to 32, 10, 50, 10, 0.1 and a pair of.0, respectively.
Contrastive-VAMB for neighborhood merging and enlargement
Contrastive-VAMB is a variation of the unique VAMB mannequin, with a modification on the loss operate to account for the communities extracted from the fastnode2vec embeddings. Contrastive-VAMB consists of an encoder, latent illustration layer μ and a decoder. Every contig represented by the concatenation of the contig coabundances alongside samples Cin, the tetranucleotide frequencies Tin and the unnormalized contig abundances Ain is handed to the encoder. Whereas Cin and Tin have been already used within the unique VAMB mannequin, the inclusion of Ain is motivated by the remark that the normalization within the computation of Cin removes abundance sign by eradicating a level of freedom. This sign is reintroduced as Ain. This fashion, the community explicitly fashions two distinct notions of abundance: how a lot abundance there’s in whole and the way it’s distributed amongst samples. The encoder tasks the contigs right into a latent regular N(μ, I) distribution parametrized by the μ layer, from which the decoder samples. The decoder is optimized to reconstruct Cin, Tin and Ain from the cases sampled from N(μ, I), lower the latent cosine distance between contigs with carefully associated fastnode2vec graph embeddings and reduce the deviance between the latent regular distribution N(μ, I) parametrized by the μ layer and the usual regular distribution used as a previous N(0, I).
Loss capabilities
The contrastive-VAMB loss will be decomposed in three phrases: reconstruction loss, contrastive loss and regularization loss. The reconstruction loss (Lrec) penalizes the reconstruction error of Ain, Tin, and Cin. Like in VAMB, the sum of squared error (SSE) loss was used for Tin and cross-entropy (CE) loss was used for Cin. For Ain, PlasMAAG additionally makes use of SSE loss. These three phrases are weighted with hyperparameters wA, wT and wC.
$${L}_{mathrm{rec}}={w}_{{bf{A}}}mathrm{CE}left({{bf{A}}}_{mathrm{in}},{{bf{A}}}_{mathrm{out}}proper)+{w}_{{bf{T}}}mathrm{SSE}left({{bf{T}}}_{mathrm{in}},{{bf{T}}}_{mathrm{out}}proper)+{w}_{{bf{C}}}mathrm{SSE}left({{bf{C}}}_{mathrm{in}},{{bf{C}}}_{mathrm{out}}proper)$$
The contrastive loss (Lcontr) penalizes the cosine distance between the VAMB-latent representations contigs extremely associated in fastnode2vec embedding house when such cosine distance overcomes a predefined margin m, the place m is a hyperparameter. For a contig ci and extremely associated fastnode2vec embedding house contigs Hci = {n0,…, nn}, Lcontr is expressed as
$${L}_{mathrm{contr}}=max left(frac{{sum }_{{n}_{i}in {H}^{ci}}mathrm{cosine},mathrm{distance}left({{rm{m}}}^{ci},{{rm{m}}}^{{n}_{i}}proper)}{left|{H}^{ci}proper|}-m,0right)$$
The regularization loss (Lreg) penalizes the deviance between the latent regular distribution N(μ, I) parametrized by the μ layer and the usual regular distribution used as a previous N(0, I) with the Kullback–Leibler divergence; because the s.d. is about to 1, Lreg will be simplified to
$${L}_{mathrm{reg}}=frac{1}{2}+sum {{rm{mu }}}^{2}$$
Lastly, the mannequin whole loss (L) was aggregated with weighting hyperparameters wLreg, and wLcontr:
$$L={L}_{mathrm{rec}}+{{w}_{{L}_{mathrm{reg}}}L}_{mathrm{reg}}+{{w}_{{L}_{mathrm{contr}}}+L}_{mathrm{contr}}$$
Clustering plasmid and organism candidates with geNomad
Two sequential methods have been carried out to cluster the latent house tailor-made to extract plasmids and nonplasmids. The clustering technique consists of two phases: neighborhood clustering and iterative medoid clustering, each utilizing latent house cosine distances. Group clustering works in 5 steps (Supplementary Fig. 11). First, for every neighborhood extracted from the fastnode2vec embeddings, contigs belonging to the identical neighborhood are linked and hyperlinks between contigs with a VAE embedding cosine distance > 0.2 are eliminated. Second, contigs inside 0.01 cosine distance are linked. Third, linked elements are extracted as bins. Fourth, doubtlessly round contigs are detected by mapping learn pairs, the place mates map to reverse contig ends inside 50 bp of the contig finish. These are extracted to their very own bins. Lastly, plasmid rating is outlined for every cluster by aggregating the geNomad plasmid contig scores with a contig size weighted imply, defining plasmid candidates when cluster scores are bigger than the outlined threshold. When the geNomad plasmid threshold is bigger than 0.5, a set geNomad plasmid threshold of 0.5 is utilized to the round contigs, accounting for the round proof relatable to plasmids. The nonplasmid clustering technique consists of two steps. First, the VAMB-latent house is clustered utilizing the iterative medoid clustering algorithm from VAMB. Second, contigs belonging to any plasmid candidate cluster as outlined by community-based clustering are eliminated. In abstract, candidate plasmids are first recognized utilizing community-based clustering mixed with geNomad cluster scoring. Contigs belonging to those plasmid bins are then faraway from the bins generated by the density-based clustering technique.
Binning benchmarking: CAMI2 reassembled
We in contrast the plasmid and organism binning efficiency of PlasMAAG model 1.0.0, VAMB model 4.1.3, CONCOCT model 1.1.0, MetaBAT2 model 2.12.1, SemiBin2 model 2.1.0, Comebin model 1.0.4, MetaDecoder model 1.0.19, SCAPP model 0.1.4 and MetaPlasmidSPAdes model 3.15.3 over the resimulated CAMI2 datasets. Binning efficiency was evaluated when it comes to the full variety of genomes recovered throughout all samples with precision ≥95% and recall ≥90% (so-called NC genomes). As PlasMAAG, VAMB, MetaBAT2, SemiBin2, Comebin and MetaDecoder carry out the binning after assembling the contigs, precision and recall of the bins have been obtained from the contig references utilizing BinBencher (model 0.3.0)51. However, SCAPP and MetaPlasmidSPAdes assemble their very own contigs. Right here, we produced a floor fact by aligning the output bins to the origin genomes utilizing NCBI BLAST 2.15.0 accepting hits with an identification >97% and a question protection > 90%, after which we benchmarked utilizing BinBencher.
Pattern benchmarking: CAMI2 reassembled
Plasmid purity price (PPR), plasmid restoration price (PRR) and F1 have been computed for every set of plasmid candidates, reflecting the plasmid characterization on the pattern degree, not on the bin degree. PPR was calculated because the fraction of candidate plasmids reconstructed above precision and recall thresholds within the whole variety of candidate plasmids output by the tactic. PRR was calculated because the fraction of candidate plasmids reconstructed above precision and recall thresholds divided by the full variety of plasmids with protection above the recall threshold. Given a pattern (s), a set of plasmid candidates (candidates), binning precision and binning recall thresholds (pre, rec) and the set of true plasmids current within the pattern (plasmids) are associated by the next expressions:
$$start{array}{l}{mathrm{PPR}}_{mathrm{candidates},mathrm{pre},mathrm{rec}=displaystyle frac{mathrm{no}.,mathrm{of},mathrm{binner},mathrm{plasmid},mathrm{candidates} > (mathrm{pre},,mathrm{rec})}{mathrm{no}.,mathrm{of},mathrm{binner},mathrm{candidates}}}finish{array}$$
$${mathrm{PRR}}_{mathrm{candidates},,mathrm{plasmids},,mathrm{pre},,mathrm{rec}}=frac{mathrm{no}.,mathrm{of},mathrm{binner},mathrm{plasmid},mathrm{candidates} > (mathrm{pre},,mathrm{rec})}{mathrm{no}.,mathrm{of},mathrm{recoverable},mathrm{plasmids},mathrm{in},mathrm{pattern}}$$
Lastly, F1 represents the harmonic imply between the PPR and PRR:
$${{F}_{1}}_{mathrm{PPR},,mathrm{PRR}}=frac{2times mathrm{PPR}occasions mathrm{PRR}}{mathrm{PPR}+mathrm{PRR}}$$
Hospital sewage pattern sequence datasets
Two organic datasets have been used on this research to evaluate the standard of plasmid binning. City sewage samples (UWSs) have been collected from comparable UWSs from Denmark and Spain situated in Odense and Santiago de Compostela, as beforehand described40. On this research, solely hospital sewage samples from every location have been used. Sewage samples have been collected within the winter and summer season of 2018 utilizing ISCO computerized samplers for 24-h stream (50 ml per 5 min) in Denmark and utilizing 24-h time-proportional samples in Spain (mixing hourly samples in accordance with stream data) (Supplementary Desk 8). Three replicates per web site and season have been collected on three consecutive days with out rain. All samples have been initially cooled with ice on web site. Then, 100 ml of every pattern was centrifuged at 10,000g for 8 min at 4 °C within the laboratory. After eradicating supernatant, pellets have been resuspended in 20% of glycerol inventory to achieve a ultimate quantity of 10 ml for storage at −80 °C. In whole, environmental DNA was extracted from all samples utilizing the NucleoSpin soil equipment (Macherey-Nagel) utilizing 500 μl of glycerol inventory materials for direct shotgun metagenomic utilizing Illumina NovaSeq utilizing 2× 150-bp paired-end mode (all 18 samples) and PacBio Sequel2e (5 samples from Denmark). PacBio libraries have been constructed from the identical DNA extracts utilizing libraries utilizing SMRTbell specific template 2.0 equipment and Sequel II binding equipment 3.2 (Pacific Bioscience) and barcoded utilizing SMRTbell barcoded adaptor plate 3.0 (Pacific Bioscience). Two libraries per 8M SMRTcell (Pacific Bioscience) have been pooled and sequenced on a PacBio Sequel2e instrument on the College of Copenhagen.
For plasmid-enriched samples, we used particular strategies to deplete nonplasmid DNA as described beforehand52,53. Briefly, hospital sewage samples have been pretreated by filtration, vortex and sonication and resuspended in TE buffer. Afterward, a prelysis cocktail of cell-wall-degrading enzymes (lysozyme, mutanolysin and lysostaphin) was used to facilitate lysis of Gram-positive micro organism throughout alkaline lysis. Prelysis was adopted by alkaline lysis to take away chromosomal DNA54, adopted by Plasmid-Protected ATP-dependent DNase (Lucigen) digestion. Plasmid-Protected DNase will digest any fragments of dsDNA with open 3′ or 5′ termini, thereby eradicating fragmented chromosomal DNA. The purified plasmid DNA was then quality-checked and libraries have been ready and sequenced on an Illumina NextSeq platform with a v2.5 sequencing equipment (Illumina) in paired-end mode.
Binning benchmarking: hospital sewage
We in contrast the binning efficiency of PlasMAAG model 1.0.0, VAMB model 4.1.3, MetaBAT2 model 2.12.1, SemiBin2 model 2.1.0, Comebin model 1.0.4, CONCOCT model 1.1.0, MetaDecoder model 1.0.19, metaplasmidSPAdes model 3.15.3 and SCAPP model 0.1.4 on the hospital sewage samples. The dataset consisted of 147,437 short-read contigs assembled from 5 hospital sewage samples. Every pattern was collected at a singular mixture of time and site. To judge binning efficiency on these samples, we generated high-quality long-read sequences from the identical samples utilizing HiFi sequencing on a Pacific BioSciences Sequel II platform. We assembled the sequences to long-read contigs with hifiasm_meta model 0.3-r073. To find out which short-read contigs corresponded to long-read contigs, we mapped short-read contigs to long-read contigs utilizing minimap2 model 2.24, dealing with rotated round contigs by concatenating every long-read contig to itself. We accepted hits with an identification > 97% and a question protection > 90%. To quantify short-read binning, we counted ‘NC long-read assemblies’ summed throughout all samples: long-read contigs the place a bin of short-read contigs existed such that 90% of the long-read contig was coated with short-read contigs from that bin and 95% of the bin’s base pairs have been mapped to no different long-read contig. To judge the general binning efficiency, the complete set of long-read contigs was used to construct the reference, whereas, to guage the plasmid binning efficiency, solely the long-read contigs that have been round or with metaplasmidomics reads protection > 50% have been used. Metaplasmidomics brief reads have been derived from the identical samples however underwent a filtering step to counterpoint for plasmids, thereby representing extremely plasmid-enriched sequencing information. NC long-read assemblies have been counted with BinBencher model 0.3.0 and NC mobile genomes have been estimated with CheckM2 model 0.1.3.
Host–plasmid and intraplasmid variety exploration
PlasMAAG was used to bin the contig sequences from hospital sewage samples from hospitals in Spain. The dataset consisted of n = 828,260 contigs assembled from n = 24 hospital sewage samples. Every pattern was collected at a singular mixture of time and site. PlasMAAG bins have been aggregated into PlasMAAG clusters and categorised as plasmids if the aggregated geNomad plasmid rating exceeded 0.75. Solely plasmids clusters with greater than 150 kb have been thought-about for the host–plasmid affiliation. Organism bin high quality was estimated with CheckM2 model 0.1.3 and solely high-quality (completeness ≥ 70% and precision ≥ 90%) bins have been stored. GTDB-tk (model 2.4.0)55 was used to estimate taxonomy for the high-quality bins, with cluster taxonomy assigned on the idea of majority vote. Abundance correlation evaluation was performed for plasmids and organism clusters with nonzero abundance over not less than 18 of 24 overlapping samples. Spearman correlation coefficients and P values have been computed utilizing SciPy’s scipy.stats.spearmanr with default settings, which checks the choice speculation of a nonzero correlation. To account for a number of testing, P values have been corrected utilizing the Benjamini–Hochberg, which makes an attempt to right the false discovery price, carried out within the statsmodels package deal. Plasmid cluster hosts have been inferred from PLSDB (model 2024_05_21_v2)56 when aligning to any PLSDB entry with >80% identification and >80% protection. Purposeful annotations of contigs have been carried out with anvi’o (model 8)57 software program, utilizing the ‘anvi-run-workflow -w contigs’ command.
Useful resource utilization
We evaluated computational useful resource utilization of all strategies utilizing the CAMI2 airways reassembled dataset and 5 samples from the hospital sewage dataset. For the airways dataset, PlasMAAG took 46 min, utilizing 8 threads and 16 GB of RAM. Against this, SCAPP, excluding the BAM file era step, took 192 min, utilizing 16 threads and 24 GB of RAM (Supplementary Desk 6). Among the many different binners, PlasMAAG was slower than VAMB, MetaDecoder and MetaBAT2. For instance, VAMB accomplished the duty in simply 8 min whereas utilizing 8 threads and 16 GB of RAM. Nevertheless, we noticed a unique development when evaluating performances on the 5 hospital sewage samples. When accounting for the extra steps of learn meeting and skim mapping required to compute abundances, PlasMAAG exhibited related runtimes to most binners, aside from SCAPP, which required rather more time. Particularly, PlasMAAG took 3,575 min, VAMB took 3,435 min, ComeBin required 4,911 min and metaplasmidSPAdes took 4,430 min (Supplementary Desk 7). Against this, SCAPP required 116,965 min, 32 occasions longer than PlasMAAG. This distinction in runtime remained constant even when excluding the learn meeting steps (Supplementary Desk 8).
Reporting abstract
Additional data on analysis design is obtainable within the Nature Portfolio Reporting Abstract linked to this text.