Spatiotemporal sampling data
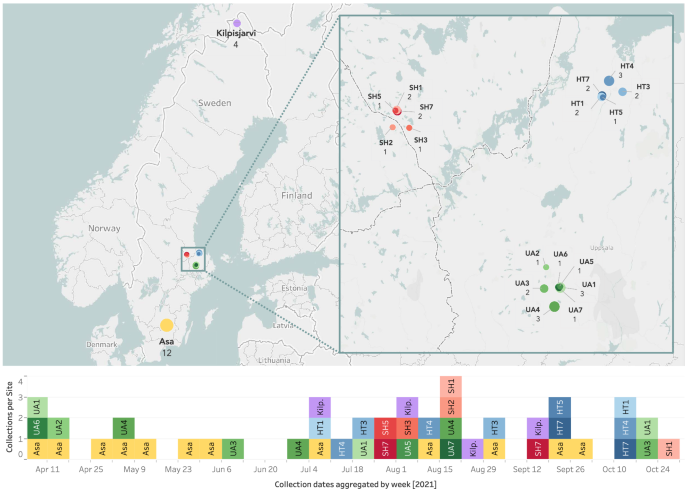
We sampled arthropod communities at 19 websites in Sweden and northern Finland utilizing Townes-style Malaise traps22. We deployed the traps repeatedly throughout 2021 and emptied them as soon as per week. The MassID45 dataset we current right here constitutes a subset of 45 of those samples, collected between March 31 and October 25, 2021 (Fig. 1). Every pattern is uniquely named with a six-character alphanumeric code and has related geographic and temporal data, together with the latitude and longitude of the sampling website, in addition to placement and assortment dates. 17 of the 19 sampling websites have been a part of a hierarchical sampling design23 and have been due to this fact clustered comparatively carefully collectively. In ecological analyses, this nested design permits us to match samples that are shut to one another (some 102 m) to websites that are farther away (106 m) via a spread of scales (Fig. 1). After assortment, samples have been shipped to the Centre for Biodiversity Genomics, Guelph, Canada, the place they have been preserved in contemporary 96% ethanol and saved at −20°C till evaluation.
Geographical distribution and assortment dates of samples. Prime: Places of the 19 sampling websites throughout Sweden and northern Finland. SH, HT, and UA are a part of a hierarchical sampling design, every together with 5–7 entice places. The scale of every circle is proportional to the variety of samples collected at that website, which can be indicated by an integer under the entice title. Backside: Temporal distribution of the MassID45 samples. Assortment dates have been aggregated by week in order that samples collected throughout the identical week are displayed in the identical column, no matter what day of the week they have been collected. Hierarchically organized websites (SH, HT, and UA) are colored with shades of the identical foremost colors to emphasise their geographical proximity.
DNA barcoding and imaging workflows
Bulk pattern evaluation
We first analyzed samples via a bulk workflow24, the place all specimens collected in a pattern have been analyzed concurrently with out prior sorting. We weighed the arthropods from the majority pattern after filtering out the ethanol to acquire the moist biomass. We then carried out non-destructive lysis for DNA extraction and picked up three technical replicates from every pattern. After extraction, we amplified a brief (418 bp) fragment inside the usual barcoding area of COI, which we then sequenced on an Illumina NovaSeq 6000. After DNA extraction, we transferred every pattern to a translucent sorting tray (44 × 39 cm) with a shallow layer of ethanol and punctiliously unfold out the specimens to reduce overlap. We positioned the tray on an LED panel inside a modified mild dice, the place the entrance panel was eliminated and a gap was added within the ceiling to suit a digital camera (Fig. 2a). To additional enhance mild circumstances, we used two ring lights positioned on opposing sides of the sunshine dice. We captured a bulk picture from above with a Canon EOS R5 digital camera and an RF 24–240 mm F4-6.3 IS USM zoom lens mounted on a big copy stand. We used the next digital camera settings: focal size of 27 mm, aperture f/20, shutter velocity 1/6 seconds, and ISO 100. Every picture included a QR code distinctive to the pattern (Fig. 2b). For 4 samples weighing greater than roughly 10 g, we divided the pattern into two sorting trays, leading to two bulk photos for every of those samples, for a complete of 49 photos.

(a) Imaging setup used to seize bulk photos of the MassID45 dataset, together with the positioning of the digital camera, mild dice and ring mild sources. (b) A consultant picture captured utilizing the described imaging setup, with the perimeters trimmed.
We manually edited the full-resolution RAW photos (45 megapixels; 8192 × 5464) in Adobe Lightroom Traditional to enhance distinction and guarantee visibility of each mild and darkish insect physique components, utilizing the next settings: we elevated publicity by 1.3 stops, set whites and highlights to -100, and shadows to +50. To revive picture distinction and color we additionally adjusted readability and saturation to twenty and elevated the white stability from 4200K to 5050K. To cut back noise and purple fringing, we utilized luminance noise discount and defringe values of 20. Lastly, we elevated sharpening to 60 and saved the photographs in JPEG format.
Particular person specimen evaluation
After bulk imaging was accomplished, we positioned every specimen from the majority samples in a separate properly in a 96-well round-bottom microplate for particular person evaluation. Specimens smaller than 5 mm have been positioned instantly within the properly and imaged utilizing a Keyence VHX-7000 Digital Microscope system10. For bigger specimens (roughly > 5 mm), we eliminated a single leg for DNA extraction and pinned the primary physique of the arthropod for imaging utilizing an computerized Imaging Rig25. We amplified and sequenced full 658-bp DNA barcodes for every specimen utilizing single-molecule real-time (SMRT) sequencing26 on a PacBio Sequel platform. The success fee of amplification and sequencing of DNA barcodes from the person specimens was 97.5%, although solely 89.6% handed high quality and contamination checks. Together with components affecting metabarcoding success, resembling primer bias and amplification of non-target DNA (e.g., intestine contents), we due to this fact count on some discrepancies between the person and bulk-level DNA barcoding knowledge. We uploaded photos and DNA barcodes to BOLD and assigned taxonomic classifications primarily based on each picture and molecular data utilizing the BOLD ID engine. We retained all specimens for future morphological reference within the pure historical past assortment of the Centre for Biodiversity Genomics (BIOUG).
Pattern-specific taxonomies. Utilizing individual-level DNA barcodes, we constructed sample-specific taxonomies to information the annotations of the corresponding bulk photos. To do that, we used a base taxonomy masking all arthropods in Sweden, which we then subset for every pattern to incorporate solely taxa noticed in that pattern. We created the bottom taxonomy containing the ranks kingdom, phylum, subphylum, class, order, suborder, infraorder, superfamily, household, subfamily, genus, and species, by beginning with the taxonomy offered by BOLD14. We supplemented the BOLD taxonomy with the ranks suborder, infraorder and superfamily from Dyntaxa, the Swedish taxonomy database (downloaded from which covers all arthropods recorded in Sweden. We subset the Dyntaxa taxonomy to Hexapoda and Arachnida and mixed it with the BOLD taxonomy by matching genus names inside phyla and courses. Any taxonomic discordances between the taxonomies have been resolved by giving the BOLD taxonomy priority as follows. We used household names as they occurred within the BOLD taxonomy. For instances the place Dyntaxa used a distinct household title, we verified that the suborder, infraorder, and superfamily from Dyntaxa nonetheless utilized to the BOLD household title by comparability with the NCBI taxonomy database27. If NCBI listed one other suborder, infraorder, or superfamily for the BOLD household title, we modified the discordant rank in our taxonomy to match NCBI. Nonetheless, we didn’t add any data from NCBI to ranks that have been empty in our taxonomy. If there was no taxonomic data for the BOLD household, we stored the data from Dyntaxa for suborder, infraorder, and superfamily. Diplura, Collembola, and Protura occurred as courses in BOLD however as orders in Dyntaxa. We stored them as courses in our taxonomy and eliminated all sub- and infraorders, as the identical taxa appeared as orders in BOLD. The orders Phthiraptera and Psocoptera within the Dyntaxa taxonomy have been mixed into the order Psocodea in BOLD. We due to this fact used the latter in our taxonomy. We additionally added “microlepidoptera” as a casual taxonomic group between the ranks of infraorder and superfamily. This group included 14 superfamilies inside the order Lepidoptera (Adeloidea, Choreutoidea, Gelechioidea, Gracillarioidea, Micropterigoidea, Nepticuloidea, Pterophoroidea, Pyraloidea, Schreckensteinioidea, Tineoidea, Tischerioidea, Tortricoidea, Urodoidea, and Yponomeutoidea). Whereas microlepidoptera shouldn’t be a real taxonomic group, it’s a helpful classification when working with insect photos with low decision. Lastly, we generated sample-specific taxonomies by utilizing the taxonomic classifications obtained from DNA barcoding of particular person specimens in every of the 45 samples to subset the bottom taxonomy.
Bulk picture annotation
We annotated the majority photos in two steps: first, creating segmentation masks round every seen specimen, and second, assigning taxonomic labels to every arthropod masks. This workflow confined the necessity for taxonomic experience to the second step. Our strategy mixed three key methods to facilitate annotation: watershed segmentation for producing preliminary masks11, the Toronto Annotation Suite (TORAS)28 for AI-assisted masks refinement, and the DNA barcoding-derived sample-specific taxonomies to limit the set of urged taxonomic labels to taxa that have been confirmed to be current in every pattern. The entire annotation workflow is described intimately under.
Annotation step 1: Create segmentation masks
To facilitate fast annotation of a lot of arthropods, we used a watershed algorithm to generate preliminary segmentation masks11. Watershed segmentation treats the picture as a topographical floor, the place pixel intensities characterize heights, and finds boundaries between areas by simulating water filling up from native minima. Contiguous areas with pixel values under a threshold (200 in 8-bit grayscale) have been grouped right into a segmentation masks. Though computationally environment friendly, this easy algorithm usually merged shut groupings of arthropods into single masks and excluded light-coloured or translucent physique components, resembling wings, and slender constructions, like legs and antennae, from the masks. Subsequently, we improved the masks by manually enhancing them utilizing TORAS, a web-based annotation software harnessing human-in-the-loop AI fashions to hurry up and enhance annotations for laptop imaginative and prescient.
To allow quick annotation of tiny objects, we applied two customized options in TORAS. First, we modified the default zoom behaviour when creating a brand new segmentation masks: as a substitute of zooming out to point out the complete picture, we maintained the present zoom stage to forestall the annotator from shedding monitor of particular person bugs. Second, we built-in a scale bar to assist annotators higher gauge object sizes inside the photos. Additional, because the bulk samples contained between 36 and 3228 arthropods every, we used a customized script to separate photos into subimages for quicker masks rendering in TORAS. We first cut up photos into 4 × 4 equally-sized subimages. To keep away from splitting arthropods throughout subimage boundaries, we used the preliminary watershed masks to find them. We calculated every masks’s centroid and assigned it to the corresponding subimage. We then adjusted subimage sizes to incorporate the complete vary of every preliminary segmentation masks, with a 100-pixel buffer to permit for handbook masks changes. This methodology of splitting resulted in some overlap, inflicting some arthropods to seem in a number of subimages. We visually tagged these arthropods in all however one of many subimages they appeared in to forestall redundant manually created masks.
We uploaded the preliminary segmentation masks to TORAS and robotically improved them with the built-in segmentation refinement software, which adjusts every masks to incorporate solely pixels belonging to the item. In comparison with the uncooked watershed masks, the robotically refined masks offered a extra exact match across the arthropods. Three annotators with fundamental information of insect morphology then manually edited the refined masks to make sure that every arthropod had a separate segmentation masks, capturing all its pixels with out together with any background. The annotators might make the most of the complete vary of instruments out there in TORAS for enhancing the masks. For instance, Paint and Erase have been generally used to manually edit the masks, whereas the Field Device, which estimates a segmentation masks primarily based on a bounding field offered by the annotator, was used to generate masks for arthropods missed by the watershed algorithm. Lastly, the annotators assigned every segmentation masks to certainly one of 4 coarse courses: arthropod (b for “bug”), particles (d), edge (e, together with tray edges and QR codes) or unidentifiable (u). For detailed annotator directions, see Supplementary Data S1.
Annotation step 2: Assign taxonomic labels
Within the second step of annotations, one “expert annotator” with expertise in arthropod identification used TORAS to assign taxonomic labels to every segmentation masks beforehand categorised as containing an arthropod (class b). To attenuate misspellings or taxonomic disagreements between ranks, the skilled annotator chosen labels from the sample-specific taxonomy constructed from the individual-level DNA-based taxonomic classifications (see Particular person specimen evaluation). Nonetheless, since there was some, albeit small, proportion of specimens for which the person DNA barcoding failed, the skilled annotator was capable of override the default set of decisions by creating and assigning labels that didn’t happen within the sample-specific taxonomy. The skilled annotator was requested to assign the bottom taxonomic group doable for every segmentation masks containing an arthropod.
To permit the skilled annotator to convey as detailed taxonomic data as doable, we distinguished between labels of excessive and low confidence utilizing two totally different strategies. First, by assigning a number of taxonomic labels at totally different ranks to a single segmentation masks: the label which belonged to the best taxonomic rank was thought of excessive confidence, whereas all lower-ranking labels have been handled as low confidence. That’s, if the skilled annotator assigned order:Lepidoptera and household:Tortricidae, Lepidoptera was thought of excessive confidence and Tortricidae low confidence. Second, by assigning a number of labels on the similar rank: their most up-to-date widespread ancestor was thought of a high-confidence label, whereas all different labels have been handled as low-confidence. For instance, if the skilled annotator assigned each household:Tortricidae and household:Geometridae, these household labels could be thought of low confidence, whereas their most up-to-date widespread ancestor, order:Lepidoptera, could be interpreted as excessive confidence, no matter whether or not the skilled annotator explicitly assigned it. These strategies allowed the skilled annotator to assign uncertainty with out requiring a separate step for assessing label confidence. In addition they prevented the issue of the skilled quantifying their confidence.
Along with assigning taxonomic labels, we requested the skilled annotator to carry out a high quality test of the segmentation masks from the primary annotation step. This was performed with a customized characteristic in TORAS, which allowed annotators to view the segmentation masks within the second annotation step. In the course of the high quality test, the skilled annotator primarily adjusted the masks if visible traits vital for the classification of the arthropod, resembling wings or antennae, weren’t included within the masks, or if a number of arthropods have been grouped in a single masks. The skilled annotator might additionally change the preliminary task to one of many 4 coarse courses, if, for instance, particles or an unidentifiable object had mistakenly been labelled as an arthropod. For detailed annotation directions, see Supplementary Data S1.4.
Annotation completeness and reliability
We have been capable of annotate nearly all of specimens within the bulk photos at rank suborder or above with excessive confidence (Desk 1). Together with low-confidence annotations elevated the variety of specimens annotated at decrease ranks, with nearly half of the specimens annotated on the superfamily stage, and greater than a 3rd on the household stage (Desk 1). At decrease ranks, resembling genus or species, taxonomic classifications from bulk photos alone turn into virtually not possible for many taxa. Morphological traits could also be obscured by the random orientation of bugs within the photos, the place, for instance, wings could be hidden underneath the primary physique, or by the low decision at which particular person specimens are depicted inside the bulk photos. Moreover, many diagnostic options are solely out there via microscopy or dissection of constructions such because the genitalia, making examination of the bodily specimen successfully vital. The energy of this dataset thus lies primarily in its utility for coaching fashions that function on the order or household stage, the place label protection and confidence are highest. Even with out species-level decision, these ranks are sometimes enough to disclose broad patterns in biodiversity and neighborhood composition29,30. Within the bulk photos, 45 specimens have been annotated with the brief label b with none further taxonomic label, leading to 17892 of 17937 bug segmentation masks being taxonomically labelled.
Inside this work, we didn’t carry out any unbiased validation of annotations past the sanity test of segmentation masks made in the course of the second annotation step, primarily resulting from restricted annotator availability throughout dataset improvement. This makes it tough to instantly consider the consistency and uncertainty related to each the segmentation masks and the taxonomic annotations. Nonetheless, the mix of bulk-level and individual-specimen knowledge in our dataset gives an inner foundation for evaluating annotated labels in opposition to anticipated patterns, which may give a sign of annotation reliability.
To judge how precisely the majority picture annotations replicate the true variety of arthropods within the samples, we in contrast the variety of segmentation masks annotated as arthropods with the precise variety of specimens remoted from every pattern (Fig. 3a). We discovered that in samples containing greater than roughly 250 arthropods, the variety of arthropods primarily based on the majority picture annotations was considerably decrease than the true depend. A few of these discrepancies occurred in samples with a excessive abundance of springtails (Collembola), which are sometimes small, pale, and tough to separate from particles within the bulk samples. Proscribing the comparability to particular person specimens categorised as Insecta or Arachnida (each of that are usually bigger and darker than springtails) decreased the distinction between annotated counts and true specimen counts. Total, absolutely the discrepancy in counts tended to extend with bigger samples, suggesting two potentialities. First, it’s inherently difficult for human annotators to detect all bugs in photos the place the whole variety of people could be very excessive. These samples additionally usually contained substantial particles, which may obscure smaller bugs (Fig. 3b). Moreover, as a result of every insect occupies solely a small proportion of the picture, particularly tiny bugs could seem visually vague or blurry, making them tough to annotate accurately. This was significantly distinguished when insect sizes diverse drastically inside a picture, because the restricted focal airplane prevented all bugs from being equally in focus. Second, annotator fatigue could set in for these giant samples, resulting in fewer corrections for arthropods lacking a segmentation masks as soon as the whole depend is already excessive. Nonetheless, this rationalization stays speculative as we didn’t file the annotation session metadata vital to check the impact of fatigue on annotation charges instantly. Recording such metadata in future dataset efforts would allow extra rigorous high quality evaluation on this respect.
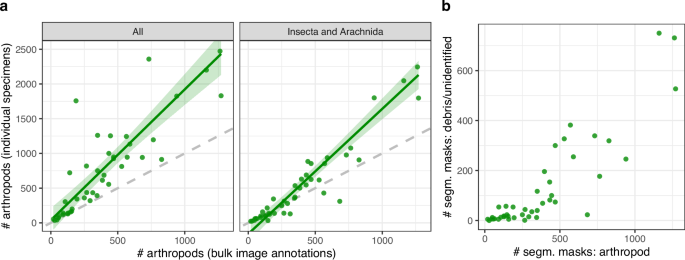
(a) For every pattern, a comparability was made between the variety of arthropods annotated within the bulk photos and the variety of particular person specimens remoted from the corresponding samples, right here proven for all taxa (left) and restricted to Insecta and Arachnida (proper). The inexperienced line represents a linear regression match with a 95% confidence interval between the 2 portions, and the dashed gray line signifies a 1:1 relationship. (b) For every pattern, a comparability was made between the variety of segmentation masks tagged as arthropods and the quantity tagged as particles or unidentifiable throughout all bulk photos.
Utilizing high-confidence taxonomic annotations on the order rank, we in contrast the variety of bulk picture annotations with the variety of specimens remoted from every pattern (Fig. 4). We discovered that the majority picture annotations usually recovered fewer specimens per order than sorted people, however that the proportion of specimens lacking from the annotations was largely constant throughout pattern sizes, suggesting that the relative proportion of taxa is reliably captured in our annotations. According to the whole depend comparability, Collembola – represented right here by the biggest order, Entomobryomorpha – have been the exception, with many specimens lacking from the majority picture annotations. This sample could once more replicate that lots of the smallest specimens have been missed throughout annotation resulting from their poor visibility. Nonetheless, the dimensions distribution of our bulk picture annotations general agrees with the anticipated distribution from Malaise entice catches. Right here, the “tiny” class corresponds to specimens with an annotated space of <1 mm2. Translating the annotation space to standard physique dimension shouldn’t be simple, because the measured space could also be inflated by constructions resembling wings, and physique size can’t be instantly inferred from space. Nonetheless, 60% of bugs in our dataset had an annotated space under 3 mm2, which has similarities to dimension distributions reported from different Malaise entice collections31.
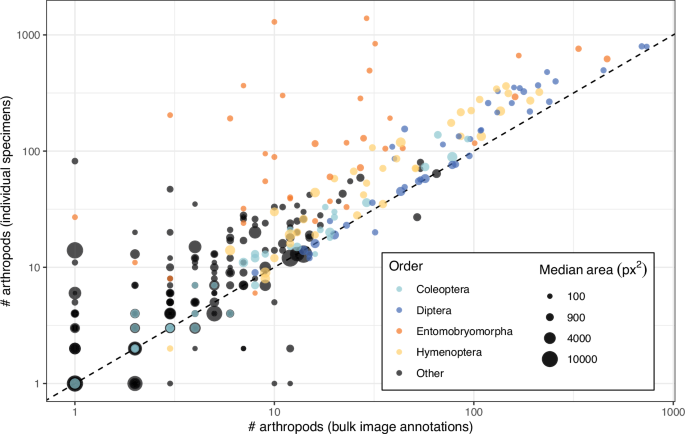
For every pattern and order, we in contrast the variety of arthropods annotated within the bulk photos and the variety of particular person specimens remoted from the corresponding samples. The dashed line exhibits a 1:1 relationship between the 2 counts. The areas of the circles characterize the median segmentation masks space for every pattern and order. The 4 most considerable orders in our datasets are proven in colors, whereas all different orders are grouped collectively and proven in black.
Machine studying dataset
MassID45 serves as a benchmark dataset as an illustration segmentation of tiny, densely packed objects. Occasion segmentation differs from different duties like object detection, which makes use of bounding containers, and semantic segmentation, which labels pixels however doesn’t distinguish between particular person objects. In distinction, occasion segmentation assigns a novel pixel-wise masks to every particular person object (Fig. 5). Utilizing the totally annotated set of 49 bulk photos, we restricted the duty to occasion segmentation of arthropod specimens (class b), i.e., excluding objects tagged as particles or unknown. This allowed us to strategy this downside as a single-class, small-instance segmentation job.

Variations between annotations for object detection (left), occasion segmentation (center), and semantic segmentation (proper).
Annotations are exported from TORAS in the identical format because the Microsoft Frequent Objects in Context (MS-COCO) dataset32, a benchmark dataset for pretraining object detection and occasion segmentation fashions. We primarily based the analysis scheme for the occasion segmentation fashions on MS-COCO conventions, together with metrics that depend on the areas of the occasion masks to be detected. Nonetheless, the MS-COCO analysis scheme was designed for objects which might be considerably bigger than these within the MassID45 dataset, with 76.5% of the arthropod masks being categorized as “small”. Subsequently, to extend the granularity in our efficiency evaluations, we as a substitute used the world thresholds from iSAID33, a dataset of distant sensing photos supposed for small object detection and occasion segmentation. Thus, we outlined object sizes as follows: “tiny” for areas < 144 pixels; “small” for areas ≥144 however < 1024 pixels; and “medium/large” for areas ≥1024 pixels. In whole, the 49 totally annotated bulk photos include segmentation masks for 17937 arthropods. Masks areas vary between 15.1 and 83182.4 pixels, with a imply and median of 1152.2 and 343.4 pixels, respectively (Fig. 6).
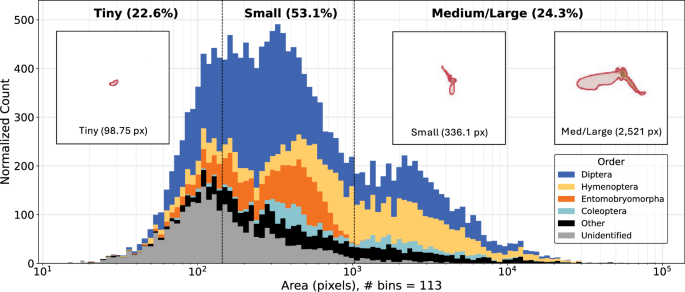
Distribution of insect masks areas for “tiny” (<144 pixels), “small” (≥144 however <1024 pixels), and (“medium/large” (≥1024 pixels) bugs. Counts are adjusted such that the world of a bar is proportional to the depend in that bin. Colors inside the stacked bars characterize the 4 most considerable orders, all different named orders grouped as “Other”, and orders with out order-level annotation as “Unidentified”. The three overlaid photos present the median masks for tiny, small, and medium/giant bugs, all on the similar magnification.
Preprocessing bulk photos
Earlier than coaching deep neural networks to carry out occasion segmentation, we preprocessed the picture and segmentation masks knowledge. We first merged the annotations from the subimages again collectively to match the unique bulk photos (Bulk picture annotation). This allowed us to interrupt up the photographs into tiles as wanted for mannequin coaching, detailed under. We used the Shapely library34 in Python to merge segmentation masks with a number of polygons and to appropriate invalid (i.e., self-intersecting) polygon masks. The insect masks have been processed as concave hulls, filling in holes (e.g., areas between legs) within the edited segmentation masks to create single polygons. For 63 of the 17937 insect masks (0.351%), these preprocessing steps resulted in a number of unconnected polygons that would not be merged by way of a unary union. In such instances, we took the polygon with the biggest space as the ultimate masks. This ensured that deep studying fashions solely wanted to foretell one polygon per annotation, simplifying the segmentation job. After cleansing the segmentation masks, we manually cropped the majority photos to solely include the areas by which bugs have been current. This resulted in finalized cropped bulk photos of various dimensions.