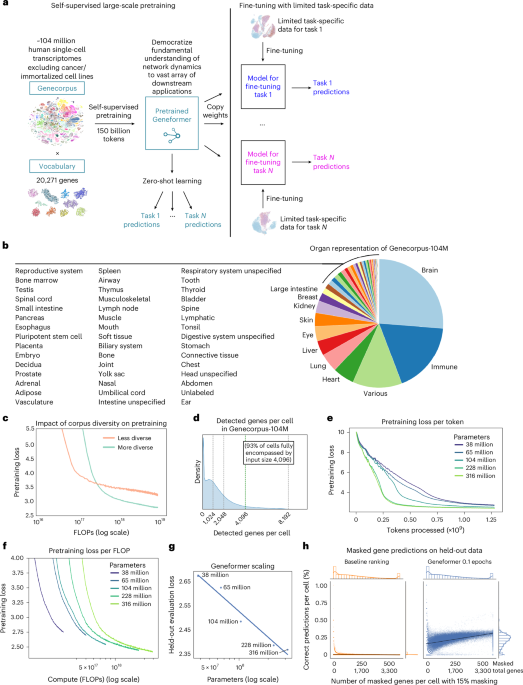
Meeting and rank worth encoding of transcriptomes in Genecorpus-104M
Meeting and uniform processing of single-cell transcriptomes
We assembled a large-scale pretraining corpus, Genecorpus-104M, comprising ~104 million human single-cell transcriptomes (post-filtering as described under) from a broad vary of tissues from 2,903 publicly accessible datasets (Fig. 1b and Supplementary Desk 1). Importantly, DOIs have been cross-referenced between all research to make sure that datasets have been distinctive to keep away from inclusion of duplicated cells throughout the corpus. Of notice, there are substantial duplications of datasets throughout public databases so the overall variety of distinctive cells can be extremely overestimated if this process weren’t carried out (Prolonged Knowledge Fig. 1a).
Publicly accessible datasets containing uncooked counts have been collected from Nationwide Middle for Biotechnology Info (NCBI) Gene Expression Omnibus (GEO), NCBI Sequence Learn Archive (SRA), CELLxGENE, Human Cell Atlas, European Molecular Biology Laboratory-European Bioinformatics Institute (EMBL-EBI) Single Cell Expression Atlas, Broad Institute Single Cell Portal, Brotman Baty Institute (BBI)-Allen Single Cell Atlases, Tumor Immune Single-cell Hub (TISCH) (excluding malignant cells), Panglao Database, 10x Genomics, College of California, Santa Cruz Cell Browser, European Genome-phenome Archive, Synapse, Riken, Zenodo, Nationwide Institutes of Well being (NIH) Figshare Archive, NCBI dbGap, Refine.bio, China Nationwide GeneBank Sequence Archive, Mendeley Knowledge, and particular person communication with authors of the unique research (Supplementary Desk 1). Extra sources for amassing details about appropriate research included Entrez Direct instruments and the dataset abstract from Svensson et al., Database 202022. Instruments utilized in conversion of information to uniform information included loompy, scanpy, anndata, scipy, numpy, pandas, Cell Ranger and LoomExperiment. Gene annotation knowledge have been retrieved from Ensembl, NCBI and HGNC (1 November 2023) databases and moreover queried by means of MyGene23. Uncooked and unfiltered knowledge information have been processed to take away empty droplets and particles utilizing STAR model 2.7.8a with the Cell Ranger2.2 (run mode –soloCellFiltered). Datasets have been moreover filtered to retain cells that contained a minimal of seven detected Ensembl-annotated protein-coding genes provided that the 15% masking used for the pretraining studying goal wouldn’t reliably masks a gene in cells with fewer detected genes. Research have been annotated as a number of of the 55 consolidated organs as listed in Fig. 1b.
Rank worth encoding of single-cell transcriptomes
Every transcriptome was introduced to the mannequin as a rank worth encoding as beforehand described6. The rank worth encodings are a non-parametric illustration of the transcriptome that takes benefit of the numerous observations of the gene’s expression throughout all the Genecorpus to prioritize genes that distinguish cell state. Particularly, this methodology will deprioritize ubiquitously extremely expressed housekeeping genes by normalizing them to a decrease rank. Conversely, genes resembling transcription elements which may be lowly expressed when they’re expressed however extremely distinguish cell state will transfer to the next rank throughout the encoding. Moreover, this rank-based method could also be extra strong in opposition to technical artifacts that will systematically bias absolutely the transcript counts worth whereas the general relative rating of genes inside every cell stays extra secure.
The rank worth encodings have been constructed as beforehand described6. The scaling issue for every gene was derived from the non-zero median worth of expression of every detected gene throughout all cells within the pretraining corpus passing high quality filtering that have been sequenced on droplet-based platforms, excluding cells with excessive mutational burdens resembling malignant cells and immortalized cell traces. After scaling the expression of every gene, the genes have been ordered by the rank of their scaled expression in that particular cell. The rank worth encoding for every single-cell transcriptome was then tokenized on the idea of a vocabulary of 20,271 protein-coding genes detected throughout the pretraining corpus. The vocabulary additionally included 4 particular tokens: a padding, masking, CLS (classification) and EOS (finish of state) token, for a complete vocabulary dimension of 20,275. A CLS and EOS token have been added to the start and finish of every rank worth encoding, respectively. The CLS token is standardly abbreviating the time period ‘classification’ as it’s used for fine-tuning the mannequin for input-level classification (on this case cell-level classification). Normally, nonetheless, the CLS token is used right here as a worldwide contextual cell illustration because it happens firstly each cell with its that means tuned by way of the bidirectional consideration to the actual cell’s context utilized throughout coaching. The tokenized dataset was saved throughout the Hugging Face Datasets construction, which relies on the Apache Arrow format that enables processing of enormous datasets with zero-copy reads with out reminiscence constraints.
Of notice, this technique can also be space-efficient because the genes are saved as ranked tokens versus the precise transcript values, and we solely retailer genes detected inside every cell relatively than the total sparse dataset that features all the undetected genes. This additionally prevents losing computation on zeros, because the mannequin learns from the absence of genes from a rank worth encoding with out having to explicitly instruct the mannequin that they’ve zero expression. That is analogous, for instance, to how pure language fashions be taught {that a} assertion could have ‘positive’ that means based mostly on the absence of ‘negative’ phrases, while not having to current the rest of the absent phrases from the pure language dictionary on the finish of each sentence to explicitly instruct the mannequin they don’t seem to be current.
Geneformer structure, pretraining and quantization
Geneformer structure
Geneformer consists of dense transformer encoder items, every composed of a self-attention layer and a feed-forward neural community. The unique Geneformer mannequin6 pretrained in June 2021 was an ~10 million parameter mannequin with enter dimension of two,048. On this work, the enter dimension was expanded to 4,096 genes per cell, which absolutely represents 93% of the cells in Genecorpus-104M when contemplating genes throughout the mannequin vocabulary of 20,271 protein-coding genes. Geneformer fashions of accelerating parameter rely have been pretrained on this work to judge the scaling legal guidelines of transcriptional masked studying. The depth of the Geneformer fashions skilled with 38 million, 65 million, 104 million, 228 million and 316 million parameters was 8, 10, 12, 16 and 18 layers, respectively. All fashions had a width-to-depth side ratio of 64, which was optimized by evaluating fashions skilled with the identical variety of parameters however various width-to-depth side ratios. All fashions had an embedding-dimension-to-attention-head ratio of 64 and feed-forward-size-to-embedding-dimension ratio of 4. For instance, the 316 million parameter mannequin was 18 layers with 1,152 embedding dimensions, 18 consideration heads and a feed-forward dimension of 4,608. Additional parameters are as follows: nonlinear activation operate, rectified linear unit; dropout chance for all absolutely linked layers, 0.02; dropout ratio for consideration chances, 0.02; customary deviation of the initializer for weight matrices, 0.02; epsilon for layer normalization layers, 1 × 10−12. Modeling was applied in pytorch and utilizing the Hugging Face Transformers library for mannequin configuration, knowledge loading and coaching.
Geneformer pretraining and efficiency optimization
Geneformer was pretrained with ~104 million (103,877,737) single-cell transcriptomes from Genecorpus-104M excluding cells with excessive mutational burdens resembling malignant cells and immortalized cell traces. Pretraining was achieved utilizing a masked studying goal, which has been proven in different informational fields3,4 to enhance generalizability of the foundational data discovered throughout pretraining for a variety of downstream fine-tuning aims. Throughout pretraining, 15% of the genes inside every transcriptome have been masked, and the mannequin was skilled to foretell which gene ought to be inside every masked place in that particular cell state utilizing the context of the remaining unmasked genes. A serious power of this method is that it’s fully self-supervised and might be achieved on fully unlabeled knowledge, which permits the inclusion of enormous quantities of coaching knowledge with out being restricted to samples with accompanying labels. Pretraining hyperparameters have been optimized to the next closing values: max studying price, 2 × 10−4; studying scheduler, cosine with warm-up; optimizer, Adam with weight decay repair; warm-up ratio, 0.007; weight decay, 0.044; efficient batch dimension (batch dimension × GPUs), 432 for all fashions besides 228 million parameter mannequin, which had an efficient batch dimension of 216. For the scaling evaluation, all fashions have been pretrained for 0.1 epochs. For the full-model pretraining, GF-104M was pretrained for 3 epochs, and GF-316M was pretrained for 1 epoch. Weights & Biases was used for experimentation monitoring.
Because the enter dimension of 4,096 is significantly massive for a full dense self-attention mannequin (for instance, BERT3,4 enter dimension of 512) and transformers have a quadratic reminiscence and time complexity O(L2) with respect to enter dimension, we applied distributed GPU coaching algorithms24,25 to permit environment friendly pretraining on the large-scale dataset utilizing Deepspeed. This method partitions parameters, gradients and optimizer states throughout accessible GPUs, offloads processing/reminiscence as potential to a central processing unit to permit extra to suit on a GPU, and reduces reminiscence fragmentation by guaranteeing that long- and short-term reminiscence allocations don’t combine. Pretraining was carried out in full precision (FP32) and distributed throughout 8–24 Nvidia H100 80GB GPUs.
Geneformer quantization and parameter environment friendly fine-tuning
Quantization of Geneformer fashions for fine-tuning was carried out utilizing QLoRA16, a method that allows parameter- and memory-efficient fine-tuning of enormous language fashions by combining low-bit quantization with low-rank adaptation. On this set-up, gradients are propagated by means of the frozen, quantized base mannequin with out updating its weights, and are used to solely replace the light-weight, task-specific low-rank adapters, enabling memory- and compute-efficient fine-tuning. For each gene-level and cell-level fine-tuning duties, Geneformer was quantized to 4-bit precision utilizing the NormalFloat4 (NF4) format, enabling fine-tuning on useful resource constrained {hardware} with out sacrificing mannequin efficiency.
QLoRA achieves parameter-efficient fine-tuning by integrating two complementary methods: quantization of the bottom mannequin and using low-rank adapters to carry out task-specific updates. As a substitute of updating all weights within the mannequin, QLoRA freezes the vast majority of the parameters and introduces trainable low-rank matrices into choose layers of the mannequin structure, sometimes throughout the consideration mechanisms (for example, the important thing, question, worth projection matrices) the place the majority of the computations happen. Low-rank adaptation (LoRA)26 replaces a full-rank weight replace with a low-rank decomposition. For a weight matrix W with form N × N, which might usually require N2 parameter updates throughout fine-tuning, LoRA approximates the replace W with two matrices of smaller rank: A of form N × r and B of form r × N, the place r is way smaller than N. This reduces the variety of trainable parameters from N2 to 2Nr, offering a considerable acquire in effectivity, significantly when N is massive, and r ≪ N. The low-rank product ΔW = AB is then added to the unique, frozen weight matrix W to protect the pretrained data whereas enabling environment friendly task-specific adaptation.
As the bottom mannequin weights are frozen, LoRA modules might be skilled independently for various duties and saved individually, which facilitates modular deployment. LoRA’s use of low-rank updates is efficient as a result of pretrained massive language fashions have low intrinsic dimensionality, that means there exists a low-dimensional subspace that may be practically as efficient for fine-tuning as optimizing over the total parameter area27. Curiously, bigger massive language fashions are inclined to have decrease intrinsic dimensionality as a result of they’ve already been extensively pretrained on a broad and various set of information, permitting them to generalize extra successfully with fewer task-specific parameter updates.
To additional enhance reminiscence effectivity, QLoRA makes use of quantization strategies to compress the bottom mannequin earlier than inserting the low-rank adapters. Quantization is the method of decreasing the numerical precision of mannequin weights, sometimes from 16- or 32-bit floating-point values to lower-bit representations resembling 8-bit or 4-bit, to scale back reminiscence utilization and speed up computation, whereas guaranteeing minimal lack of data from the unique weights. QLoRA makes use of blockwise quantization using the NF4 format (which is extra correct than customary 4 bit floats or integers) that preserves the distributional properties of full-precision weights whereas enabling substantial compression. For fine-tuning, Geneformer was quantized from 32-bit (FP32) to 4-bit (NF4), additionally utilizing double quantization, which makes use of a second nested quantization to save lots of an extra 0.4 bits per parameter.
For inference, the full-precision (FP32) fashions have been quantized to 8-bit precision. The quantization configuration loaded the mannequin in 8-bit precision with out utilizing low-rank matrices A and B for fine-tuning. The total-precision weights have been instantly transformed to a decrease precision with out extra low-rank factorization, such that the unique structure and weights remained intact through the quantization course of.
To supply a sensible estimate of inference time and price for in silico perturbation, the GF-V2-316M mannequin was examined both as full precision or quantized within the job of genome-wide in silico deletion for cells with 4,096 genes detected every, which for 30,000 cells ends in 122,910,000 complete cells processed throughout inference together with each unperturbed and simulated perturbed cells. With the identical batch dimension of 64, genome-wide in silico perturbation for 100 circumstances (for example, cell sorts or developmental levels) with 300 cells every on 12 Nvidia 80G A100 GPUs would take 32.8 days for the full-precision mannequin however solely 5.9 days for the quantized mannequin. With the present worth from a low-cost business supplier of US$2.56 per Nvidia 80G A100 GPU hour, the experiment would value US$24,188 for the full-precision mannequin however solely US$4,361 for the quantized mannequin. On condition that the quantization additionally saves reminiscence, rising the batch dimension to make the most of the freed reminiscence would end in even higher time and price advantages. Of notice, compute time and price depends upon a number of elements together with dataset dimension, variety of genes detected per cell, variety of perturbations examined, which mannequin is used, which GPU distribution strategies are used, the reminiscence and velocity of the GPU sources accessible, the price of the particular GPU sources at that given time limit, and so forth.
Masked studying analysis
Geneformer fashions with rising parameters have been evaluated on 100,000 held-out cells randomly subsampled from Genecorpus-104M to retain the total variety of that dataset. For the comparability with baseline masked gene predictions, a Geneformer mannequin of 104 million parameters was utilized, pretraining for 0.1 epochs and evaluating on held-out cells randomly subsampled from Genecorpus-104M. Efficiency on the masked studying goal will increase with pretraining, as proven within the reducing loss over coaching time in Fig. 1f; 0.1 epochs was chosen for this analysis because it represents the compute optimum frontier. The share of appropriate predictions per cell was quantified whereas masking 15% of the genes. The variable variety of masked positions per cell displays the variable variety of genes detected per cell, with the proportion masked all the time remaining stably at 15%.
The baseline masked gene predictions have been derived by figuring out the gene mostly ranked at every place throughout the rank worth encoding in the identical 10% of the pretraining corpus noticed by the Geneformer mannequin throughout pretraining. The baseline method predicts the most definitely gene for every rank throughout the 15% masked genes. This technique compares the baseline method for the equivalently difficult job that the Geneformer mannequin is required to carry out, specifically, figuring out which gene throughout the 20,271 gene vocabulary ought to be predicted for every rank throughout the 15% masked genes. This method additionally ensures that the distribution for the baseline predictions shouldn’t be drawn from the analysis knowledge itself, as some earlier research have completed. Moreover, the analysis knowledge are of equal variety because the pretraining corpus, guaranteeing an equitable comparability provided that basis fashions are tasked with predicting genes inside masked positions within the full variety of tissues, developmental levels and illness states within the pretraining corpus. This ensures that the baseline predictions aren’t based mostly on a distribution drawn from a slender analysis dataset itself, as on the excessive a baseline evaluated on a single cell can be 100% correct if the analysis knowledge have been used to generate the baseline predictions, which is problematic.
Contextual gene embedding extraction and robustness to batch-dependent technical artifacts
For every single-cell transcriptome introduced to Geneformer, the mannequin embeds every gene into an N-dimensional area that encodes the traits of the gene particular to the context of that cell. Contextual Geneformer gene embeddings are extracted because the hidden state weights for the N embedding dimensions for every gene throughout the given single-cell transcriptome evaluated by ahead go by means of the Geneformer mannequin. Gene embeddings analyzed on this research have been extracted from the second-to-last layer of the fashions as the ultimate layer is thought to embody options extra instantly associated to the educational goal prediction whereas the second-to-last layer is a extra generalizable illustration. Contextual zero-shot gene embeddings for the quantized versus full-precision fashions have been evaluated for cosine similarity between the identical versus completely different genes throughout the similar or completely different cell subclasses as annotated inside a random subsample of 248,569 regular grownup cells inside uniformly annotated CELLxGENE knowledge.
To quantify the affect of widespread batch-dependent technical artifacts on Geneformer gene embeddings, we in contrast (1) the cosine similarity of embeddings from two randomly chosen genes from the identical cell (which we count on to have low cosine similarity), (2) the cosine similarity of embeddings from the identical gene from two completely different cells of the identical cell sort from the identical batch (which we count on to have excessive cosine similarity), and (3) the cosine similarity of embeddings from the identical gene from two completely different cells of the identical cell sort from completely different batches (which we count on to have excessive cosine similarity if the gene embeddings are strong to batch-dependent technical artifacts). We carried out the above process to quantify (1) platform-related results utilizing 500 cells iPSCs assayed in parallel on the Drop-seq (single cell) or DroNc-seq (single nucleus) platform28, (2) preservation-related results utilizing 330 contemporary versus frozen pure killer (NK) cells from the identical donor ( and and (3) preprocessing-related results utilizing 4,000 peripheral blood mononuclear cells29 aligned to genome meeting GRCh37 versus GRCh38 or processed utilizing Cell Ranger variations 2.2.0, 3.1.0 or 7.1.030. Evaluation used cell-type annotations supplied by authentic authors for all datasets. Quantification of cosine similarity was carried out with n = 100 per situation examined (100 cells per batch per cell sort). Of notice, though we discovered that Geneformer was strong to the widespread batch-dependent technical artifacts that we examined, Geneformer was not designed particularly for scRNA-seq batch integration. As such, customers could elect to preprocess their restricted task-specific knowledge with different batch integration strategies earlier than utilizing that knowledge for fine-tuning Geneformer towards their downstream job in the event that they discover their dataset to be persistently affected by such batch-dependent artifacts.
As well as, separability of the gene embeddings was quantified by the geometric separability index, which was calculated by extracting gene embeddings from a randomly subsampled set of 1,000 cells from the identical cell sort from the 4,000 PBMCs from evaluation 3 above and for genes detected at the least 20 instances in these cells, measuring whether or not their nearest neighbor within the embedding area was an embedding of the identical gene. The proportion of genes the place the closest neighbor was an embedding of the identical gene was 1.0.
Contextual cell embedding extraction and robustness to batch-dependent technical artifacts
Geneformer cell embeddings, which encode traits of the state of that single cell, are generated by extracting the hidden state weights for the N embedding dimensions for the CLS token firstly of the given cell’s rank worth encoding evaluated by ahead go by means of the Geneformer mannequin. For all functions on this research, we used the second-to-last layer embeddings as mentioned above. Contextual zero-shot cell embeddings for the quantized versus full-precision fashions have been evaluated for cosine similarity between the identical versus completely different cell subclasses as annotated inside a random subsample of 248,569 regular grownup cells inside uniformly annotated CELLxGENE knowledge.
To quantify the affect of widespread batch-dependent technical artifacts on Geneformer cell embeddings, we in contrast (1) the cosine similarity of embeddings from two randomly chosen cells of various cell sorts (which we count on to have low cosine similarity), (2) the cosine similarity of embeddings from two randomly chosen cells of the identical cell sort from the identical batch (which we count on to have excessive cosine similarity), and (3) the cosine similarity of embeddings from two randomly chosen cells of the identical cell sort from completely different batches (which we count on to have excessive cosine similarity if the cell embeddings are strong to batch-dependent technical artifacts). We carried out the above process to quantify (1) platform-related results utilizing 500 cells iPSCs assayed in parallel on the Drop-seq (single -cell) or DroNc-seq (single nucleus) platform28, (2) preservation-related results utilizing 330 contemporary versus frozen NK cells from the identical donor ( and and (3) preprocessing-related results utilizing 4,000 peripheral blood mononuclear cells29 aligned to genome meeting GRCh37 versus GRCh38 or processed utilizing Cell Ranger variations 2.2.0, 3.1.0 or 7.1.0 (CR2, 3 or 7, respectively)30. We repeated the process of quantifying the similarity of cell pairs throughout the comparisons 1, 2 and three above for a complete n = 1,120,000, 79,200 and 79,200, respectively, for genome reference; n = 840,000, 79,200 and 39,600, respectively, for CR2 versus CR3; n = 1,400,000, 158,400 and 39,600, respectively, for CR2/3 versus CR7; n = 40,000, 20,000 and 19,800, respectively, for the preservation methodology. Evaluation used cell-type annotations supplied by authentic authors for all datasets. As well as, as a result of the preprocessing-related results allowed comparability for a similar actual cell below completely different circumstances, we quantified the affect of the genome reference and Cell Ranger model on the embedding of the very same cell by cosine similarity, which once more we count on to have excessive cosine similarity if the cell embeddings are strong to batch-dependent technical artifacts. Of notice, the evaluation of batch-dependent technical artifacts targeted on the identical and completely different cell kinds of iPSCs and cardiomyocyte subtype 1 as cardiomyocyte subtype 2 annotated by the unique authors is each comparable and completely different from cardiomyocyte subtype 1 so couldn’t be clearly categorized as comparable or completely different for the needs of this batch artifact analysis.
We additionally examined the affect of fine-tuning on the batch-dependent technical artifacts associated to the decrease variety of genes detected by single-nuclear versus single-cell sequencing platforms in contrast with evaluation 1 above. GF-104M was fine-tuned to differentiate cell sorts (as annotated by authentic authors) utilizing solely cells sequenced on the single-cell (Drop-seq) sequencing platform. The fine-tuned embedding area was then evaluated by the identical process as above. The ultimate embedding layer is thought to embody options extra instantly associated to the educational goal prediction whereas the second-to-last layer is a extra generalizable illustration. As such, the uniform manifold approximation and projections (UMAPs) mirror the second-to-last layer for the zero-shot mannequin and the final layer for the fine-tuned mannequin because the fine-tuning studying goal was certainly designed to handle the separability of cell sorts versus the final self-supervised pretraining goal. The unique knowledge UMAP was generated in response to the procedures outlined within the scanpy clustering tutorial, with or with out normalization by ComBat17 or Concord18 strategies as indicated.
The zero-shot Geneformer embedding area improved the separation by cell sort in contrast with the unique knowledge, even after batch correction by ComBat or Concord, and fine-tuning to differentiate cell sort utilizing knowledge from just one platform (single-cell platform) additional oriented the embedding area to separate cells by cell sort greater than by sequencing platform. Nonetheless, as mentioned above, Geneformer was not designed particularly for scRNA-seq batch integration. As such, customers could elect to preprocess their restricted task-specific knowledge with different batch integration strategies earlier than utilizing that knowledge for fine-tuning Geneformer towards their downstream job in the event that they discover their dataset to be persistently affected by such batch-dependent artifacts.
Zero-shot, few-shot and fine-tuning analysis in gene and cell-level duties
Gene-level job analysis
We examined the full-precision, quantized and different fashions throughout a various panel of biologically significant duties together with distinguishing (1) illness genes (dosage-sensitive versus -insensitive transcription elements31,32,33), (2) downstream targets of a transcription issue with out perturbation knowledge (NOTCH1 targets versus non-targets1,2), (3) chromatin dynamics from transcriptomic knowledge alone (bivalent versus Lys4-only methylated promoters34) and 4) transcription issue regulatory vary with no data of genomic distance (transcription elements that act in lengthy versus quick vary with their targets35).
The gene set labels and analysis knowledge have been as beforehand described6. Activity 1 is related to deciphering copy quantity variants in genetic prognosis to find out that genes are delicate to adjustments of their dosage. On this evaluation, the illness gene job used cells from Genecorpus-30M to make sure that the cells have been noticed by each the unique and present fashions throughout pretraining. Of notice, it was beforehand proven that downstream job efficiency was not impacted by whether or not the info have been included or excluded from the pretraining corpus provided that the downstream duties are distinct from the pretraining job of the masked studying goal6.
Activity 2 is related to predicting the direct downstream targets of transcription elements to map gene networks regulating cell states even within the absence of task-specific perturbation knowledge. Activity 3 is related to predicting chromatin regulation from transcriptomic knowledge alone and distinguishing bivalent marker promoters, that are identified to mark key developmental elements in embryonic stem cells and keep their promoters poised for activation. Efficiency was evaluated for the 56 extremely conserved loci during which bivalent marks have been initially described34 in Fig. 2b and for genome-wide bivalent chromatin marks36 in Fig. 2c.
Activity 4 is related to figuring out the genomic distances over which transcription issue binding influences downstream expression, which is effective for deciphering regulatory variants and inferring goal genes from transcription issue genome occupancy knowledge. Others beforehand systematically built-in 1000’s of transcription issue binding and histone modification profiles assayed by ChIP–seq with 1000’s of gene expression profiles to establish two courses of transcription issue with distinct ranges of regulatory affect. We examined Geneformer’s capacity to differentiate these long- versus short-range transcription elements utilizing solely single-cell transcriptomes from cells present process iPSC-to-cardiomyocyte differentiation with no related ChIP–seq or genomic distance knowledge. This higher-order transcription issue property of regulatory vary is a very difficult attribute to deduce from transcriptional knowledge alone.
Zero-shot evaluations have been carried out by coaching a linear layer on frozen mannequin embeddings to judge separability of the zero-shot gene embedding area. Few-shot evaluations have been carried out by fine-tuning with solely 100 instance cells. Effective-tuning was carried out with 10,000 instance cells. All layers have been unfrozen to permit weight updates in each the few-shot and fine-tuning evaluations. Different fashions together with help vector machines, random forest and logistic regression have been fine-tuned as beforehand described6. Geneformer fashions used a most studying price of 0.0005 for GF-30M and all quantized fashions and 0.0001 for GF-104M and GF-316M, as bigger fashions usually require decrease studying charges for secure coaching. In any other case no hyperparameter optimization was carried out, and all fashions have been skilled for 1 epoch with warm-up ratio of 1%, studying price decay on a cosine schedule and batch dimension of 1 (standardized for time measurements in contrast with quantization). QLoRA fashions have been skilled with a rank of 128 and alpha of 256. Effective-tuning was examined with 3 completely different seeds for all fashions, coaching on 80% of the genes and predicting on the held-out 20% of genes.
Cell-level job analysis
We examined the separability of the zero-shot versus fine-tuned cell embedding area of full-precision and quantized fashions by tissue and cell-type attributes and illness states. Zero-shot evaluations have been carried out by coaching a linear layer on frozen mannequin embeddings whereas fine-tuning was allowed to replace weights in all layers of the mannequin. Hyperparameter optimization was carried out utilizing Ray/HyperOpt for 15 trials for the tissue and cell-type job and 40 trials for the illness job, optimizing studying price, warm-up ratio, dropout price, gradient normalization and weight decay for every mannequin. QLoRA fashions have been skilled with a rank of 32 and alpha of 64. Efficiency utilizing the Geneformer embedding area to differentiate tissues and cell sorts was in contrast with utilizing scikit-learn’s implementation of help vector machines, random forest and logistic regression skilled on both uncooked counts or extremely variable genes chosen from the coaching knowledge.
Relatively than evaluating every tissue individually, the embedding area was collectively evaluated for separability throughout all 159 courses of concatenated tissue and cell subclass labels inside a random subsample of regular, grownup cells throughout the uniformly annotated CELLxGENE corpus. We used 434,994 cells for coaching, a separate 144,999 cells have been used for validation throughout hyperparameter tuning and a separate 248,569 cells have been held-out within the check set used for the ultimate reported analysis metrics. For the illness classification, the embedding area was evaluated for separability throughout 78 courses (77 illness courses, 1 regular class) inside a random subsample of cells throughout the uniformly annotated CELLxGENE corpus. We used 20,000 cells for coaching, a separate 10,000 cells have been used for validation throughout hyperparameter tuning and a separate 10,000 cells have been held-out within the check set used for closing reported analysis metrics. The ailments included 23 most cancers sorts and 54 non-cancer ailments.
In silico perturbation
We carried out in silico perturbation experiments as beforehand described6. In silico deletion was modeled by eradicating the given gene from the rank worth encoding of the given single-cell transcriptome and quantifying the cosine similarity between the unique and perturbed gene embeddings of the remaining genes within the single-cell transcriptome to find out which genes have been predicted to be most delicate to in silico deletion of the given gene. Of notice, the cosine similarity metric doesn’t instantly decode to numerical values of gene expression, however as an alternative represents the mannequin’s prediction of the extent of affect of the given perturbation on one another gene.
We examined the affect on iPSC-derived cardiomyocytes19 of in silico deleting GATA4, a identified congenital coronary heart illness gene, on housekeeping genes37 in contrast with GATA4 direct versus oblique goal genes. GATA4 direct goal genes have been outlined as any gene considerably dysregulated in response to the GATA4 variant the place GATA4 certain inside 20 kb of the gene’s transcriptional begin web site by ChIP–seq within the iPSC illness mannequin20, whereas oblique goal genes have been genes considerably dysregulated in response to the GATA4 variant with out GATA4 binding inside 20 kb of their transcriptional begin web site. We examined the affect of in silico perturbation on these genes throughout the high two quartiles of detections within the cells.
Reporting abstract
Additional data on analysis design is obtainable within the Nature Portfolio Reporting Abstract linked to this text.