Improvement of Omnitrap UVPD, ECD and EID LC-MS strategies
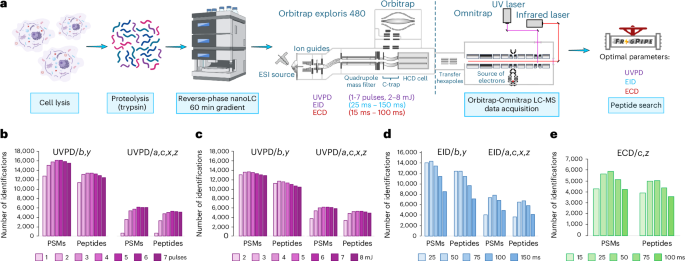
The outcomes of our latest improvement and characterization of UVPD, EID and ECD on the Omnitrap platform36 recommended that it could possibly be deployed in an LC-MS configuration for the evaluation of complicated peptide mixtures. On condition that the circumstances in direct-infusion experiments from our earlier work, similar to variety of obtainable ions, injection occasions and ion switch logistics, are usually extra relaxed than in automated LC-MS evaluation, an investigation is required to find out the optimum parameters for all dissociation methods. Direct-infusion experiments reported beforehand36 had been targeted on greater decision and signal-to-noise ratio with no regard to responsibility cycle. On condition that acquisition of spectra with this configuration has restricted parallelization potential (Prolonged Knowledge Fig. 1a), we initially focused on decreasing scan size to extend pace of spectra acquisition to deal with the complexity of proteomes. The Omnitrap design requires ions to be cooled by way of a gasoline pulse previous to any ion manipulation. The unique design used a single gasoline valve that had a most repetition price of 10 Hz (Prolonged Knowledge Fig. 1b). To enhance the utmost price of the Omnitrap we applied the usage of two valves, working alternately, for gasoline injection, which may probably double the pace (Prolonged Knowledge Fig. 1b). Subsequently, we optimized the potentials for ion switch within the Omnitrap to scale back the background collisional fragmentation (Supplementary Notes and Prolonged Knowledge Fig. 1c–f). We then targeted on rising the identification price in LC-MS experiments by way of software of pragmatic parameters for acquisition (Fig. 1a). Except in any other case specified, human Expi293F cell lysate digests had been used because the analyte. We started with the characterization of UVPD. We first various the variety of laser pulses at a hard and fast vitality of three mJ per pulse after which various the vitality for a hard and fast variety of pulses. For knowledge evaluation, we began with utilizing solely b and y ions for identification, which had been beforehand proven to be probably the most considerable in UVPD of tryptic peptides6,37,38. Evaluation reveals that rising the variety of laser pulses results in a larger variety of recognized peptide–spectrum matches (PSMs) and peptide sequences till a most is reached at 4 pulses (Fig. 1b). Additional will increase within the variety of laser pulses used for dissociation leads to a drop of the identification price, both because of secondary fragmentation or diminished scan price. We chosen 4 pulses for additional investigation and various the vitality of every pulse. On this sequence of experiments, the utmost of recognized PSMs and peptide sequences was noticed at distinct energies relying on the kind of fragment ions used for identification (Fig. 1c). Utilizing solely b and y fragments, the utmost is noticed at 5 mJ per pulse, whereas when different varieties of fragment attribute of UVPD are used, specifically a, c, x, z (ref. 4) (see Supplementary Desk 1 for constructions and definitions of fragment ions thought-about on this work), the utmost is situated at 6 mJ per pulse. On condition that a, c, x, z in distinction to b, y are extra distinctive to UVPD, we opted to make use of 6 mJ per pulse in future experiments.
a, Experimental workflow. b–e, Variety of PSMs and peptides recognized in UVPD experiments various the variety of UV laser pulses at 3 mJ pulse−1 (b), UVPD experiments utilizing 4 laser pulses and ranging the heartbeat vitality (c), EID experiments various the irradiation time at 25 eV of electron vitality (d), and ECD experiments various the irradiation time at ~1 eV of electron vitality (e). In UVPD and EID, b, y or a, c, x, z fragments had been used for knowledge evaluation; c and z ions had been used within the evaluation of ECD knowledge. Schematic diagram in a created in BioRender; Govender Kirkpatrick, M. (2025).
Subsequent, we studied the optimum response occasions for ExD. In typical ExD experiments, ions are transferred into the response chamber and bear irradiation by electrons emitted by a heated filament35 throughout a specified period of time (Prolonged Knowledge Fig. 1g,h). In EID experiments, we various the irradiation time from 25 ms to 150 ms and measured the variety of recognized PSMs and peptides. We noticed that b and y ions will be probably the most distinguished ions in EID. When utilizing solely these two ions for evaluation, the variety of PSMs and of peptides reaches the utmost worth at 50 ms of irradiation (Fig. 1d). At longer irradiation occasions, these numbers begin to drop. Curiously, the profile of peptide identification reveals a way more distinctive dependence on the kind of ions used for evaluation in contrast with UVPD (Fig. 1d). At shorter irradiation occasions, a, c, x, z fragments are underrepresented in contrast with these of b, y, and the most important variety of PSMs and peptides was noticed at 75 ms (Fig. 1d). To maintain scan charges excessive within the curiosity of absolute variety of identifications, we selected to proceed with the 50 ms irradiation time. Lastly, we discovered 50 ms of irradiation to be optimum in ECD utilizing c and z fragments for the information evaluation (Fig. 1e). We didn’t examine different main-series varieties of fragments, as a result of the vast majority of the merchandise of ECD of comparatively quick peptides are c and z ions7,8. On condition that ECD is thought to be a charge-dependent course of favoring greater cost states, the worth of fifty ms obtained utilizing primarily doubly charged and fewer steadily triply charged precursors of tryptic peptides will be thought-about conservative. To characterize the fragmentation habits of ECD, UVPD and EID, a bigger and extra various vary of peptides is required.
Giant-scale multi-enzyme LC-MS evaluation
We elevated the range of peptide sequences by way of the usage of extra proteases, and we elevated peptide depth by using offline reverse-phase high-pH fractionation (Fig. 2a). We selected trypsin, LysC, GluC, chymotrypsin and LysN as a result of they’ve been proven to supply complementary outcomes when it comes to peptide size, protein sequence protection, and frequencies and positions of amino acid residues throughout the peptide spine39. Subsequent, we fractionated every digest40 into 20 pooled fractions and analyzed all of them utilizing ECD, EID, beam kind CID (known as higher-energy CID or ‘HCD’ on Thermo instrumentation) and UVPD LC-MS. The selection of liquid chromatography gradient time for the dissociation methods was primarily based on their most sequencing price to make sure that all of them produced the same variety of scans.
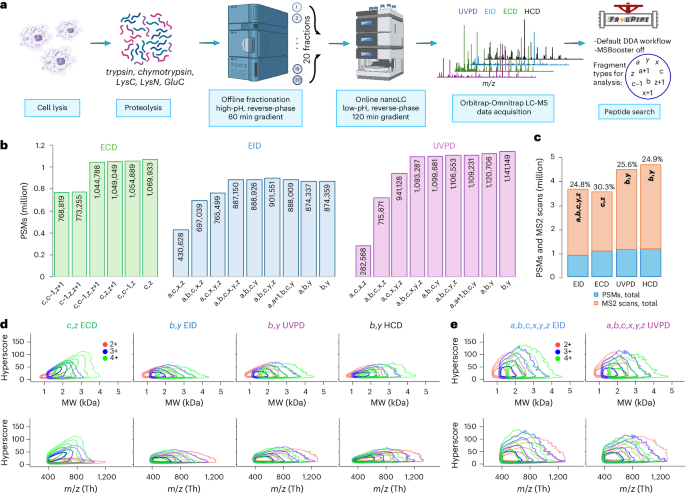
a, Experimental workflow. b, Whole numbers of PSMs in ECD, EID, and UVPD experiments recognized utilizing completely different mixtures of fragment varieties. c, Highest variety of PSMs from b (blue) and whole variety of acquired MS2 scans (orange) in ECD, EID, UVPD and HCD experiments; the speed of PSM identification is proven above every corresponding bar. d, Density contour plots of hyperscore distributions of two+, 3+ and 4+ cost states of distinctive PSMs (distinctive mixture of amino acid sequence, cost and modification chosen by highest hyperscore) acquired in ECD utilizing c and z fragments and in EID, UVPD and HCD utilizing b and y fragments. e, Density contour plots of hyperscore distributions of two+, 3+ and 4+ cost states of distinctive PSMs acquired in EID and UVPD utilizing a,b,c,x,y,z fragments. Contour strains demarcate the smallest areas to comprise 50%, 80%, 95% and 99% of factors. Schematic diagram in a created in BioRender; Govender Kirkpatrick, M. (2025).
The evaluation of UVPD, EID and ECD knowledge will not be as simple as that of HCD knowledge. The key merchandise of HCD are effectively characterised, with a, b, y ions dominating knowledge. In distinction, UVPD and EID are identified to supply all main-series varieties of peptide fragments in addition to some radical a + 1, x + 1 ions,4,15,41 with the final two largely understudied. The typical proportion of every kind of main-series fragment has been reported for UVPD6,37,38; nevertheless, the consequences of utilizing these ions and their mixtures within the automated knowledge evaluation haven’t been extensively mentioned. We due to this fact analyzed the acquired uncooked knowledge utilizing a number of distinctive mixtures of the anticipated fragment varieties with the aim to maximise the variety of recognized PSMs whereas sustaining the identical 1% false discovery price (FDR). For ECD, crucial ions for sturdy identification had been c and z (Fig. 2b). The addition of c − 1 or z + 1 had a minimal and barely detrimental impact. Analogously, b and y had been the dominant ion varieties for each EID and UVPD. Nevertheless, a, a + 1, c, z ions had been useful for enhancing identification charges for EID, whereas b, y produced the perfect leads to UVPD. The numbers when damaged right down to the person enzyme degree are just like the worldwide end result, though tryptic and LysC peptides improve the formation of z + 1 ions whereas impairing the formation of c − 1 in ECD, and favor the era of y ions in EID and UVPD in contrast with different enzymes (Supplementary Fig. S1). The outcomes for UVPD and EID appear to be strongly depending on y ions and to a smaller diploma on b ions. Whereas no in depth literature exists for EID, our UVPD knowledge agree with earlier findings. Others additionally discovered that b, y fragments are probably the most considerable varieties of ions in 193 nm UVPD of tryptic peptides, and the ion present of y fragments is roughly double that of b (refs. 6,37). Equally, b, y fragments dominate the spectra in 213 nm UVPD of tryptic peptides, and the typical variety of annotated y fragments is twice that of b ions38.
In whole, every fragmentation method produced between roughly 3.5 million and 4.5 million MS2 spectra throughout 5 enzymes, 20 fractions per enzyme (Fig. 2c). EID knowledge had the least variety of PSMs ( ~900,000), whereas UVPD, which has the quickest acquisition price amongst all Omnitrap methods studied right here ( ~6.3 MS2 scans per second on common), had 1,141,000 (Fig. 2c). Surprisingly, charge-dependent ECD got here closest to UVPD with 1,070,000 PSMs, though its scan price ( ~5.2 MS2 spectra per second) was basically the identical as in EID. HCD confirmed the very best numbers with 1,160,000 PSMs acquired utilizing 60 minute gradients on the price of, on common, ~13 MS2 scans per second. Pleasingly, the effectivity of peptide sequencing by EID (24.8%) and UVPD (25.6%), expressed because the ratio of the variety of confidently recognized PSMs to that of acquired MS2 scans, is actually the identical as by HCD (24.9%), whereas the effectivity of sequencing by ECD (30.3%) was the perfect (Fig. 2c). This was stunning contemplating the relative inefficiency of ECD for doubly charged peptides, which symbolize a considerable subset of recognized peptides (Prolonged Knowledge Fig. 2a).
The MSFragger hyperscore can function an oblique measure of the variety of fragments present in a spectrum, just like a spectrum high quality rating42. We plotted density contour plots for hyperscores of all distinctive precursors (that’s, distinctive mixtures of amino acid sequences, cost states and modifications, Prolonged Knowledge Fig. 2b,c) per cost state utilizing c, z fragments in ECD and b, y fragments in UVPD, EID and HCD (Fig. second and Supplementary Figs. S2 and S3). Expectedly, the distribution of hyperscores in ECD is strongly cost dependent, with doubly charged precursors assigned considerably decrease values. Moreover, the hyperscore distributions for 3+ and 4+ precursors in ECD have an obvious most at 800 Th. An identical pattern was reported earlier by Good et al. for ETD of tryptic and LysC peptides, by which the % of bonds cleaved by ETD begins to drop at roughly 600 Th for 3+ precursors and 650 Th for 4+ ones13. When analyzing solely b, y ion sequence, EID, UVPD and HCD all produce very related hyperscore distributions for a similar cost states of precursors (Fig. second). UVPD has marginally greater hyperscores within the low-m/z vary than HCD, and EID produces decrease hyperscores within the high-m/z vary than UVPD and HCD. The higher boundary of hyperscore distributions for these dissociation methods begins to drop past roughly 2,000–2,500 Da for two+ and three+ precursors and a pair of,500–3,000 Da for 4+ precursors. We interpret these observations because the discount of the signal-to-noise ratio that follows the spreading of obtainable fragment sign throughout a bigger variety of produced fragments in spectra of lengthy and extremely charged peptides, that’s, sign splitting. The distinction in variety of identifications with the identical 1% FDR was marginal for UVPD and EID after we elevated the variety of fragment varieties all the best way as much as a, b, c, x, y, z, so long as the b, y fragments had been included (Fig. 2b). We due to this fact investigated how the selection of kind of fragment for evaluation impacts hyperscores (Fig. 2e and Supplementary Fig. S4). Clearly, including extra varieties of fragments leads to vastly improved hyperscores for each EID and UVPD, indicating a bigger variety of dissociated bonds and data-rich spectra.
Deep studying modeling of UVPD, EID and ECD fragment intensities
PSM scoring will be improved considerably if carried out towards experimental or in silico-generated spectral libraries32. Deep studying fashions have demonstrated promising leads to predicting CID-based spectra of peptides utilizing solely peptide sequence, cost state and collision vitality as enter26,27,28,31, however no such fashions exist for different fragmentation methods as a result of lack of enormous quantities of high-quality knowledge for coaching. We due to this fact got down to use the datasets generated on this work to coach a deep studying mannequin in a position to predict fragment ion intensities. To create a extra complete mannequin we then generated the same dataset for electron-transfer/collision-induced dissociation (ETciD) on a Thermo Tribrid instrument (Supplementary Notes). Coaching a deep mannequin requires changing the uncooked knowledge right into a dataset containing accurately annotated peak intensities. This means that we have to remedy potential clashes similar to, for instance, a + 1 ion, which is a radical a ion coupled with a further hydrogen atom, versus the 13C peak for an a ion. For all datasets, we carried out an automatic annotation of main fragment varieties anticipated in EID, ECD, ETciD and UVPD (Supplementary Desk 1) utilizing the Oktoberfest framework30. The comparability of [a + 1]/[a] ratio in HCD, EID and UVPD means that a big proportion of a + 1 in EID and UVPD spectra originate from gas-phase electron- and photon-based chemistries (Fig. 3a, Prolonged Knowledge Fig. 3, Supplementary Figs. S5–S9 and Supplementary Notes). With the annotated spectra in hand, we outlined our mannequin’s ion dictionary and curated coaching and validation datasets. The unique Prosit mannequin27 structure was designed round a structured output house consisting of b and y fragments with lengths 1–29 and expenses +1 to +3. Against this, the mannequin educated on our knowledge has an unstructured output house, with fragment ions chosen primarily based on frequency of prevalence (≧100 occurrences, Supplementary Figs. S5–S9). The mannequin additionally takes the specific fragmentation kind as enter; provided that the HCD knowledge had been acquired on a single instrument, it was pointless to make use of collision vitality as extra enter to the mannequin, as was carried out for earlier Prosit fashions27. Our mannequin shares similarity with the unique Prosit mannequin in that the sequence and metadata are individually encoded into latent areas and mixed within the inside of the community, however the metadata have barely modified, and the mannequin outputs predicted intensities of 815 fragment ions of assorted size, cost and fragment kind (Fig. 3b). Outcomes present little or no overtraining: the median Pearson correlations for ECD, UVPD, HCD and EID are 0.919, 0.931, 0.950 and 0.897, respectively, on the coaching set, and the corresponding scores for the check set are solely ~0.005 decrease for every fragmentation methodology (Fig. 3c and Prolonged Knowledge Fig. 4). Moreover, we observe that precursor cost is consequential for prediction efficiency, with precursor expenses larger than 2 having an more and more wide selection of Pearson correlations, prone to be as a result of sparsity of excessive cost precursors within the coaching set and more and more complicated fragment ions current within the spectra. Pleasingly, we see that conditioned on the fragmentation methodology the mannequin reliably assigns considerable depth solely to these fragments anticipated for every fragmentation methodology, for instance b, y for HCD and c, z for ECD (Fig. 3d,e). The mannequin can be in a position to predict intensities of b, y and minor fragments, similar to a, a + 1, x, x + 1, c, z in UVPD and EID, though predictions of low-intensity ions for the latter appear barely much less correct (Fig. 3f,g). We carried out a sequence of extra assessments to validate the robustness and correctness of our mannequin (Supplementary Notes and Supplementary Fig. S10).
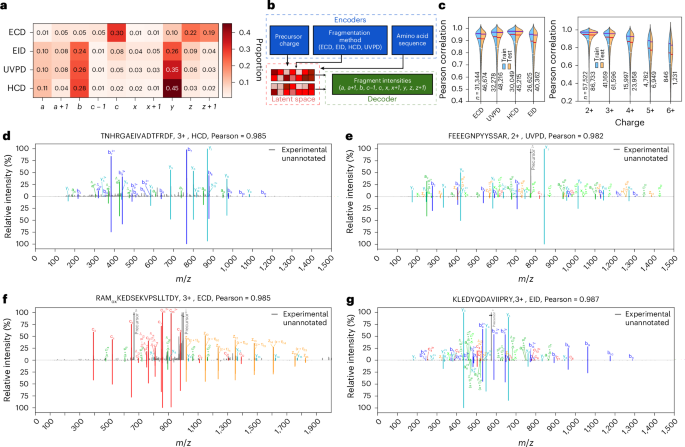
a, Heatmap of imply proportion of every kind of fragment ion amongst all annotated peaks in ECD, EID, HCD and UVPD spectra acquired throughout all enzymes, not reflecting relative depth of ions. Annotation was carried out for 10 ion varieties: a, a + 1, b, c − 1, c, x, x + 1, y, z, z + 1 (Supplementary Desk 1). b, The modified Prosit deep studying structure for prediction of fragment ion intensities in ECD, EID, HCD and UVPD spectra. The enter parameters (peptide sequences, precursor cost state and fragmentation methodology) are encoded right into a latent illustration (latent house). This illustration is then decoded to foretell fragment ion intensities. c, Pearson correlation coefficients between predicted and experimental spectra in coaching and check units separated by fragmentation methodology (left) and cost state (proper). Horizontal white, crimson, and blue strains correspond to 25%, 50% and 75% percentiles, respectively. n signifies pattern dimension. Distributions extending past 1.0 are plotting artefacts. d–g, Mirror plots of chosen precursors in HCD (d), UVPD (e), ECD (f) and EID (g) knowledge. Every mirror plot compares experimental (high) and predicted (backside) fragment intensities, with every fragment kind uniquely coloured.
Rescoring of different fragmentation knowledge utilizing fragment depth predictions
An environment friendly management of FDR in database looking out is vital for identification of true-positive peptide matches. Beforehand, we confirmed that data-driven rescoring of CID knowledge utilizing the Prosit mannequin vastly improved quantity and accuracy of peptide identifications27. We hypothesized that predicting fragment ion depth can be useful for enhancing the outcomes of the database searches of UVPD, EID and ECD knowledge as effectively. Utilizing the optimized MSFragger outcomes we first calculated the ratio of the variety of all noticed to that of all attainable theoretical fragment ions in every recognized spectrum (Fig. 4a and Prolonged Knowledge Fig. 5, higher distributions). The ensuing distributions for goal and decoy (a priori false-positive) PSMs had been closely intermixed and shifted in direction of smaller ratios. EID and UVPD ratios had been significantly small because of a lot of theoretical ions. We then calculated the identical ratios however allowed solely fragments predicted by Prosit (Fig. 4a and Prolonged Knowledge Fig. 5, decrease distributions). The inclusion of solely predicted fragments cut up the distribution of ratios of goal PSMs, by which the bulk shifted in direction of greater values with a bigger portion being above 0.8, and the rest had been basically unchanged. On the similar time, the ratio of decoy PSMs remained clustered at decrease values. This means a considerable enchancment within the alignment between the noticed and predicted fragment ions.
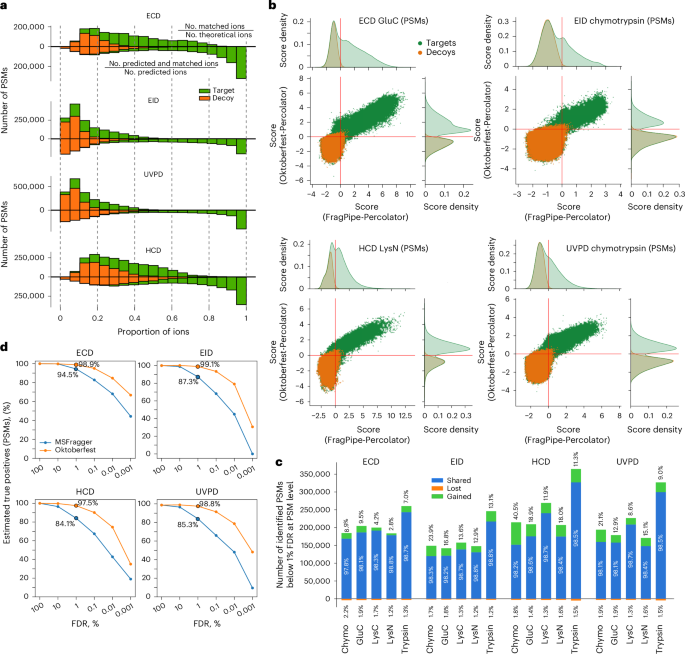
a, Histogram of the ratio of experimentally noticed ions to all theoretically attainable fragments (higher distributions); and histogram of the ratio of predicted and experimentally noticed ions to all predicted ions (decrease distributions). b, Correlation of Percolator scores for all goal and decoy PSMs obtained from the rescoring of the MSFragger (high) and Oktoberfest (proper) units of scores for chosen mixtures of enzyme and fragmentation method. The crimson stable strains point out the 1% PSM-level FDR cut-offs. For database search scores, the perfect mixtures of fragment varieties from Fig. 2b had been used; for Oktoberfest scoring, most steadily annotated fragment varieties ( >4% of all annotated ions throughout all spectra) had been used for every dissociation methodology (Prolonged Knowledge Fig. 3). c, Variety of shared, gained and misplaced PSMs recognized at 1% PSM-level FDR utilizing the Oktoberfest set of scores in comparison with the unique MSFragger seek for every fragmentation method per enzyme. The numbers correspond to the information from b and Supplementary Figs. S11–S14. Chymo, chymotrypsin. d, Proportion of the variety of true-positive PSMs to the estimated most variety of true-positive PSMs acquired utilizing unique MSFragger and Oktoberfest scores at completely different values of PSM-level FDR for every fragmentation method, all enzymes mixed.
Subsequent, we utilized data-driven rescoring utilizing the Oktoberfest framework, which advantages from the here-developed fragment ion depth prediction mannequin by producing fragment intensity-dependent scores moderately than relying solely on the presence or absence of any theoretical fragments. Together with Percolator43, these scores are aggregated right into a single rating that maximizes the separation of appropriate and incorrect matches. The ensuing Oktoberfest scores had been then in comparison with the Percolator-derived scores from MSFragger (Fig. 4b and Supplementary Figs. S11–S15), which don’t embody fragment intensity-based options. For MSFragger database searches, we selected the perfect mixture of ion varieties for every fragmentation methodology from Determine 2b, and for rescoring in Oktoberfest we used all the most steadily annotated varieties of fragments ( >4% of annotated ions in a spectrum, averaged throughout all spectra) for every fragmentation method (Prolonged Knowledge Fig. 3). Each units of scores had been filtered to 1% FDR utilizing Percolator43. Whereas rescoring led to outstanding separation of decoys from targets for almost all of enzyme–fragmentation methodology pairs (Fig. 4b and Supplementary Figs. S11–S15), ECD generally demonstrated adequate separation in database searches, such that rescoring delivers solely marginal enhancements in identification (Supplementary Fig. S11). This partly explains the very best identification price noticed for ECD within the preliminary database searches (Fig. 2c). We attribute this to the relative cleanliness of ECD spectra that consist primarily of c, z fragments, precursor ions and charge-reduced species, thus decreasing possibilities for random false matches. Curiously, ECD was the one method by which it was attainable to discriminate the distributions of cost states amongst goal PSMs after rescoring, which displays the distinct charge-dependent kinetics of this course of (Supplementary Fig. S16). Utilizing rescoring, we had been in a position to salvage a considerable variety of PSMs in all mixtures of enzyme and dissociation methodology (quadrant II in Fig. 4b and Supplementary Figs. S11–S15). On the similar time, a excessive variety of PSMs initially recognized had been discarded (quadrant IV in Fig. 4b and Supplementary Figs. S11–S15).
To guage how this separation of scores translated into positive factors and losses of PSMs and peptides, we in contrast the outcomes of the database search and rescoring at each 1% PSM-level (Fig. 4c) and 1% peptide-level FDR (Supplementary Figs. S17 and S18). The variety of gained PSMs various (relying on the enzyme and fragmentation methodology) between roughly 3% and 40.5%, with chymotrypsin HCD knowledge producing a notable achieve of 40.5%. The latter remark is in keeping with our earlier findings27. Remarkably, chymotrypsin was additionally the primary beneficiary of rescoring in UVPD and EID knowledge. This demonstrates the usefulness of rescoring for expanded search areas characterised by an elevated variety of attainable cost states, allowed missed cleavages and diminished enzyme specificity, all of that are typical for chymotrypsin (Prolonged Knowledge Fig. 2a). In step with the rating distributions (Fig. 4b and Supplementary Figs. S11–S15), ECD had the bottom variety of gained PSMs and peptides no matter protease amongst all fragmentation methods (Fig. 4c and Supplementary Fig. S17). Additional investigation of ECD knowledge reveals that prediction of retention time and of fragment depth generated related positive factors, every including roughly 6.5% of PSMs (Supplementary Notes and Prolonged Knowledge Fig. 6). Such a comparatively modest contribution of retention time predictions reveals that enhancements noticed after rescoring of different mixtures of enzyme and fragmentation method are primarily pushed by the brand new Prosit mannequin.
To discover the explanations for the various variety of positive factors noticed, we investigated the restoration of estimated true-positive PSMs. We in contrast the variety of estimated true positives throughout a spread of FDR thresholds (by subtracting the variety of decoy PSMs from the variety of goal PSMs at completely different FDR cut-offs) earlier than and after rescoring with the full variety of estimated true positives within the dataset that could possibly be recovered from the preliminary search outcomes, by subtracting the full variety of decoys from the full variety of goal PSMs (Fig. 4d and Supplementary Fig. S19). At 1% PSM-level FDR, rescored ECD, EID and UVPD searches recovered greater than 97% of attainable true positives, whereas the unique database searches extracted roughly 95% in ECD, 87% in EID, 85% in UVPD, and 84% in HCD. At a stricter FDR of 0.01%, the outcomes after rescoring nonetheless captured greater than 75% of all estimated attainable true positives, with ECD exhibiting the very best proportion approaching 85%. On the similar FDR degree, preliminary database searches recognized lower than 70% of attainable true positives in ECD and fewer than 55% in all different dissociation strategies (Fig. 4d). The evaluation reveals that data-driven rescoring utilizing the pan-fragmentation Prosit mannequin considerably will increase the proportion of estimated true-positive PSMs retained at stringent thresholds, approaching saturation of the set of PSMs recoverable from the preliminary MSFragger search outcomes. It is very important observe that additional appropriate identifications, for instance from modified peptides not thought-about within the preliminary search, can’t be thought-about within the estimation of the variety of true positives.
The rescoring knowledge offered a possibility to examine the efficacy of every enzyme and dissociation method for proteome evaluation (Supplementary Notes, Prolonged Knowledge Figs. 7 and eight and Supplementary Figs. S20–S24). Trypsin, as anticipated, recognized probably the most PSMs, peptides and proteins for each fragmentation method. Chymotrypsin had the subsequent greatest end result, with LysC and LysN barely additional behind (Prolonged Knowledge Fig. 7a and Supplementary Fig. S20a), replicating earlier developments noticed for CID and ETciD knowledge44,45,46. The enzyme GluC clustered with LysN, showing to be barely superior or inferior relying on the dissociation method. Common protein sequence protection was related for every fragmentation method (Prolonged Knowledge Fig. 8). To evaluate complementarity on the protein sequence degree we represented our knowledge on the amino acid degree. Normally phrases, when evaluating the complementarity of trypsin towards its options, we noticed substantial enhancements in proteome protection for all fragmentation methods (Prolonged Knowledge Fig. 7b and Supplementary Fig. S20b); in reality, the distinctive mixed protection for LysN, LysC, GluC and chymotrypsin was greater than that for trypsin. These observations echo earlier work demonstrating the complementarity of enzymes for enhancing sequence protection39,44,45,46. It ought to be famous that every trypsin fraction was basically analyzed with LC-MS 4 occasions, and a extra exhaustive LC-MS evaluation wouldn’t considerably improve proteome protection, and therefore the quantity of study time for the opposite enzymes versus trypsin will not be an necessary issue within the comparability. Additional evaluation of distinctive protection for every fragmentation method confirmed that UVPD produced probably the most quantity of distinctive knowledge, with HCD and ECD shut behind, and EID the least (Prolonged Knowledge Fig. 7c). Nevertheless, UVPD had vital overlap with EID, which could be a purpose for the weak distinctive proteome protection end result for EID (Prolonged Knowledge Fig. 7c).
Utility of data-independent acquisition in all fragmentation methods
The spectral prediction mannequin created on this work is moveable and freely obtainable as ’Prosit_2025_intensity_MultiFrag’ on the Koina mannequin repository47, and will be interfaced from inside any software program suite. We applied our mannequin inside FragPipe as a part of MSBooster29. We reanalyzed the deep proteome knowledge in MSFragger to match the outcomes with and with out MSBooster and located very related positive factors to these noticed utilizing Oktoberfest at each the PSM and peptide ranges (Prolonged Knowledge Fig. 9). Mixed with the optimization of search parameters in FragPipe, we will now carry out each data-dependent and data-independent acquisition (DDA and DIA, respectively) analyses (pseudo-DDA by way of the usage of DIA-Umpire) for all activation methods. The flexibility to now make the most of these activation methods with DIA approaches led us to create DIA methodologies for the Orbitrap-Omnitrap. The change in ion inhabitants, each when it comes to ion density and distribution of cost states, required adjustment of the acquisition parameters for every dissociation method each on the Exploris and Omnitrap degree (see Strategies). We carried out LC-MS analyses on unfractionated tryptic cell lysate digests from Homo sapiens (Expi293F), Arabidopsis thaliana and Escherichia coli cells. We launched the final two varieties of cells to evaluate the universality of the Prosit mannequin. To optimize responsibility cycle, we selected to make use of the ‘normal isolation window’ method with MS1 vary sure to retention time48. MSBooster, utilizing the here-developed Prosit mannequin, elevated identification price on the PSM, peptide and protein ranges for all three cell varieties. The A. thaliana and H. sapiens lysate samples had the most important enhancements, buying and selling high place relying on actual context. On common, ECD had the bottom positive factors throughout all samples, with the worst end result being 1.0%, 1.7% and three.0% on the three ranges for E. coli, whereas EID demonstrated the most important enhancements throughout all three varieties of samples, with the perfect end result being 31.4%, 20.9% and 22.6% on the three ranges for the A. thaliana pattern (Fig. 5).
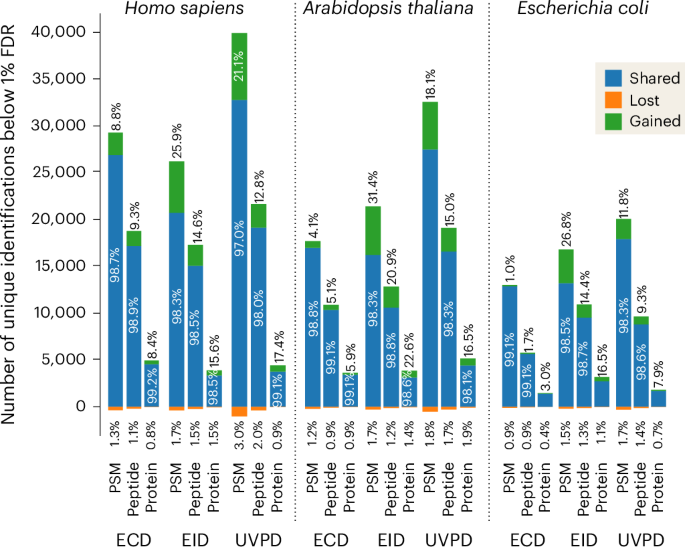
Variety of PSMs, peptides and proteins recognized at 1% FDR within the UVPD, EID and ECD DIA knowledge of unfractionated tryptic digests of human, A. thaliana and E. coli proteins. The evaluation was carried out within the FragPipe platform utilizing the MSFragger search engine with Prosit predictions of fragment ion intensities applied throughout the MSBooster module. The numbers of shared, gained and misplaced identifications correspond to the evaluation with MSBooster ’on’ as in contrast with the outcomes obtained with MSBooster ’off’.