Information preprocessing
Single-modal information preprocessing
For spatial area identification, the gene expression counts had been log-transformed and normalized by library dimension by way of the SCANPY bundle62. The highest 3,000 extremely variable genes (HVGs) had been chosen and used as enter to the encoder. For the alignment of spatial slices, we reference SLAT22 and use SVD-based cross-dataset matrix decomposition technique to right inter-sample batch results.
Multimodal information preprocessing
To preprocess the transcriptomic information, pixels expressing fewer than 200 genes and genes expressing fewer than 200 pixels had been filtered out. Subsequent, the gene expression counts had been log-transformed and normalized by library dimension by way of the SCANPY bundle62. The highest 3,000 HVGs had been chosen and used as enter to principal-component evaluation (PCA) for dimensionality discount. For consistency with the chromatin peak information, the primary 50 principal parts had been retained and used as enter to the encoder. For the chromatin peak information, we used latent semantic indexing63 to scale back the uncooked chromatin peak counts information to 50 dimensions. For the antibodies (ADT) information, we first filtered out genes expressed in fewer than ten spots. The filtered gene expression counts had been then log-transformed and normalized by library dimension utilizing the SCANPY bundle. Lastly, the highest 3,000 HVGs had been chosen and used as enter for PCA. We utilized centered log ratio normalization to the uncooked protein expression counts. PCA was then carried out on the normalized information and the highest 30 principal parts had been used as enter to the encoder.
The 3d-OT framework
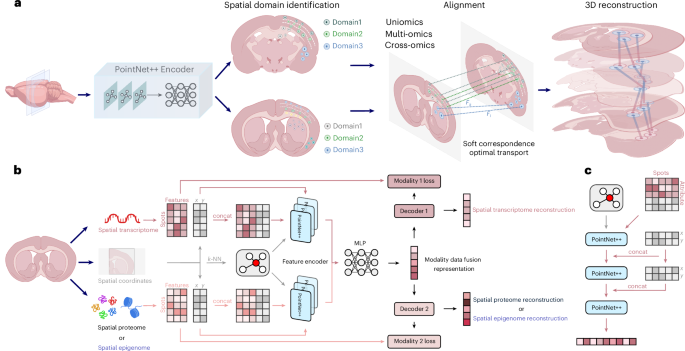
3d-OT is a brand new mannequin based mostly on multi-module integration, designed to leverage the spatial location info and expression information from a number of omics modalities throughout totally different spatial slices to attain fine-grained spatial area segmentation and correct heterogeneous slice alignment. The mannequin consists of two distinct modules: (1) PointNet++ Encoder; and (2) SCOT. Benefiting from the modular design, 3d-OT readily extends to spatial multi-omics information with greater than two modalities.
PointNet++ Encoder
PointNet++ Encoder relies on the purpose cloud deep-learning mannequin PointNet++. Initially designed to seize the native geometric info of level clouds, PointNet++ improved the classification, segmentation, and different geometric evaluation duties of 3D objects. By way of its hierarchical construction, progressively extracted options from native to international scales. This strategy successfully captured native gene expression and its spatial location correlations in spatial omics information. The mixing of two-dimensional (2D) spatial info allowed the mannequin to extra precisely distinguish complicated geometric constructions and object boundaries.
Development of neighbor graph
For every spatial omics slice (), we constructed a spatial neighborhood graph (G=(V,E)) utilizing the okay-NN algorithm, the place (V) and (E) denote the units of spots and edges, respectively. The adjacency matrix of (G) is denoted as (A), with (_ =1) if an edge exists between spots i and (j), and (0) in any other case. Based mostly on the expression function matrix derived from every slice and the neighborhood indices supplied by (G), we constructed a neighborhood function matrix (^) for every spot (_) in (). The (j)-th row, denoted as ( Q_^ Nleft(xright)in {}^{_{{rm}}}), represents the expression function vector of the (j)-th nearest neighbor of spot ( _ P). By stacking all spot-specific neighborhood matrices, we obtained a worldwide neighboring function tensor ({bf Q}in {{mathbb{R}}}^{Ntimes ktimes {d}_{{rm{modality}}}}) the place (N) denotes the entire variety of spots and ({d}_{{rm{modality}}}) represents the dimensionality of the omics options for the slice. This tensor served because the enter for subsequent integrative evaluation.
PointNet++ Encoder for particular person modality
Completely different omics information exhibit distinct function distributions. To encode these information right into a latent area, we employed an encoder framework constructed upon the PointNet++ structure. To include spatial info, we concatenated the precise spatial coordinates ({rm{spatial}}=(x,y)) of every neighboring spot to the corresponding native function matrix ({{bf{X}}}_{i}), during which ({{bf{X}}}_{i}) represents the i-th matrix of the tensor ({bf{X}}). Formally, for every spot i (=1,ldots ,N), the enter function matrix for the (l)-th layer of the PointNet++ Encoder is outlined as:
$${{bf{X}}}_{i}^{l}={rm{concat}}left({{bf{X}}}_{i},left(x,yright)proper)$$
(1)
the place (l) represents the layer index of the encoder and ((x,y)) are the vectors of spatial coordinates for the neighbors of spot i. This strategy is conceptually just like studying graph representations from the spatial graph (G), which captures bodily proximity between spots. By explicitly together with spatial coordinates, the mannequin is healthier in a position to understand the general distribution sample of the spots, thereby integrating native expression options with tissue spatial construction.
A PointNet++ Encoder layer consists of three successive convolutional layers, every adopted by batch normalization, and a last max pooling operation that yields the latent illustration. Particularly, after the preprocessing step for the (l)-th layer, the built-in attribute tensor ({{bf{X}}}^{l}in {{mathbb{R}}}^{Ntimes ktimes left({d}_{{rm{modality}}}+2right)}) is supplied as enter to the next transformation:
$${X}^{l}=textual content{MaxPooling}left({rm{BN}}left({rm{LeakyReLU}}left({rm{Conv}}left({{bf{X}}}^{l}proper)proper)proper)proper)$$
(2)
the place ({X}^{l}) denotes the latent illustration produced by the (l)-th PointNet++ layer. In our 3d-OT framework, three PointNet++ Encoder layers are stacked, and the ultimate illustration of the (m)-th modality is denoted as ({{rm{modality}}}_{m}{X}^{3}). The latent representations from totally different modalities are fed into an MLP to study a unified illustration:
$${Z}_{{rm{fused}}}={rm{MLP}}left({rm{concat}}left({{rm{modality}}}_{1}{X}^{3},{{rm{modality}}}_{2}{X}^{3}proper)proper)$$
(3)
the place ({Z}_{{rm{fused}}}) denotes the fused-modality illustration output by the MLP and ({{rm{modality}}}_{1}{X}^{3}) and ({{rm{modality}}}_{2}{X}^{3}) symbolize the latent representations from totally different modalities.
Module coaching in PointNet++ Encoder
To implement the realized latent illustration to protect the expression profiles from totally different modalities, we design a person decoder for every modality to reverse the built-in illustration ({Z}_{{rm{fused}}}) again into the normalized expression area. Particularly, by leveraging ({Z}_{{rm{fused}}}) from the between-modality MLP layer as enter, the representations ({X}_{{rm{output}}}^{l}) for every modality (min {mathrm{1,2}}) generated by the decoder on the (l)-th layer are formulated as follows:
$${mathrm{modality}}_{m}{X}_{mathrm{output}}^{l}=sigma left({X}_{mathrm{output}}^{l-1}{mathrm{modality}}_{m}{W}^{,l}+{{mathrm{modality}}_{m}b}^{l}proper)$$
(4)
the place ({W}^{,l}) and ({b}^{l}) correspond to the trainable modality-specific weights and biases at every layer, respectively, with (sigma) denoting the Rectified Linear Unit (ReLU) activation operate. The enter to the primary decoder layer is (Z_{rm{fused}}), and the output of the ultimate layer is (X^3_{rm{recon}}). The PointNet++ Encoder module is educated to attenuate the expression reconstruction loss:
$$start{array}{l}mathrm{Loss}=frac{1}{{it{N}}}mathop{sum }limits_{{it{N}}}^{{it{i}}=1},{left({mathrm{modality}}_{1}{{X}}_{{it{i}}}-{mathrm{modality}}_{1}{{X}}_{mathrm{recon},{it{i}}}^{3}proper)}^{2}+frac{1}{{it{N}}}mathop{sum }limits_{{it{N}}}^{{it{i}}=1} {left({mathrm{modality}}_{2}{{X}}_{{it{i}}}-{mathrm{modality}}_{2}{{X}}_{mathrm{recon},{it{i}}}^{3}proper)}^{2}finish{array},,,$$
(5)
the place ({{rm{modality}}}_{1}X) and ({{rm{modality}}}_{2}X) symbolize the preprocessed unique options of the modalities 1 and a couple of, respectively.
Alignment module
Impressed by earlier analysis64,65, we carried out an optimum transport-based framework to attain mushy correspondence alignment between supply and goal slices. Optimum transport supplies a mathematical framework for aligning chance distributions, whereas preserving their geometric properties, addressing the problem of redistributing mass from a supply to a goal distribution with minimal price.
To make the spatial construction of the estimated distribution higher match the goal distribution, we employed a mushy correspondence technique. Fairly than imposing strict one-to-one correspondence between spatial places, we leveraged chamfer distance inside an unsupervised optimization course of, prioritizing structural and spatial geometric properties because the optimization goal to attain versatile and correct alignment.
Fixing optimum transport for mushy correspondence
The optimum transport computation requires a function matrix (Phi in {{mathbb{R}}}^{ntimes d}), the place (n) denotes the variety of spatial spots and (d) is the function dimension. The matrix (Phi) will be instantiated in two methods:
(1) Because the fused illustration:
$$Phi ={Z}_{{rm{fused}}}in {{mathbb{R}}}^{ntimes d}$$
(6)
which is obtained from the MLP information fusion module; or
(2) Because the reconstructed single-modality illustration:
$$Phi ={{rm{modality}}}_{m}{X}_{{rm{recon}}}^{3}in {{mathbb{R}}}^{ntimes d}$$
(7)
the place ({{rm{modality}}}_{m}{X}_{{rm{recon}}}^{3}) denotes the reconstructed illustration of modality (m) after three successive PointNet++ Encoder layers.
Let ({Phi }_{{mathcal{X}}}) and ({Phi }_{{mathcal{Y}}}) symbolize the function matrices extracted from two enter spatial slices, ({mathcal{X}}) and ({mathcal{Y}}), respectively. We provoke by computing the spot-to-spot similarity matrix, given by:
$${S}_{i,j}=frac{{Phi }_{{x}_{i}}cdot {Phi }_{{y}_{j}}^{T}}{parallel {Phi }_{{x}_{i}}{parallel }_{2}parallel {Phi }_{{y}_{j}}{parallel }_{2}}$$
(8)
the place ({Phi }_{{x}_{i}}) represents the function of the i-th spot in ({Phi }_{{mathcal{X}}}) and ({Phi }_{{y}_{j},}) represents the function of the (j)-th spot in ({Phi }_{{mathcal{Y}}}). For every supply spot ({x}_{i}{mathscr{in }}{mathcal{X}}), we assigned an equal mass of (frac{1}{left|{mathcal{X}}proper|}) to make sure normalization. We then thought-about its transport to the goal spot ({y}_{i}{mathscr{in }}{mathcal{Y}}) with the associated fee matrix outlined as:
$${C}_{i,,j}=1{-S}_{i,,j}$$
(9)
On this formulation, a better price corresponds to a decrease similarity within the function area. We then apply the entropy-regularized Sinkhorn algorithm33 to establish the optimum transport plan ({T}^{* }):
$$start{array}{l}{T}^{* }=mathop{{rm{argmin}}}limits_{Tin {{mathbb{R}}}_{+}^{n}}left[mathop{sum }limits_{i,j}left(exp left(-displaystylefrac{{C}_{i,j}}{epsilon }right)times {{rm{support}}}_{i,j}right){T}_{i,j}right.left.qquad,+epsilon timesdisplaystylefrac{gamma +epsilon }{gamma }mathop{sum }limits_{i,j}{T}_{i,j}left(log {T}_{i,j}-1right)right]finish{array}$$
(10)
the place (gamma ={e}^{{rm{gamma}}}) and (epsilon ={e}^{{rm{epsilon}}}), with ({rm{gamma}}) and ({rm{epsilon}}) being learnable parameters optimized throughout coaching. The operator ({mathrm{help}}_{i,j}=)({(1-frac{sigma }{parallel {S}_{i,j}parallel })}_{+}odot {S}_{i,j}) acts as a thresholding operate. Right here, (epsilon) controls the power of entropy regularization, whereas (gamma) serves as a further tuning issue, modulating the convergence habits of the transport matrix by adjusting the step dimension of the Sinkhorn iterations.
The optimum transport plan ({T}^{* }) yields the mushy correspondence weight of spot ({x}_{i}) to identify ({y}_{j}):
$${W}_{i,j}=frac{{e}^{{T}_{i,j}^{* }}}{{sum }_{kin {{mathcal{M}}}_{{mathcal{Y}}}left({x}_{i}proper)},{e}^{{T}_{i,okay}^{* }}}$$
(11)
the place ({{mathcal{M}}}_{{mathcal{Y}}}({x}_{i})) denotes the set of (L) candidate spots from ({mathcal{Y}}) akin to the top-(L) values of ({T}_{i,j}^{* }), with (L) being a tunable hyperparameter. Based mostly on these weights, the estimated place of spot ({y}_{j}^{* }) is obtained as:
$${y}_{j}^{* }=mathop{sum }limits_{{y}_{j}{mathscr{in }}{mathcal{Y}}},{W}_{i,j}{y}_{j}$$
(12)
Lastly, the alignment move ({F}_{i}) from spot ({x}_{i}) to identify ({y}_{j}) is set as:
$${F}_{i}={y}_{j}^{* }-{x}_{i}$$
(13)
Self-supervised losses
As manually linking spots between spatial slices is notably intricate, we undertake self-supervised loss operate to coach the alignment module.
Reconstruction loss
A core precept guiding self-supervised alignment move studying is the truth that ({y}_{j}^{* }) and ({y}_{j}) must be comparable. The chamfer distance is a typical metric used to measure the form dissimilarity between level clouds in level cloud completion. We undertake it as our reconstruction loss:
$${L}_{{rm{recon}}}=frac{1}{| {y}^{{prime} }| }mathop{sum }limits_{{y}_{j}^{{prime} }in y},mathop{min }limits_{{y}_{j}in y}parallel {y}_{j}^{{prime} }-{y}_{j}{parallel }^{2}$$
(14)
the place ({y}^{{prime} }) represents the estimated spatial slice fashioned by ({y}_{j}) and (y) is the goal spatial slice fashioned by ({y}_{j}). By minimizing the chamfer distance, we make sure the consistency between the mushy matching connection goal and the estimated spatial slice location.
Clean loss
The precept of smoothing loss relies on native consistency and continuity, making certain that the alignment between adjoining slices stays easy and no abrupt modifications or discontinuities happen. Particularly in spatial transcriptomics or picture alignment issues, continuity of knowledge and easy transitions of spatial places are essential. The sleek loss is constructed as follows:
$${L}_{mathrm{easy}}=mathop{sum }limits_{{x}_{i}{mathscr{in }}{mathcal{X}}},mathop{sum }limits_{kin {{mathscr{N}}}_{l}left({x}_{i}proper)},frac{{F}_{i}-{F}_{okay}|_{1}}{left|{mathcal{X}}proper|left|{{mathscr{N}}}_{l}left({x}_{i}proper)proper|}$$
(15)
the place ({mathcal{X}}) represents the spatial slice fashioned by(,{x}_{i}), ({{mathscr{N}}}_{l}({x}_{i})) represents the index set of the (l) closest spots to(,{x}_{i}) and ({F}_{i}) and ({F}_{okay}) denote the estimated alignment move at spot ({x}_{i}) and ({x}_{okay}), respectively. As proven in Supplementary Fig. 2. The alignment course of is optimized by penalizing the alignment error between adjoining spots, making certain that the spatial location modifications easily all through the alignment course of.
Zero-divergence loss
Zero-divergence loss is just like easy loss in that it computes spatial gradients and requires the norm of the gradient to be small, basically penalizing the case that neighboring alignment move vectors are irrelevant; nevertheless, easy regularization is just too strict for spatial slice. Whereas the divergence constraint solely requires the entire divergence to be zero, it doesn’t necessitate that any two vectors be oriented in the identical route, thus permitting for regionally intricate spatial location variations (Supplementary Fig. 2b). In follow, we set the neighborhood set dimension for calculating easy loss to be a lot bigger than that for calculating zero-divergence loss, as a result of smoothness is a extra coarse-scale regularization. Lastly, impressed by 3D particle monitoring analysis28, we make use of an environment friendly splatter-based methodology for calculating the zero-divergence loss.
Splat-based implementation
To compute divergence, we want the partial by-product of the sector. The irregular association of spots in 2D spatial slice complicates this. Thus, we enhance upon earlier28, we splatting unstructured alignment move estimates onto a uniform 2D grid, then making use of zero-divergence regularization at these grid factors, as proven in Supplementary Fig. 2c. In formal phrases, the dense grid is denoted by ({({sj},{sk})}^{T}), with (j,kin {mathbb{Z}}) indicating the 2D indices of the grid level. The parameters (s) correspond to the grids spacing. Then, given a grid, we make use of the inverse squared distance as interpolation weights to approximate the alignment move at that spot:
$$Fleft(xright)=frac{1} Nleft(xright)mathop{sum }limits_{{x}_{i}in Nleft(xright)}frac{{F}_{i}}{{leftVert {x}_{i}-xrightVert }_{2}^{2}+epsilon }$$
(16)
the place ({F}_{i}) is the estimated alignment move worth at spot ({x}_{i}) and (N(x)) denotes the neighborhood among the many spot set of spatial slices ({mathcal{X}}) for grid level (x).
Divergence calculation
As soon as splatting has been employed, the divergence at that spot, specified by ({x}={({sj},{sk})}^{T}), will be outlined as:
$$left(nabla cdot {bf{F}}proper)left(xright)=mathop{sum }limits_{okay=1}^{2},frac{{bf{f}}left(x+s{u}_{okay}proper)-{bf{f}}left(x-s{u}_{okay}proper)}{2s}$$
(17)
the place ({u}_{okay}) is a unit vector with 1 on the (okay)-th entry. Lastly, the zero-divergence regularization will be formulated as:
$${L}_{{rm{div}}}=frac{1}{{JK}}mathop{sum }limits_{j=0}^{J-1},mathop{sum }limits_{okay=0}^{Okay-1},{leftVert (nabla cdot {bf{F}})({({sj},{sk})}^{T})rightVert }_{1}$$
(18)
the place (J) and (Okay) symbolize the variety of grid factors alongside the respective dimensions.
Due to this fact, the general loss operate used for alignment module coaching is outlined in keeping with:
$${L}_{{rm{prepare}}}={L}_{{rm{recon}}}+{lambda }_{{rm{easy}}}{L}_{{rm{easy}}}+{lambda }_{{rm{div}}}{L}_{{rm{div}}}$$
(19)
the place ({lambda }_{{rm{easy}}}) and ({lambda }_{{rm{div}}}) are weight elements which can be utilized to steadiness the contribution of various loss.
Neighborhood enrichment
We calculated neighborhood enrichment with the Squidpy bundle66 to evaluate the spatial relationships between clusters.
Chamfer distance
The chamfer distance is used to guage the similarity between two units of factors. It calculates the sum of the minimal squared Euclidean distances from every level in a single set to the closest level within the different set.
$${D}_{{rm{chamfer}}}left(P,Qright)=frac{1} Pmathop{sum }limits_{pin P},mathop{min }limits_{qin Q}parallel p-q{parallel }^{2}+frac{1} mathop{sum }limits_{qin Q},mathop{min }limits_{pin P}parallel q-p{parallel }^{2}$$
(20)
the place (P={{p}_{1},{p}_{2},ldots ,{p}_{m}) and (Q={{q}_{1},{q}_{2},ldots ,{q}_{n}}) symbolize two units of factors. The space is computed by summing the minimal squared distances between the factors within the two units and averaging over the variety of factors in every set. Particularly, we calculate the chamfer distance between the goal aligned factors and the precise aligned factors. It’s notable that 3d-OT nonetheless exhibits superior efficiency when utilizing the reconstructed aligned place info as an alternative of calculations based mostly on the unique place info (beneath the identical variety of alignments, calculations utilizing the unique coordinates end in a smaller chamfer distance).
Spatial clustering
Clustering was carried out utilizing the mclust R bundle by way of a Python–R interface. Particularly, mclust67 was employed to establish clusters based mostly on both the single-modality illustration generated by the PointNet++ Encoder or the fused-modality illustration built-in by way of the MLP module. The variety of clusters was manually specified for benchmarking totally different strategies. To seize identified organic constructions and cell sorts, we additional iteratively examined totally different cluster numbers to attain optimum efficiency.
3D reconstruction
We carried out 3D reconstruction utilizing the 3d-OT alignment pipeline. By pairwise alignment of carefully associated spatial transcriptomic slices, we set up a exact mapping of corresponding cells, reflecting developmental development when samples differ primarily in ontogenetic phases. By way of sequential alignment of a sequence of corresponding tissue slices representing distinct developmental phases throughout the identical pipeline, we reconstruct complete spatiotemporal developmental trajectories for the goal organism.
Parameters of 3d-OT
3d-OT was educated for 1,150 epochs for single-omics area identification. For the DLPFC, breast most cancers and mouse visible cortex datasets, the highest 3,000 spatially variable genes (SVGs) had been chosen, and the standardized expression matrix was used as enter. The spatial graph was constructed with neighbors = 16. For multi-omics area identification, coaching settings diversified by dataset. For spatial ATAC–RNA-seq and spatial CUT&Tag–RNA-seq datasets, the mannequin was educated for 300 epochs with neighbors = 16 and studying fee = 0.001. Simulated multi-omics datasets had been educated for 600 epochs with neighbors = 16 and studying fee = 0.001. For the human lymph node A1 dataset, 500 epochs, neighbors = 6, and studying fee = 0.001 had been used. These parameter selections had been optimized to steadiness coaching stability and convergence pace.
Within the alignment module of 3d-OT, a Euclidean distance-based threshold operator was utilized to filter candidate finish spots. For slices with comparatively easy spatial construction, the 50 nearest neighbors on the goal slice had been chosen as candidates, from which the highest 5 most comparable spots had been used to reconstruct the ultimate finish place. For slices with substantial decision variations or weak spatial smoothness, 500–1,000 nearest neighbors had been thought-about as candidates, and the highest 5 most comparable spots had been retained for reconstruction. The alignment module was educated with the next parameters: studying fee = 0.0001, easy loss weight = 1, divergence loss weight = 1, divergence neighbors = 8, easy neighbors = 32 and most iteration steps within the Sinkhorn algorithm = 100.
Parameters of baseline strategies
For all baseline strategies, we adhered to the parameter settings advisable of their unique publications at any time when doable. For parameters not explicitly specified or for datasets requiring adaptation, we carried out a restricted grid search inside an inexpensive vary to make sure aggressive and truthful efficiency in opposition to our proposed methodology.
GraphST was educated for 600 epochs with default parameters, utilizing the total gene expression matrix as enter.
STAGATE was educated for 1,000 epochs with standardized expression values of the highest 3,000 SVGs, and the spatial graph was constructed with a radial cutoff of 150.
SpaGCN was educated for as much as 200 epochs utilizing the highest 3,000 SVGs as enter, with standardized expression values. The next parameters had been specified: p = 0.5, begin = 0.01, finish = 1,000, tol = 0.01, max_run = 100, and r_seed = t_seed = n_seed = 200.
SpaNCMG was educated for 1,000 epochs utilizing the highest 3,000 SVGs with standardized expression values, with two spatial graphs constructed utilizing rad_cutoff = 150 and k_cutoff = 6.
GAAEST was educated for 600 epochs utilizing the total gene expression matrix, with hyperparameters set as (alpha) = 10, (beta) = 1, (gamma) = 1 and (lambda) = 1.
SEDR was educated for 200 epochs with the highest 2,000 SVGs as enter; the highest 200 principal parts after standardization had been used, and the spatial graph was constructed with 12 neighbors.
SpatialGlue was educated for 1,600 epochs (neighbors = 3) in spatial ATAC–RNA-seq and spatial CUT&Tag–RNA-seq, for 200 epochs (neighbors = 6) in simulation information and for 200 epochs (neighbors = 6) in human lymph node.
COSMOS was educated with spatial_regularization_strength = 0.01, z_dim = 50, studying fee = 0.001, wnn_epoch = 500, total_epoch = 1,000, max_patience_bef = 10, max_patience_aft = 30, min_stop = 200, regularization_acceleration = True, edge_subset_sz = 1,000,000 and neighbors = 10.
For spatial slices alignment, SLAT constructed spatial graphs with k_cutoff = 10 and used function = ‘dpca’ and be part of = ‘inner’ to account for batch results. The ensuing embeddings had been subsequently utilized in spatial matching.
Concord was run with be part of = ‘inner’, max_iter_harmony = 20, and PCA preprocessing, adopted by spatial matching with the ensuing embeddings.
PASTE was run with (alpha) = 0.1, whereas PASTE2 was run with (alpha)= 0.1 and s = 0.7.
SANTO was educated with studying fee = 0.001, 20 epochs, okay = 10, (alpha)= 0.9, mode = ‘align’ and dimension=2.
Information and detailed strategies
Particulars on the datasets, downstream analyses, competing strategies and metrics used can be found in Supplementary Tables 1–4.
Reporting abstract
Additional info on analysis design is accessible within the Nature Portfolio Reporting Abstract linked to this text.